Tumor Suppressor Genes
Introduction
Tumor Suppressor Genes: A Historical Perspective
Complications of Tumor Suppression
Moderate and Low Penetrance Cancer Susceptibility Loci
Haploinsufficiency
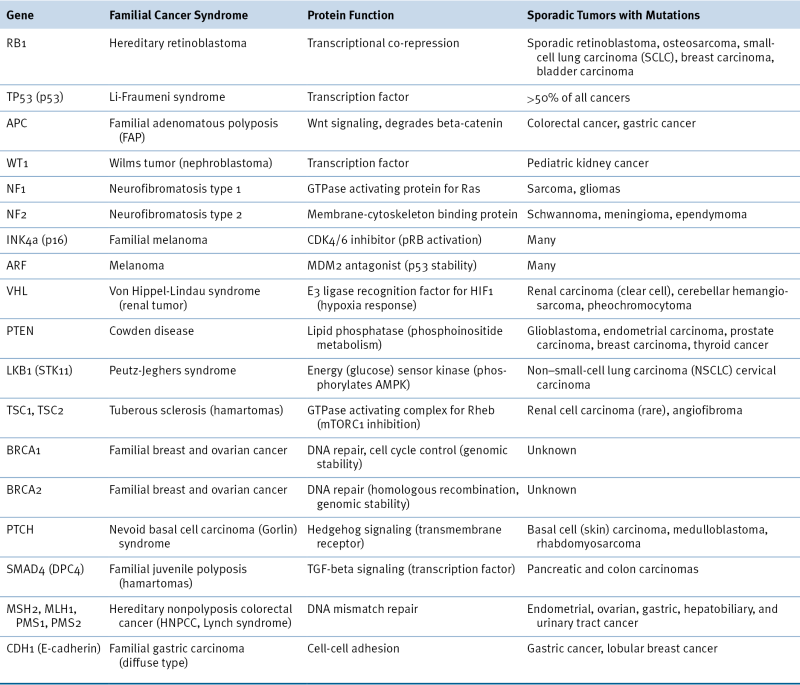
Table 3-1
Representative Tumor Suppressor Genes
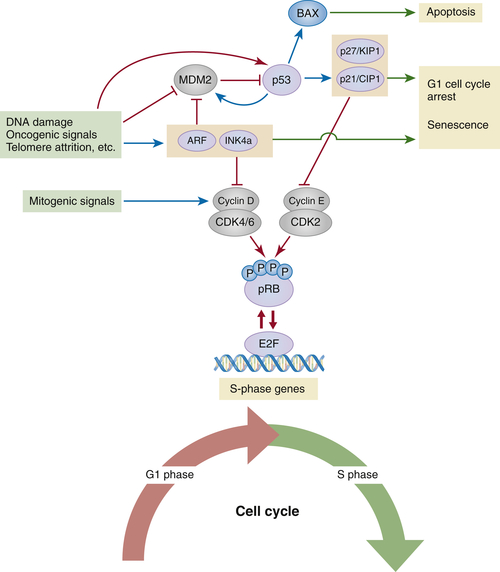
Interconnecting the pRB, p53, and mTORC1 Pathways
The pRB Tumor Suppressor Pathway
The p53 Tumor Suppressor Pathway
pRB and p53 Pathways in Action: Cellular Senescence
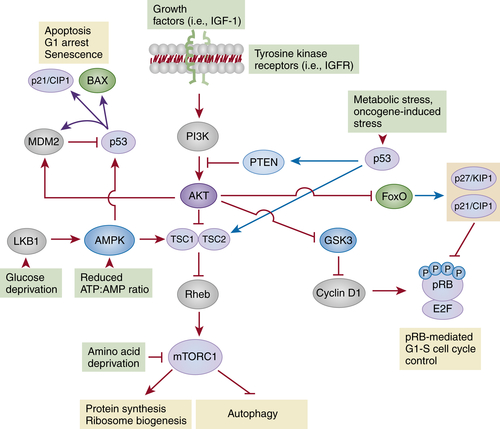
The mTORC1-Dependent Pathways
Epigenetic Modifications and Tumor Suppression
DNA Methylation
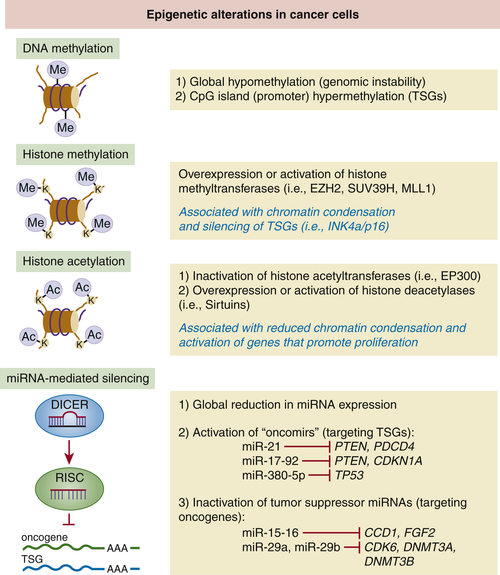
Histone Modifications
Micro-RNAs
Conclusions
References
1. Hanahan D., Weinberg R.A. Hallmarks of cancer: the next generation. Cell. 2011;144:646–674.
2. Vogelstein B., Kinzler K.W. Cancer genes and the pathways they control. Nat Med. 2004;10:789–799.
3. Kastan M.B., Bartek J. Cell-cycle checkpoints and cancer. Nature. 2004;432:316–323.
4. Sherr C.J. Principles of tumor suppression. Cell. 2004;116:235–246.
5. Hoeijmakers J.H. DNA damage, aging, and cancer. N Engl J Med. 2009;361:1475–1485.
6. Hoeijmakers J.H. Genome maintenance mechanisms for preventing cancer. Nature. 2001;411:366–374.
7. Steele R.J., Lane D.P. P53 in cancer: a paradigm for modern management of cancer. Surgeon. 2005;3:197–205.
8. Berger A.H., Knudson A.G., Pandolfi P.P. A continuum model for tumour suppression. Nature. 2011;476:163–169.
9. Fletcher O., Houlston R.S. Architecture of inherited susceptibility to common cancer. Nat Rev. 2010;10:353–361.
10. Balmain A. Cancer genetics: from Boveri and Mendel to microarrays. Nat Rev. 2001;1:77–82.
11. Harris H., Miller O.J., Klein G., Worst P., Tachibana T. Suppression of malignancy by cell fusion. Nature. 1969;223:363–368.
12. Allderdice P.W., Miller O.J., Miller D.A. et al. Chromosome analysis of two related heteroploid mouse cell lines by quinacrine fluorescence. J Cell Sci. 1973;12:263–274.
13. Klein G., Bregula U., Wiener F., Harris H. The analysis of malignancy by cell fusion. I. Hybrids between tumour cells and L cell derivatives. J Cell Sci. 1971;8:659–672.
14. Fearon E.R. Human cancer syndromes: clues to the origin and nature of cancer. Science. 1997;278:1043–1050.
15. Knudson A.G. Two genetic hits (more or less) to cancer. Nat Rev. 2001;1:157–162.
16. Cavenee W.K., Dryja T.P., Phillips R.A. et al. Expression of recessive alleles by chromosomal mechanisms in retinoblastoma. Nature. 1983;305:779–784.
17. Knudson Jr. A.G., Meadows A.T., Nichols W.W., Hill R. Chromosomal deletion and retinoblastoma. N Engl J Med. 1976;295:1120–1123.
18. Friend S.H., Bernards R., Rogelj S. et al. A human DNA segment with properties of the gene that predisposes to retinoblastoma and osteosarcoma. Nature. 1986;323:643–646.
19. Burkhart D.L., Sage J. Cellular mechanisms of tumour suppression by the retinoblastoma gene. Nat Rev. 2008;8:671–682.
20. Knudsen E.S., Knudsen K.E. Tailoring to RB: tumour suppressor status and therapeutic response. Nat Rev. 2008;8:714–724.
21. Levine A.J., Oren M. The first 30 years of p53: growing ever more complex. Nat Rev. 2009;9:749–758.
22. Goh A.M., Coffill C.R., Lane D.P. The role of mutant p53 in human cancer. J Pathol. 2011;223:116–126.
23. Meijers-Heijboer H., van den Ouweland A., Klijn J. et al. Low-penetrance susceptibility to breast cancer due to CHEK2(∗)1100delC in noncarriers of BRCA1 or BRCA2 mutations. Nat Genet. 2002;31:55–59.
24. Rahman N., Seal S., Thompson D. et al. PALB2, which encodes a BRCA2-interacting protein, is a breast cancer susceptibility gene. Nat Genet. 2007;39:165–167.
25. Anderson C.A., Pettersson F.H., Barrett J.C. et al. Evaluating the effects of imputation on the power, coverage, and cost efficiency of genome-wide SNP platforms. Am J Hum Genet. 2008;83:112–119.
26. Hao K., Schadt E.E., Storey J.D. Calibrating the performance of SNP arrays for whole-genome association studies. PLoS Genet. 2008;4:e1000109.
27. Chen Z., Trotman L.C., Shaffer D. et al. Crucial role of p53-dependent cellular senescence in suppression of Pten-deficient tumorigenesis. Nature. 2005;436:725–730.
28. Alimonti A., Carracedo A., Clohessy J.G. et al. Subtle variations in Pten dose determine cancer susceptibility. Nat Genet. 2010;42:454–458.
29. Bellacosa A., Godwin A.K., Peri S. et al. Altered gene expression in morphologically normal epithelial cells from heterozygous carriers of BRCA1 or BRCA2 mutations. Cancer Prev Res (Phila). 2010;3:48–61.
30. Venkatachalam S., Shi Y.P., Jones S.N. et al. Retention of wild-type p53 in tumors from p53 heterozygous mice: reduction of p53 dosage can promote cancer formation. EMBO J. 1998;17:4657–4667.
31. Malumbres M., Barbacid M. To cycle or not to cycle: a critical decision in cancer. Nat Rev. 2001;1:222–231.
32. Massague J. G1 cell-cycle control and cancer. Nature. 2004;432:298–306.
33. Malumbres M., Barbacid M. Cell cycle, CDKs and cancer: a changing paradigm. Nat Rev. 2009;9:153–166.
34. Ruas M., Peters G. The p16INK4a/CDKN2A tumor suppressor and its relatives. Biochim Biophys Acta. 1998;1378:F115–F177.
35. Lowe S.W., Sherr C.J. Tumor suppression by Ink4a-Arf: progress and puzzles. Curr Opin Genet Dev. 2003;13:77–83.
36. Larsson L.G. Oncogene- and tumor suppressor gene-mediated suppression of cellular senescence. Semin Cancer Biol. 2011;21(6):367–376.
37. Polager S., Ginsberg D. p53 and E2f: partners in life and death. Nat Rev. 2009;9:738–748.
38. Kuilman T., Michaloglou C., Mooi W.J., Peeper D.S. The essence of senescence. Genes Dev. 2010;24:2463–2479.
39. Feldser D.M., Greider C.W. Short telomeres limit tumor progression in vivo by inducing senescence. Cancer Cell. 2007;11:461–469.
40. Nardella C., Clohessy J.G., Alimonti A., Pandolfi P.P. Pro-senescence therapy for cancer treatment. Nat Rev. 2011;11:503–511.
41. Collado M., Serrano M. The power and the promise of oncogene-induced senescence markers. Nat Rev. 2006;6:472–476.
42. Michaloglou C., Vredeveld L.C., Mooi W.J., Peeper D.S. BRAF(E600) in benign and malignant human tumours. Oncogene. 2008;27:877–895.
43. Michaloglou C., Vredeveld L.C., Soengas M.S. et al. BRAFE600-associated senescence-like cell cycle arrest of human naevi. Nature. 2005;436:720–724.
44. Jones R.G., Thompson C.B. Tumor suppressors and cell metabolism: a recipe for cancer growth. Genes Dev. 2009;23:537–548.
45. Sabatini D.M. mTOR and cancer: insights into a complex relationship. Nat Rev. 2006;6:729–734.
46. Engelman J.A. Targeting PI3K signalling in cancer: opportunities, challenges and limitations. Nat Rev. 2009;9:550–562.
47. Cairns R.A., Harris I.S., Mak T.W. Regulation of cancer cell metabolism. Nat Rev. 2011;11:85–95.
48. Vousden K.H., Ryan K.M. p53 and metabolism. Nat Rev. 2009;9:691–700.
49. Taby R., Issa J.P. Cancer epigenetics. CA Cancer J Clin. 2010;60:376–392.
50. Feinberg A.P., Tycko B. The history of cancer epigenetics. Nat Rev. 2004;4:143–153.
51. Rodriguez-Paredes M., Esteller M. Cancer epigenetics reaches mainstream oncology. Nat Med. 2011;17:330–339.
52. Calin G.A., Croce C.M. MicroRNA signatures in human cancers. Nat Rev. 2006;6:857–866.
53. Lujambio A., Lowe S.W. The microcosmos of cancer. Nature. 2012;482:347–355.
54. Kasinski A.L., Slack F.J. Epigenetics and genetics. MicroRNAs en route to the clinic: progress in validating and targeting microRNAs for cancer therapy. Nat Rev. 2011;11:849–864.