9 Antigen-Specific Immune Responses
Antigen-specific immune responses provided by T cells and antibody expand the host protections provided by innate responses. The antigen-specific immune system is a randomly generated, coordinately regulated, inducible, and activatible system that ignores self-proteins but specifically responds to and protects against infection. When not working properly, the immune response can be unregulated, overstimulated, uncontrolled, reactive to self-proteins, unresponsive or poorly responsive to infections and become the cause of pathogenesis and disease. Almost any molecule has the potential to initiate an immune response. Once specifically activated by exposure to a new antigen, the immune response rapidly expands in strength, cell number, and specificity. For proteins, immune memory develops to allow more rapid recall upon rechallenge.
Antibody and the antibody-like T-cell receptor (TCR) molecules recognize antigens and act as receptors to activate the growth and functions of the cells expressing that molecule. Soluble forms of antibody in the blood, body fluids or secreted from mucosal membranes can inactivate and promote the elimination of toxins and microbes, especially when they are in the blood (bacteremia, viremia). T cells are important for activating and regulating innate and immune responses and for direct killing of cells expressing inappropriate antigens.
Although some molecules elicit only a limited antibody response (carbohydrates), proteins and protein-conjugated molecules (including carbohydrates) elicit a more complete immune response that includes T cells. Activation of a complete immune response is highly controlled because it uses a large amount of energy, and, once initiated, it develops memory and remains for most of a lifetime. Development of an antigen-specific immune response progresses from the innate responses through dendritic cells (DCs), which direct the T cells to tell other T cells, B cells, and other cells to grow and activate the necessary responses (Figure 9-1). Cell-receptor and cytokine-receptor interactions provide the necessary signals to activate cell growth and respond to the challenge. T cells tell the B cell which type of antibody to produce (IgG, IgE, IgA) and promote memory cell development.
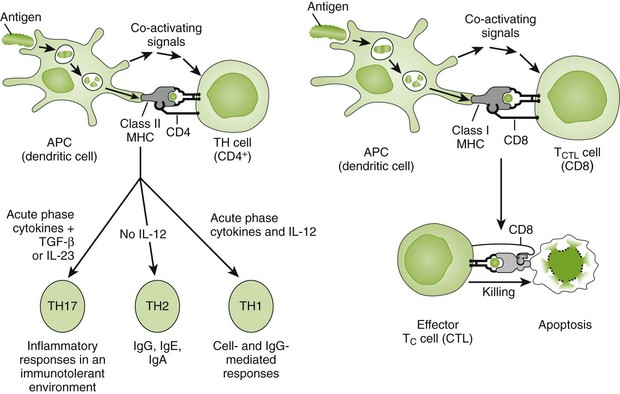
Figure 9-1 Activation of T-cell responses. The interaction of dendritic cells with CD4 or CD8 T cells initiates different immune responses, depending upon the cytokines produced by the dendritic cell. CD4 T cells mature to provide help to other cells with cytokine-mediated instructions. CD8 T cells can mature into cytolytic T cells (CTL). APC, Antigen-presenting cell; IL, interleukin; MHC, major histocompatibility complex; TGF-β, transforming growth factor-β.
(From Rosenthal KS, Tan M: Rapid reviews in microbiology and immunology, ed 3, Philadelphia, 2010, Elsevier.)
Immunogens, Antigens, and Epitopes
Almost all of the proteins and carbohydrates associated with an infectious agent, whether a bacterium, fungus, virus, or parasite, are considered foreign to the human host and have the potential to induce an immune response. A protein or carbohydrate that is recognized and sufficient to initiate an immune response is called an immunogen (Box 9-1). Immunogens may contain more than one antigen (e.g., bacteria). An antigen is a molecule that is recognized by specific antibody or the TCR on T cells. An epitope (antigenic determinant) is the actual molecular structure that interacts with a single antibody molecule or TCR. Within a protein, an epitope may be formed by a specific sequence (linear epitope) or a three-dimensional structure (conformational epitope). The TCR can recognize only linear epitopes. Antigens and immunogens usually contain several epitopes, each capable of binding to a different antibody molecule or TCR. As described later in this chapter, a monoclonal antibody recognizes a single epitope.
Box 9-1
Definitions
Adjuvant: substance that promotes immune response to immunogen
Antigen: substance recognized by immune response
Carrier: protein modified by hapten to elicit response
Epitope: molecular structure recognized by immune response
Hapten: incomplete immunogen that cannot initiate response but can be recognized by antibody
Immunogen: substance capable of eliciting an immune response
T-dependent antigens: antigens that must be presented to T and B cells for antibody production
T-independent antigens: antigens with large, repetitive structures (e.g., bacteria, flagellin, lipopolysaccharide, polysaccharide)
Not all molecules are immunogens. In general, proteins are the best immunogens, carbohydrates are weaker immunogens, and lipids and nucleic acids are poor immunogens. Haptens (incomplete immunogens) are often too small to immunize (i.e., initiate a response) an individual but can be recognized by antibody. Haptens can be made immunogenic by attachment to a carrier molecule, such as a protein. For example, dinitrophenol conjugated to bovine serum albumin is an immunogen for the dinitrophenol hapten.
During artificial immunization (e.g., vaccines), an adjuvant is used to enhance the response to antigen. Adjuvants usually prolong the presence of antigen in the tissue, promote uptake of the immunogen or activate DCs, macrophages, and lymphocytes. Some adjuvants mimic the activators (e.g., microbial ligands for Toll-like receptors) present in a natural immunization.
Some molecules will not elicit an immune response in an individual. During growth of the fetus, the body develops central immune tolerance toward self-antigens and any foreign antigens that may be introduced before maturation of the immune system. Later in life, peripheral tolerance develops to other proteins to prevent uncontrolled or autoimmune responses. For example, our immune response is tolerant of the food we eat; alternatively, eating steak would induce an antimuscle response.
The type of immune response initiated by an immunogen depends on its molecular structure. A primitive but rapid antibody response can be initiated toward bacterial polysaccharides (capsule), peptidoglycan, or flagellin. Termed T-independent antigens, these molecules have a large, repetitive structure that is sufficient to activate B cells directly to make antibody without the participation of T-cell help. In these cases, the response is limited to production of IgM antibody and fails to stimulate an anamnestic (booster) response. The transition from an IgM response to an IgG, IgE, or IgA response results from a big change in the B cell and is equivalent to differentiation of the cell. This requires help provided by T-cell interactions and cytokines. The antigen, therefore, must be recognized and stimulate both T and B cells. T-dependent antigens are proteins; they generate all five classes of immunoglobulins and can elicit memory and an anamnestic (secondary-booster) response.
In addition to the structure of the antigen, the amount, route of administration, and other factors influence the type of immune response, including the types of antibody produced. For example, oral or nasal administration of a vaccine across mucosal membranes promotes production of a secretory form of IgA (sIgA) that would not be produced on intramuscular administration.
T Cells
T cells were initially distinguished from B cells on the basis of their ability to bind sheep red blood cells through the CD2 molecule and form rosettes. These cells communicate through direct cell-to-cell interactions and with cytokines. T cells are defined through the use of antibodies that distinguish their cell surface molecules. The T-cell surface proteins include (1) the TCR, (2) the CD4 and CD8 co-receptors, (3) accessory proteins that promote recognition and activation, (4) cytokine receptors, and (5) adhesion proteins. All of these proteins determine the types of cell-to-cell interactions for the T cell and therefore the functions of the cell.
Development of T Cells
T-cell precursors are continuously developing into T cells in the thymus (Figure 9-2). Contact with the thymic epithelium and hormones, such as thymosin, thymulin, and thymopoietin II in the thymus, promote extensive proliferation and differentiation of the individual’s T-cell population during fetal development. While the T-cell precursors are in the thymus, each cell undergoes recombination of sequences within its TCR genes to generate a TCR unique to that cell. The epithelial cells in the thymus have a unique capacity to express most of the proteins of the human genome so that the developing T cells can be exposed to the normal repertoire of human proteins. T cells bearing nonfunctional TCRs, TCRs that cannot interact with major histocompatibility complex (MHC) molecules, or those that react too strongly with self-protein peptides (self-reactive) are forced into committing suicide (apoptosis). The surviving T cells differentiate into the subpopulations of T cells (Box 9-2). T cells can be distinguished by the type of T-cell antigen receptor, either consisting of γ and δ chains or α and β chains, and for α/β T cells, the presence of CD4 or CD8 co-receptors. T cells can be further distinguished by the cytokines they produce.
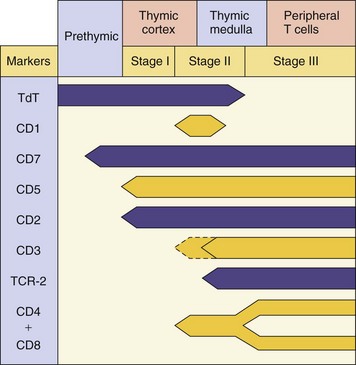
Figure 9-2 Human T-cell development. T-cell markers are useful for the identification of the differentiation stages of the T cell and for characterizing T-cell leukemias and lymphomas. TCR, T-cell receptor; TdT, cytoplasmic terminal deoxynucleotidyl transferase.
Box 9-2
T Cells
α/β T Cells
CD4: α/β TCR reactive with peptides on MHC II on antigen-presenting cell
CD4 CD25 Treg cells: control and limit expansion of immune response; promote tolerance and memory cell development
CD8: α/β TCR reactive with peptides presented on MHC I
NKT cells: α/β TCR reactive with glycolipids (mycobacteria) on CD1 molecules
MHC, Major histocompatibility complex; NK, natural killer; TCR, T-cell receptor.
T cells expressing the γ/δ TCR are present in blood, mucosal epithelium, and other tissue locations and are important for stimulating innate and mucosal immunity. These cells make up 5% of circulating lymphocytes but expand to between 20% and 60% of T cells during certain bacterial and other types of infections. The γ/δ TCR senses unusual microbial metabolites and initiates cytokine-mediated immune responses.
The α/β TCR is expressed on most T cells, and these cells are primarily responsible for antigen-activated immune responses. T cells with the α/β TCR are distinguished further by the expression of either a CD4 or a CD8 molecule.
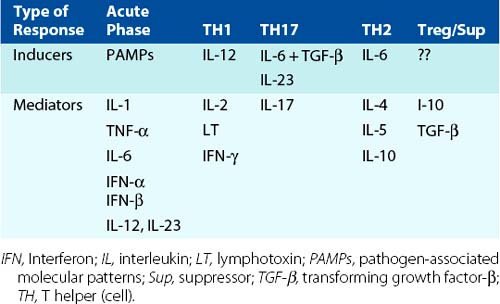
The helper T cells (CD4) activate and control immune and inflammatory responses by specific cell-to-cell interactions and by releasing cytokines (soluble messengers). Helper T cells interact with peptide antigens presented on class II MHC molecules expressed on antigen-presenting cells (APCs) (DCs, macrophages, and B cells) (see Figure 9-1). The repertoire of cytokines secreted by a specific CD4 T cell in response to antigenic challenge defines the type of CD4 T cell. Initially, TH0 cells produce cytokines to promote expansion of the cellular response and then can be converted to T cells producing other responses. TH1 cells produce interferon-γ (IFN-γ) to activate macrophages and DCs and promote responses that are especially important for controlling intracellular (mycobacterial and viral) and fungal infections and promoting certain subtypes of IgG antibody production. TH2 cells promote antibody responses. TH17 cells secrete interleukin (IL)-17 to activate neutrophils and promote antibacterial, antifungal responses and inflammation. T-regulator (Treg) cells express CD4 and CD25, prevent spurious activation of T cells, and control the immune response. The cytokines produced by each of these T-cell responses reinforce their own but may antagonize other responses. CD4 T cells can also kill target cells with its Fas ligand surface protein.
CD8 T cells are categorized as cytolytic and suppressor T cells but can also make cytokines similar to CD4 cells. Activated CD8 T cells “patrol” the body for virus-infected or tumor cells, which are identified by antigenic peptides presented by class I MHC molecules. Class I MHC molecules are found on all nucleated cells.
Cell Surface Receptors of T Cells
The TCR complex is a combination of the antigen recognition structure (TCR) and cell-activation machinery (CD3) (Figure 9-3). The specificity of the TCR determines the antigenic response of the T cell. Each TCR molecule is made up of two different polypeptide chains. As with antibody, each TCR chain has a constant region and a variable region. The repertoire of TCRs is very large and can identify a tremendous number of antigenic specificities (estimated to be able to recognize 1015 separate epitopes). The genetic mechanisms for the development of this diversity are also similar to those for antibody (Figure 9-4). The TCR gene is made up of multiple V (V1V2V3 … Vn), D, and J segments. In the early stages of T-cell development, a particular V segment genetically recombines with one or more D segments, deleting intervening V and D segments, and then recombines with a J segment to form a unique TCR gene. Like antibody, random insertion of nucleotides at the recombination junctions increases the potential for diversity and the possibility of producing inactive TCRs. Unlike antibody, somatic mutation does not occur for TCR genes. Only cells with functional TCRs will survive. Each T-cell clone expresses a unique TCR.
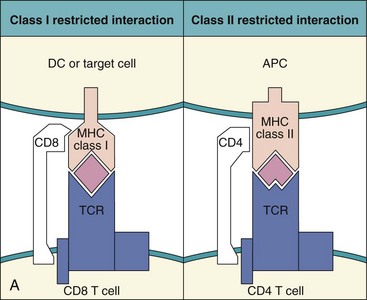
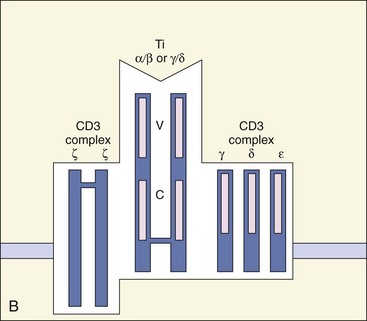
Figure 9-3 Major histocompatibility complex (MHC) restriction and antigen presentation to T cells. A, Left, Antigenic peptides bound to class I MHC molecules are presented to the T-cell receptor (TCR) on CD8 T-killer/suppressor cells. Right, Antigenic peptides bound to class II MHC molecules on the antigen-presenting cell (APC) (B cell, dendritic cell [DC], or macrophage) are presented to CD4 T-helper cells. B, T-cell receptor. The TCR consists of different subunits. Antigen recognition occurs through the α/β or γ/δ subunits. The CD3 complex of γ, δ, ε, and ζ subunits promotes T-cell activation. C, Constant region; V, variable region.
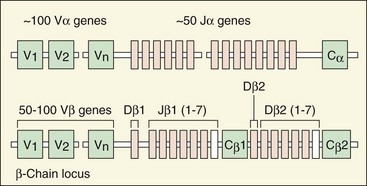
Figure 9-4 Structure of the embryonic T-cell receptor gene. Note the similarity in structure to the immunoglobulin genes. Recombination of these segments also generates a diverse recognition repertoire. C, Connecting sequences; J and D, segments; V, variable segments.
Unlike antibody molecules, the TCR recognizes a linear peptide epitope held within a cleft on the surface of either the MHC I or MHC II molecules. Presentation of the antigenic peptide requires specialized proteolytic processing of the protein (see later) and attachment to MHC II molecules by the antigen-presenting cell or MHC I molecules by all nucleated cells.
The CD3 complex is found on all T cells and consists of the γ-, δ-, ε-, and ζ-polypeptide chains. The CD3 complex is the signal transduction unit for the TCR. Tyrosine protein kinases (ZAP-70, Lck) associate with the CD3 complex when antigen is bound to the TCR complex, promote a cascade of protein phosphorylations, activation of phospholipase C (PLC), and other events. The products of cleavage of inositol triphosphate by PLC cause the release of calcium and activate protein kinase C and calcineurin, a protein phosphatase. Calcineurin is a target for the immunosuppressive drugs cyclosporine and tacrolimus. Activation of membrane G-proteins, such as Ras, and the consequences of the previously described cascades result in the activation of specific transcription factors in the nucleus, the activation of the T cell, and production of IL-2 and its receptor, IL-2R. These steps are depicted in Figure 9-5.
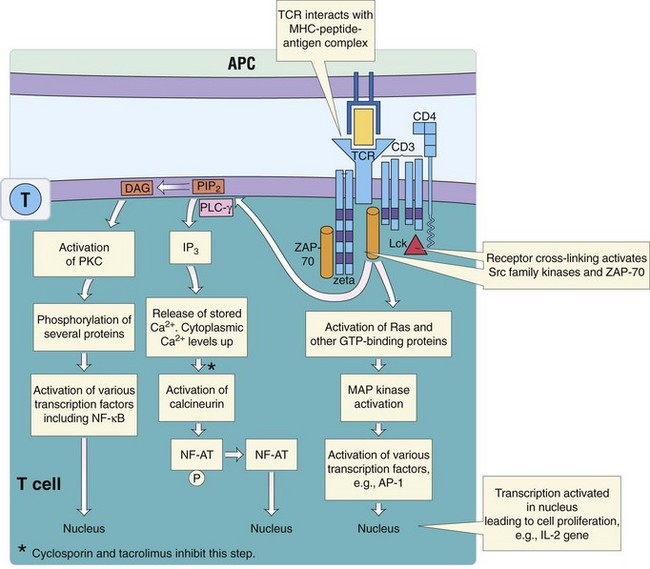
Figure 9-5 Activation pathways for T cells. Binding of major histocompatibility complex (MHC) II-peptide to CD4 and the T-cell receptor (TCR) activate kinase cascades and phospholipase C to activate the nuclear factor of activated T cells (NF-AT), nuclear factor-kappa B (NF-κβ), activation protein 1 (AP-1), and other transcription factors. APC, Antigen-presenting cell; DAG, diacylglycerol; GTP, guanosine triphosphate; IL-2, interleukin-2; IP3, inositol 1,4,5-triphosphate; Lck, lymphocyte-specific tyrosine protein kinase; MAP kinase, mitogen-activated protein kinase; PIP2, phosphatidylinositol 4,5-bisphosphate; PKC, protein kinase C; PLC-γ, phospholipase C-γ; ZAP, zeta-associated protein.
(Modified from Nairn R, Helbert M: Immunology for medical students, ed 2, Philadelphia, 2007, Mosby.)
The CD4 and CD8 proteins are co-receptors for the TCR because they facilitate the interaction of the TCR with the antigen-presenting MHC molecule and can enhance the activation response. CD4 binds to class II MHC molecules on the surface of APCs. CD8 binds to class I MHC molecules on the surface of APCs and target cells. Class I MHC molecules are expressed on all nucleated cells (see more on MHC later in this chapter). The cytoplasmic tails of CD4 and CD8 associate with a protein tyrosine kinase (Lck), which enhances the TCR-induced activation of the cell on binding to the APC or target cell. CD4 or CD8 is found on α/β T cells but not on γ/δ T cells.
Accessory molecules expressed on the T cell include several protein receptors on the cell surface that interact with proteins on APCs and target cells, leading to activation of the T cell, promotion of tighter interactions between the cells, or facilitation of the killing of the target cell. These accessory molecules are as follows:
1. CD45RA (native T cells) or CD45RO (memory T cells), a transmembrane protein tyrosine phosphatase (PTP)
2. CD28 or cytotoxic T-lymphocyte–associated protein 4 (CTLA-4) that binds to the B7 protein on APCs to deliver a co-stimulation or an inhibitory signal to the T cell
3. CD154 (CD40L), which is present on activated T cells and binds to CD40 on DCs, macrophages, and B cells to promote their activation
4. FasL, which initiates apoptosis in a target cell that expresses Fas on its cell surface.
Adhesion molecules tighten the interaction of the T cell with the APC or target cell and may also promote activation. Adhesion molecules include leukocyte function–associated antigen-1 (LFA-1), which interacts with the intercellular adhesion molecules (ICAM-1, ICAM-2, and ICAM-3) on the target cell. CD2 was originally identified by its ability to bind to sheep red blood cells (erythrocyte receptors). CD2 binds to LFA-3 on the target cell and promotes cell-to-cell adhesion and T-cell activation. Very late antigens (VLA-4 and VLA-5) are expressed on activated cells later in the response and bind to fibronectin on target cells to enhance the interaction.
T cells express receptors for many cytokines that activate and regulate T-cell function (Table 9-1). The cytokine receptors activate protein kinase cascades on binding of cytokine, to deliver their signal to the nucleus. The IL-2 receptor (IL-2R) is composed of three subunits. β/γ subunits are on most T cells (also natural killer [NK] cells) and have intermediate affinity for IL-2. The α subunit (CD25) is induced by cell activation to form a high-affinity α/β/γ IL-2R. Binding of IL-2 to the IL-2R initiates a growth-stimulating signal to the T cell, which also promotes the production of more IL-2 and IL-2R. CD25 is expressed on activated, growing cells, including the Treg subset of CD4 T cells (CD4+CD25+). Chemokine receptors distinguish different T cells and guide the cell to where it will reside in the body.
Initiation of T-Cell Responses
Antigen Presentation to T Cells
DCs provide the bridge between the innate and the immune responses, and the cytokines they produce determine the nature of the T-cell response. DCs are the only antigen-presenting cell that can initiate an antigen-specific T-cell response (see Box 9-2). DCs have octopus-like arms with large surface area (dendrites), produce cytokines, and have an MHC-rich cell surface to present antigen to T cells. Macrophages and B cells can present antigen to T cells but cannot activate a naïve T cell to initiate a new immune response.
Activation of an antigen-specific T-cell response requires a combination of cytokine and cell-to-cell receptor interactions (Table 9-2) initiated by the interaction of the TCR with MHC- bearing antigenic peptides. Class I and II MHC molecules provide a molecular cradle for the peptide. The CD8 molecule on cytolytic/suppressor T cells binds to and promotes the interaction with class I MHC molecules on target cells (see Figure 9-3A). The CD4 molecule on helper/delayed-type hypersensitivity (DTH) T cells binds to and promotes interactions with class II MHC molecules on APCs. The MHC molecules are encoded within the MHC gene locus (Figure 9-6). The MHC contains a cluster of genes important to the immune response.
Table 9-2 Antigen-Specific T-Cell Responses
| APC Activation of Naïve T Cells | ||
| Activation of the T Cell Requires Antigen, Co-Receptor, and Cytokine Interactions | ||
| DC | CD4 T Cell | Function |
| MHC II–peptide complex | TCR/CD4 | Antigen specificity |
| B7 | CD28 or CTLA4 | Activation or suppression |
| IL-1 | IL-1R | Activation |
| IL-6 | IL-6R | Overcomes Treg-induced tolerance |
| T-Cell Activation of APC | ||
| Enhanced Antigen Presentation of APCs, Enhanced Antimicrobial Activity of Macrophages, and Class Switch of Immunoglobulin Production by the B Cell Requires Antigen, Co-Receptor, and Cytokine Interactions | ||
| DC, Macrophage, or B Cell | CD4 T Cell | Function |
| MHC II–peptide complex | CD4T cell: TCR/CD4 | Antigen specificity |
| B7-1, B7-2 | CD28 | Activation of T cell |
| CD40 | CD40L | Activation of other functions in APC |
| IL-12 | Activation/reinforcement of TH1 responses | |
| IFN-γ | Activation of macrophages and B-cell class switch | |
| IL-4 | TH2 functions: growth and B-cell class switch | |
| IL-5 | TH2 functions: B-cell class switch | |
APC, Antigen-presenting cell; CTL, cytotoxic lymphocyte; DC, dendritic cell; IFN-γ, interferon-γ; IL, interleukin; MHC II, major histocompatibility complex II; TCR, T-cell receptor; TH, T helper (cell).
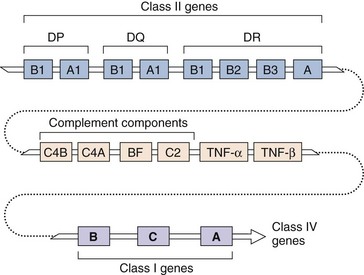
Figure 9-6 Genetic map of the major histocompatibility complex (MHC). Genes for class I and class II molecules, as well as complement components and tumor necrosis factor (TNF), are within the MHC gene complex.
Class I MHC molecules are found on all nucleated cells and are the major determinant of “self.” The class I MHC molecule, also known as HLA for human and H-2 for mouse, consists of two chains, a variable heavy chain and a light chain (β2-microglobulin) (Figure 9-7). Differences in the heavy chain of the HLA molecule between individuals (allotypic differences) elicit the T-cell response that prevents graft (tissue) transplantation. There are three major HLA genes: HLA-A, HLA-B, and HLA-C and other minor HLA genes. Each cell expresses a pair of different HLA-A, HLA-B, and HLA-C proteins, one from each parent, providing six different clefts to capture antigenic peptides. The heavy chain of the class I MHC molecule forms a closed-ended cleft, like a pita bread pocket, that holds a peptide of eight to nine amino acids. The class I MHC molecule presents antigenic peptides from within the cell (endogenous) to CD8-expressing T cells. Up-regulation of class I MHC molecules makes the cell a better target for T-cell action. Some cells (brain) and some virus infections (herpes simplex virus, cytomegalovirus) down-regulate the expression of MHC I antigens to reduce their potential as targets for T cells.
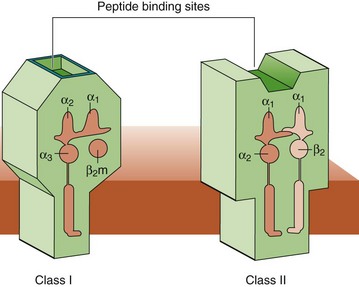
Figure 9-7 Structure of class I and class II major histocompatibility complex (MHC) molecules. The class I MHC molecules consist of two subunits, the heavy chain, and β2-microglobulin. The binding pocket is closed at each end and can only hold peptides of 8 to 9 amino acids. Class II MHC molecules consist of two subunits, α and β, and hold peptides of 11 or more amino acids.
Class II MHC molecules are normally expressed on antigen-presenting cells, cells that interact with CD4 T cells (e.g., macrophages, DCs, B cells). The class II MHC molecules are encoded by the DP, DQ, and DR loci and, like MHC I, are also co-dominantly expressed to produce six different molecules. The class II MHC molecules are a dimer of α and β subunits (see Figure 9-7). The chains of the class II MHC molecule form an open-ended peptide-binding cleft that resembles a hot-dog bun and holds a peptide of 11 to 12 amino acids. The class II MHC molecule presents ingested (exogenous) antigenic peptides to CD4-expressing T cells.
CD1 MHC molecules resemble MHC I molecules, have a heavy chain and a light chain (β2-microglobulin), but bind glycolipids rather than peptides. CD1 molecules are primarily expressed on DC and present antigen to the TCR on NKT (CD4−CD8−) cells. CD1 molecules are especially important for defense against mycobacterial infections.
Peptide Presentation by Class I and Class II MHC Molecules
Unlike antibodies that can recognize conformational epitopes, T-cell antigenic peptides must be linear epitopes. A T-cell antigen must be a peptide of 8 to 12 amino acids with a hydrophobic backbone that binds to the base of the molecular cleft of the class I or class II MHC molecule and exposes a T-cell epitope to the TCR. Because of these constraints, there may be only one T-cell antigenic peptide in a protein. All nucleated cells proteolytically process a set of intracellular proteins and display the peptides to CD8 T cells (endogenous route of antigen presentation) to distinguish “self,” “nonself,” inappropriate protein expression (tumor cell), or the presence of intracellular infections, whereas APCs process and present peptides from phagocytized proteins to CD4 T cells (exogenous route of antigen presentation) (Figure 9-8). DCs can cross these routes (cross-presentation) to present exogenous antigen to CD8 T cells to initiate antiviral and antitumor responses.
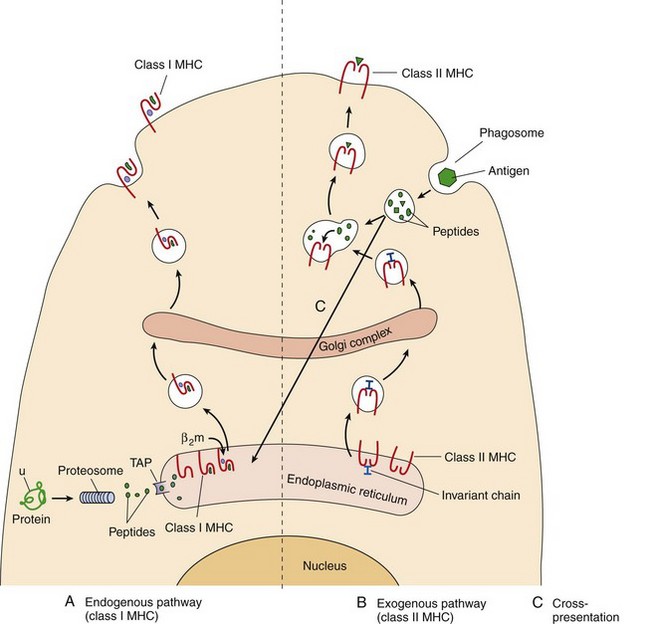
Figure 9-8 Antigen presentation. A, Endogenous: Endogenous antigen (produced by the cell and analogous to cell trash) is targeted by attachment of ubiquitin (u) for digestion in the proteosome. Peptides of eight to nine amino acids are transported through the transporter associated with antigen processing (TAP) into the endoplasmic reticulum (ER). The peptide binds to a groove in the heavy chain of the class I major histocompatibility complex (MHC) molecule, and the β2-microglobulin (β2m) binds to the heavy chain. The complex is processed through the Golgi apparatus and delivered to the cell surface for presentation to CD8 T cells. B, Exogenous: class II MHC molecules assemble in the ER with an invariant chain protein to prevent acquisition of a peptide in the ER. They are transported in a vesicle through the Golgi apparatus. Exogenous antigen (phagocytosed) is degraded in lysosomes, which then fuse with a vesicle containing the class II MHC molecules. The invariant chain is degraded and displaced by peptides of 11 to 13 amino acids, which bind to the class II MHC molecule. The complex is then delivered to the cell surface for presentation to CD4 T cells. C, Cross-presentation: Exogenous antigen enters the ER of dendritic cells and is presented on MHC I molecules to CD8 T cells.
Class I MHC molecules bind and present peptides that are degraded from cellular proteins by the proteosome (a protease machine) in the cytoplasm. These peptides are shuttled into the endoplasmic reticulum (ER) through the transporter associated with antigen processing (TAP). Most of these peptides come from misfolded or excess proteins (trash) marked by attachment of the ubiquitin protein. The antigenic peptide binds to the heavy chain of the class I MHC molecule. Then the MHC heavy chain can assemble properly with β2-microglobulin, exit the ER, and proceed to the cell membrane.
During a viral infection, large quantities of viral proteins are produced and degraded into peptides and become the predominant source of peptides occupying the class I MHC molecules to be presented to CD8 T cells. Transplanted cells (grafts) express peptides on their MHC molecules, which differ from those of the host and therefore may be recognized as foreign. Tumor cells often express peptides derived from abnormal or embryonic proteins, which may elicit responses in the host because the host was not tolerized to these proteins. Expression of these “foreign” peptides on MHC I at the cell surface allows the T cell to “see” what is going on within the cell.
Class II MHC molecules present peptides from exogenous proteins that were acquired by macropinocytosis, pinocytosis, or phagocytosis and then degraded in lysosomes by APCs. The class II MHC protein is also synthesized in the ER, but unlike MHC I, the invariant chain associates with MHC II to prevent acquisition of a peptide. MHC II acquires its antigenic peptide as a result of a merging of the vesicular transport pathway (carrying newly synthesized class II MHC molecules) and the lysosomal degradation pathway (carrying phagocytosed and proteolyzed proteins). The antigenic peptides displace a peptide from the invariant chain and associate with the cleft formed in the class II MHC protein; the complex is then delivered to the cell surface.
Cross-presentation of antigen is used by dendritic cells to present antigen to naïve CD8 T cells to initiate the response to viruses and tumor cells. After picking up antigen (including debris from apoptotic cells) in the periphery, the protein is degraded, its peptides enter the cytoplasm and are then shuttled through the TAP into the ER to bind to MHC I molecules.
The following analogy might aid in the understanding of antigen presentation: All cells degrade their protein “trash” and then display it on the cell surface on class I MHC trash cans. CD8 T cells “policing” the neighborhood are not alarmed by the normal, everyday peptide trash. A viral intruder would produce large amounts of viral peptide trash (e.g., beer cans, pizza boxes) displayed on class I MHC molecular garbage cans, which would alert the policing CD8 T cells. APCs (DCs, macrophages, and B cells) are similar to garbage collectors or sewage workers; they gobble up the neighborhood trash or lymphatic sewage, degrade it, display it on class II MHC molecules, and then move to a lymph node to present the antigenic peptides to the CD4 T cells in the “police station.” Foreign antigens would alert the CD4 T cells to release cytokines and activate an immune response.
Activation of CD4 T Cells and Their Response to Antigen
Activation of naïve T-cell responses is initiated by DCs and then expanded by other APCs. CD4 helper T cells are activated by the interaction of the TCR with antigenic peptide presented by class II MHC molecules on the APC (Figure 9-9A). The interaction is strengthened by the binding of CD4 to the class II MHC molecule and the linkage of adhesion proteins on the T cell and the APC. A co-stimulatory signal mediated by binding of B7 molecules on the macrophage, dendritic, or B-cell APC to CD28 molecules on the T cell is required to induce growth of the T cell as a fail-safe mechanism to ensure legitimate activation. B7 also interacts with CTLA4, which delivers an inhibitory signal. Activated APCs express sufficient B7 to fill up all the CTLA4 and then bind to the CD28. Cytokine signals (e.g., IL-1, IL-2, IL-6) are also required to initiate growth and overcome regulatory suppression of the cell. Proper activation of the helper T cell promotes production of IL-2 and increases expression of IL-2Rs on the cell surface, enhancing the cell’s own ability to bind and maintain activation by IL-2. Once activated, the IL-2 sustains the growth of the cell, and other cytokines influence whether the helper T cell matures into a TH1-, TH17-, or TH2-helper cell (see following section).
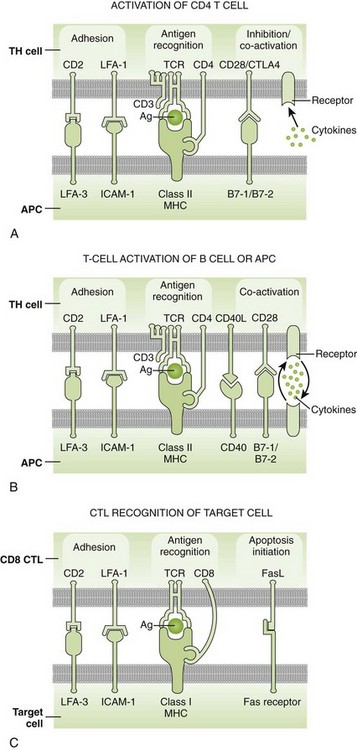
Figure 9-9 The molecules involved in the interaction between T cells and antigen-presenting cells (APCs). A, Initiation of a CD4 T-cell response. Initiation of a CD8 T-cell response is similar, but CD8 and the T-cell receptor (TCR) interact with peptide major histocompatibility complex (MHC) I and the peptide that it holds. B, CD4 T-cell helper binding to a B cell, dendritic cell, or macrophage. C, CD8 T-cell binding to target cell. The Fas–FasL interaction promotes apoptosis. Cell surface receptor-ligand interactions and cytokines are indicated with the direction of their action. Ag, Antigen; CTLA4, cytotoxic T lymphocyte A4; ICAM-1, intercellular adhesion molecule-1; LFA-1, leukocyte function–associated antigen-1.
(From Rosenthal KS, Tan M: Rapid reviews in microbiology and immunology, ed 3, Philadelphia, 2010, Elsevier.)
Partial activation (TCR interaction with MHC peptide) without co-stimulation leads to anergy (unresponsiveness) or apoptotic death (cell suicide) of the T cell. This is a mechanism for (1) eliminating self-reactive T cells in the thymus and (2) promoting the development of tolerance to self-proteins. In addition, binding of the CTLA-4, instead of CD28, on T cells with B7 on target or APC cells can result in anergy toward the antigen.
Once activated, the CD4 T cells exit the T-cell sites of the lymph node and enter the blood or move to B-cell zones of the lymph nodes and spleen. Antigen presentation initiates close interactions between the T cell and APC that allow the CD40L and CD28 molecules on the T cell to bind CD40 and B7 molecules on the APC. These interactions stimulate the mutual activation of the T cell and the APC (Figure 9-9B). This interaction and the cytokines produced by the T cell will determine the function of the macrophages and DC and which immunoglobulin the B cell will produce.
CD4 T-Helper Cell Functions
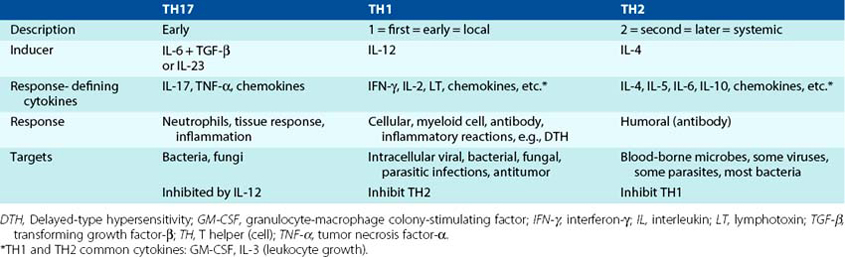
The CD4 T cells promote the expansion of the immune response with cell growth–promoting cytokines and define the nature of the response with other cytokines. CD4 T cells start as a TH0 cell that can develop into TH1, TH2, TH17, and other TH cells with different functions, as determined by the initial DC and cytokine interactions. The different types of TH cells are defined by the cytokines they secrete and thus the responses that they induce (Figure 9-10 and Table 9-3; also see Figure 9-1 and Table 9-1).
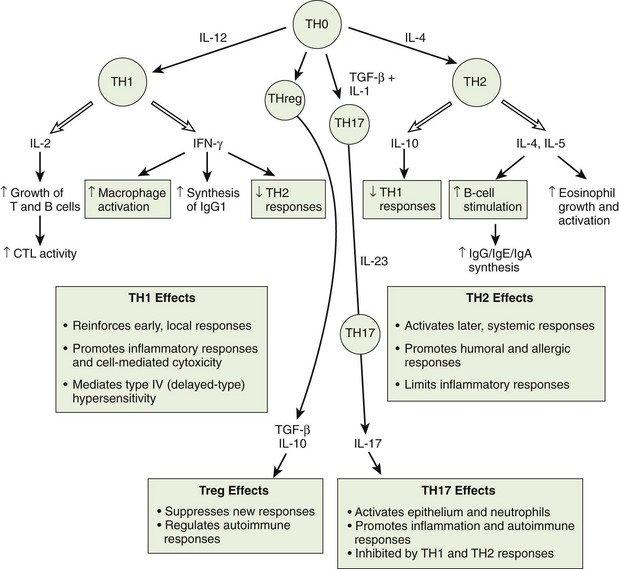
Figure 9-10 T-cell responses are determined by cytokines. Dendritic cells initiate and determine the type of CD4 T-cell responses by the cytokines that they produce. Similarly, T cells tell other cells what to do with other cytokines. The response-defining cytokines are indicated. ↑, Increase; ↓, decrease; CTL, cytotoxic T lymphocyte; IFN-γ, interferon-γ; IgG/IgE/IgA, immunoglobulin G/E/A; IL, interleukin; TGF-β, transforming growth factor-β; TH, T helper (cell).
(From Rosenthal KS, Tan M: Rapid reviews in microbiology and immunology, ed 3, Philadelphia, 2010, Elsevier.)
The primary role of the TH0 cells is to expand the immune response by producing cytokines that promote lymphocyte growth and activate DCs, including IL-2, IFN-γ, and IL-4. Once activated, the TH1 and TH2 cells produce cytokines that expand innate and immune responses (granulocyte-macrophage colony-stimulating factor [GM-CSF], tumor necrosis factor-α [TNF-α], and IL-3) and response-defining cytokines that expand the response (autocrine), but they inhibit the development of the other type of CD4 T cell.
Activation of TH1 responses requires IL-12 produced by DCs and macrophages and antigen presentation to CD4 T cells. TH1 cells are characterized by secretion of IL-2, IFN-γ, and TNF-β (lymphotoxin [LT]). These cytokines stimulate inflammatory responses and the production of a specific subclass of IgG that binds to Fc receptors on neutrophils and NK cells and can fix complement. IFN-γ, also known as macrophage activation factor, reinforces TH1 responses by promoting more IL-12 production, creating a self-sustaining cycle. TNF-β can activate neutrophils. TH1 cells are inhibited by IL-4 and IL-10, which is produced by TH2 cells. Activated TH1 cells also express the FasL ligand, which can interact with the Fas protein on target cells to promote apoptosis (killing) of the target cell and the CCR5 chemokine receptor that promotes relocation to sites of infection.
The TH1 response (1 meaning early) usually occurs early in response to an infection and activates both cellular and antibody responses. The TH1 responses amplify local inflammatory reactions and DTH reactions by activating macrophages, NK cells, and CD8 cytotoxic T cells and also expand the immune response by stimulating growth of B and T cells with IL-2. The inflammatory responses and antibody stimulated by TH1 responses are important for eliminating intracellular infections (e.g., viruses, bacteria, and parasites) and fungi but are also associated with cell-mediated autoimmune inflammatory diseases (e.g., multiple sclerosis, Crohn disease).
Initial antibacterial and antifungal responses are mediated by the TH17 cells. These are CD4 T-helper cells stimulated by IL-6 plus transforming growth factor (TGF)-β or IL-23 instead of IL-12. IL-23 is in the IL-12 family of cytokines. TH17 cells make cytokines, such as IL-17, IL-22, IL-6, and TNF-α, and proinflammatory chemokines, which activate neutrophils and promote inflammatory responses. TH17 responses would also provide protection in immunoprivileged sites, such as the eye, where there is an abundance of TGF-β. TH17 responses are associated with cell-mediated autoimmune inflammatory diseases, such as rheumatoid arthritis.
The TH2 response (2 meaning second) occurs later in response to infection and acts systemically through antibody-mediated responses. The TH2 response occurs in the absence of an IL-12/IFN-γ signal from innate responses, and then IL-4 reinforces the continuation of TH2 responses. TH2 cell development is inhibited by IFN-γ. The TH2 response may be stimulated later in an infection, when antigen reaches the lymph nodes and is presented by DCs, macrophages, and B cells. B cells expressing specific cell surface antibody can capture, process, and present antigen to TH2 cells to establish an antigen-specific circuit, stimulating the growth and clonal expansion of the helper T cells and B cells, which recognize the same antigen. TH2 cells release IL-4, IL-5, IL-6, and IL-10 cytokines that promote humoral (systemic) responses. These cytokines stimulate the B cell to undergo recombination events within the immunoglobulin gene to switch from production of IgM and IgD to production of specific subtypes of IgG, IgE, or IgA. TH2 responses are associated with production of IgE, which is useful for antihelminth responses but mediates allergies. TH2 responses can exacerbate an intracellular infection (e.g., Mycobacterium leprae, Leishmania) by prematurely shutting off protective TH1 responses.
Treg cells expressing CD4+CD25+ are antigen-specific suppressor cells. These cells prevent the development of autoimmune responses by producing TGF-β and IL-10, help to keep T-cell responses under control, and promote memory cell development. Other TH responses, such as TH9, TH22, and TFH (T-follicular helper), have been described, and their names refer to the primary cytokine that they produce or the functions promoted by the cytokine. TFH cells provide help to B cells within the follicles of the lymph node.
CD8 T Cells
CD8 T cells include cytotoxic T lymphocytes (CTLs) and suppressor cells. CTLs are part of the TH1 response and are important for eliminating virally infected cells and tumor cells. CD8 T cells can also secrete TH1-like cytokines. Less is known about suppressor cells.
The CTL response is initiated when naïve CD8 T cells in the lymph node are activated by antigen-presenting DCs and cytokines produced by TH1 CD4 T cells, including IL-2 (similar to activation of CD4 T cells as in Figure 9-9). Presentation of the antigen on MHC I may be the result of a virus infection or by cross-presentation of an antigen acquired at the site of infection or tumor by a DC. The activated CD8 T cells divide and differentiate into mature CTLs. During a viral challenge of mice, the numbers of specific CTLs will increase up to 100,000 times. When the activated CTL finds a target cell, it binds tightly through interactions of the TCR with antigen-bearing class I MHC proteins and adhesion molecules on both cells (similar to the closing of a zipper). Granules containing toxic molecules, granzymes (esterases), and a pore-forming protein (perforin) move to the site of interaction and release their contents into the pocket (immune synapse) formed between the T cell and target cell. Perforin generates holes in the target cell membrane to allow the granule contents to enter and induce apoptosis (programmed cell death) in the target cell. CD8 T cells can also initiate apoptosis in target cells through the interaction of the FasL on the T cell with the Fas protein on the target cell surface. FasL is a member of the TNF family of proteins, and Fas is a member of the TNF receptor family of proteins. Apoptosis is characterized by degradation of the target cell DNA into discrete fragments of approximately 200 base pairs and disruption of internal membranes. The cells shrink into apoptotic bodies, which are readily phagocytosed by macrophages and DCs. Apoptosis is a clean method of cell death, unlike necrosis, which signals neutrophil action and further tissue damage. TH1 CD4 T cells and NK cells also express FasL and can initiate apoptosis in target cells.
Suppressor T cells provide antigen-specific regulation of helper T-cell function through inhibitory cytokines and other means. Like CTLs, suppressor T cells interact with class I MHC molecules.
NKT Cells
NKT cells are like a hybrid between NK cells and T cells. They express an NK cell marker, NK1.1 and an α/β TCR. Unlike other T cells, the TCR repertoire is very limited. They may express CD4, but most lack CD4 and CD8 molecules (CD4−CD8−). The TCR of most NKT cells reacts with CD1 molecules, which present microbial glycolipids and glycopeptides. Upon activation, NKT cells release large amounts of IL-4 and IFN-γ. NKT cells help in the initial responses to infection and are very important for defense against mycobacterial infections.
B Cells and Humoral Immunity
The primary molecular component of the humoral immune response is antibody. B cells and plasma cells synthesize antibody molecules in response to challenge by antigen. Antibodies provide protection from rechallenge by an infectious agent, block spread of the agent in the blood, and facilitate elimination of the infectious agent. To accomplish these tasks, an incredibly large repertoire of antibody molecules must be available to recognize the tremendous number of infectious agents and molecules that challenge our bodies. In addition to interacting specifically with foreign structures, the antibody molecules must also interact with host systems and cells (e.g., complement, macrophages) to promote clearance of antigen and activation of subsequent immune responses (Box 9-3). Antibody molecules also serve as the cell surface receptors that stimulate the appropriate B-cell antibody factories to grow and produce more antibody in response to antigenic challenge.
Box 9-3
Antimicrobial Actions of Antibodies
Are opsonins: promote ingestion and killing by phagocytic cells (lgG)
Neutralize (block attachment) bacteria, toxins, and viruses
Agglutinate bacteria: may aid in clearing
Render motile organisms nonmotile
Combine with antigens on the microbial surface and activate the complement cascade, thus inducing an inflammatory response, bringing fresh phagocytes and serum antibodies into the site
Combine with antigens on the microbial surface, activate the complement cascade, and anchor the membrane attack complex involving C5b to C9
Immunoglobulin Types and Structures
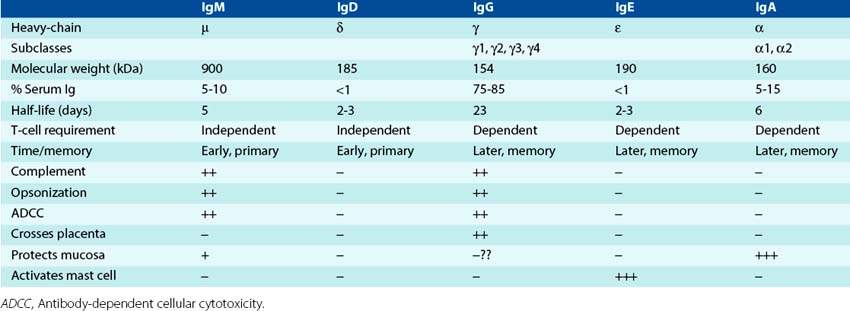
Immunoglobulins are composed of at least two heavy chains and two light chains, a dimer of dimers. They are subdivided into classes and subclasses based on the structure and antigenic distinction of their heavy chains. IgG, IgM, and IgA are the major antibody forms, whereas IgD and IgE make up less than 1% of the total immunoglobulins. The IgA and IgG classes of immunoglobulin are divided further into subclasses based on differences in the Fc portion. There are four subclasses of IgG, designated as IgG1 through IgG4, and two IgA subclasses (IgA1 and IgA2) (Figure 9-11).
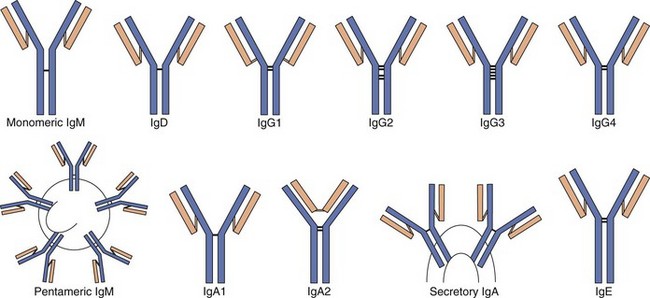
Figure 9-11 Comparative structures of the immunoglobulin classes and subclasses in humans. IgA and IgM are held together in multimers by the J chain. IgA can acquire the secretory component for the traversal of epithelial cells.
Antibody molecules are Y-shaped molecules with two major structural regions that mediate the two major functions of the molecule (see Figure 9-11; Table 9-4). The variable-region/antigen-combining site must be able to identify and specifically interact with an epitope on an antigen. A large number of different antibody molecules, each with a different variable region, are produced in every individual to recognize the seemingly infinite number of different antigens in nature. The Fc portion (stem of the antibody Y) interacts with host systems and cells to promote clearance of antigen and activation of subsequent immune responses. The Fc portion is responsible for fixation of complement and binding of the molecule to cell surface immunoglobulin receptors (FcR) on macrophages, NK cells, T cells, and other cells. For IgG and IgA, the Fc portion interacts with other proteins to promote transfer across the placenta and the mucosa, respectively (Table 9-5). In addition, each of the different types of antibody can be synthesized with a membrane-spanning portion to make it a cell surface antigen receptor.
Table 9-5 Fc Interactions with Immune Components
| Immune Component | Interaction | Function |
|---|---|---|
| Fc receptor | Macrophages | Opsonization |
| Polymorphonuclear neutrophils | Opsonization | |
| T cells | Activation | |
| Natural killer cells (antibody-dependent cellular cytotoxicity) | Killing | |
| Mast cells for immunoglobulin E | Allergic reactions, antiparasitic | |
| Complement | Complement system | Opsonization, killing (especially bacteria), activation of inflammation |
IgG and IgA have a flexible hinge region rich in proline and susceptible to cleavage by proteolytic enzymes. Digestion of IgG molecules with papain yields two Fab fragments and one Fc fragment (Figure 9-12). Each Fab fragment has one antigen-binding site. Pepsin cleaves the molecule, producing an F(ab′)2 fragment with two antigen-binding sites and a pFc′ fragment.
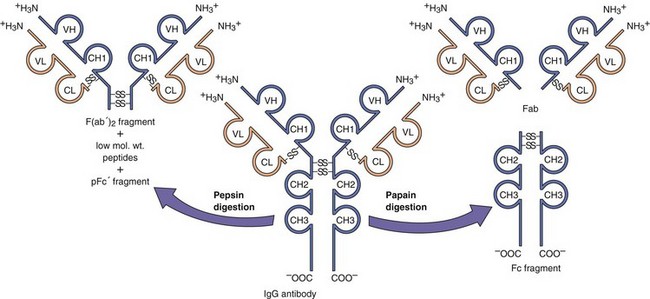
Figure 9-12 Proteolytic digestion of IgG. Pepsin treatment produces a dimeric F(ab′)2 fragment. Papain treatment produces monovalent Fab fragments and an Fc fragment. The F(ab′)2 and the Fab fragments bind antigen but lack a functional Fc region. The heavy chain is depicted in blue; the light chain in orange. mol. wt., Molecular weight.
The different types and parts of immunoglobulin can also be distinguished using antibodies directed against different portions of the molecule. Isotypes (IgM, IgD, IgG, IgA, IgE) are determined by antibodies directed against the Fc portion of the molecule (iso meaning the same for all people.) Allotypic differences occur for antibody molecules with the same isotype but contain protein sequences that differ from one person to another (in addition to the antigen-binding region). (All [“allo”] of us have differences.) The idiotype refers to the protein sequences in the variable region that generate the large number of antigen-binding regions. (There are many different idiots in the world.)
On a molecular basis, each antibody molecule is made up of heavy and light chains encoded by separate genes. The basic immunoglobulin unit consists of two heavy (H) and two light (L) chains. IgM and IgA consist of multimers of this basic structure. The heavy and light chains of immunoglobulin are fastened together by interchain disulfide bonds. Two types of light chains—κ and λ—are present in all five immunoglobulin classes, although only one type is present in an individual molecule. Approximately 60% of human immunoglobulin molecules have κ light chains, and 40% have λ light chains. There are five types of heavy chains, one for each isotype of antibody (IgM, µ; IgG, γ; IgD, δ; IgA, α; and IgE, ε). Intrachain disulfide bonds define molecular domains within each chain. Light chains have a variable and a constant domain. The heavy chains have a variable and three (IgG, IgA) or four (IgM, IgE) constant domains. The variable domains on the heavy and light chains interact to form the antigen-binding site. The constant domains from each chain make up the Fc portion, provide the molecular structure to the immunoglobulin and define the interaction of the antibody molecule with host systems, hence its ultimate function. The heavy chain of the different antibody molecules can also be synthesized with a membrane-spanning region to make the antibody an antigen-specific cell surface receptor for the B cell.
Immunoglobulin D
IgD, which has a molecular mass of 185 kDa, accounts for less than 1% of serum immunoglobulins. IgD exists primarily as membrane IgD, which serves with IgM as an antigen receptor on early B-cell membranes to help initiate antibody responses by activating B-cell growth. IgD and IgM are the only isotypes that can be expressed together by the same cell.
Immunoglobulin M
IgM is the first antibody produced in response to antigenic challenge and can be produced in a T-cell–independent manner. IgM makes up 5% to 10% of the total immunoglobulins in adults and has a half-life of 5 days. It is a pentameric molecule with five immunoglobulin units joined by disulfide bonds and the J chain, with a total molecular mass of 900 kDa. Theoretically, this immunoglobulin has 10 antigen-binding sites. IgM is the most efficient immunoglobulin for fixing (binding) complement. A single IgM pentamer can activate the classical complement pathway. Monomeric IgM is found with IgD on the B-cell surface, where it serves as the receptor for antigen. Because IgM is relatively large, it remains in the blood and spreads inefficiently from the blood into tissue. IgM is particularly important for immunity against polysaccharide antigens on the exterior of pathogenic microorganisms. It also promotes phagocytosis and promotes bacteriolysis by activating complement through its Fc portion. IgM is also a major component of rheumatoid factors (autoantibodies).
Immunoglobulin G
IgG comprises approximately 85% of the immunoglobulins in adults. It has a molecular mass of 154 kDa, based on two L chains of 22,000 Da each and two H chains of 55,000 Da each. The four subclasses of IgG differ in structure (see Figure 9-11), relative concentration, and function. Production of IgG requires T-cell help. IgG, as a class of antibody molecules, has the longest half-life (23 days) of the five immunoglobulin classes, crosses the placenta, and is the principal antibody in the anamnestic (booster) response. IgG shows high avidity (binding capacity) for antigens, fixes complement, stimulates chemotaxis, and acts as an opsonin to facilitate phagocytosis.
Immunoglobulin A
IgA comprises 5% to 15% of the serum immunoglobulins and has a half-life of 6 days. It has a molecular mass of 160 kDa and a basic four-chain monomeric structure. However, it can occur as monomers, dimers, trimers, and multimers combined by the J chain (similar to IgM). In addition to serum IgA, a secretory IgA appears in body secretions and provides localized immunity. IgA production requires specialized T-cell help and mucosal stimulation. Adjuvants, such as cholera toxin and attenuated Salmonella bacteria, can promote an IgA response. IgA binds to a poly-Ig receptor on epithelial cells for transport across the cell. The poly-Ig receptor remains bound to IgA and is then cleaved to become the secretory component when secretory IgA is secreted from the cell. An adult secretes approximately 2 gm of IgA per day. Secretory IgA appears in colostrum, intestinal and respiratory secretions, saliva, tears, and other secretions. IgA-deficient individuals have an increased incidence of respiratory tract infections.
Immunoglobulin E
IgE accounts for less than 1% of the total immunoglobulins and has a half-life of approximately 2.5 days. Most IgE is bound to Fc receptors on mast cells, on which it serves as a receptor for allergens and parasite antigens. When sufficient antigen binds to the IgE on the mast cell, the mast cell releases histamine, prostaglandin, platelet-activating factor, and cytokines. IgE is important for protection against parasitic infection and is responsible for anaphylactic hypersensitivity (type 1) (rapid allergic reactions).
Immunogenetics
The antibody response can recognize as many as 108 structures but can still specifically amplify and focus a response directed to a specific challenge. The mechanisms for generating this antibody repertoire and the different immunoglobulin subclasses are tied to random genetic events that accompany the development (differentiation) of the B cell (Figure 9-13).
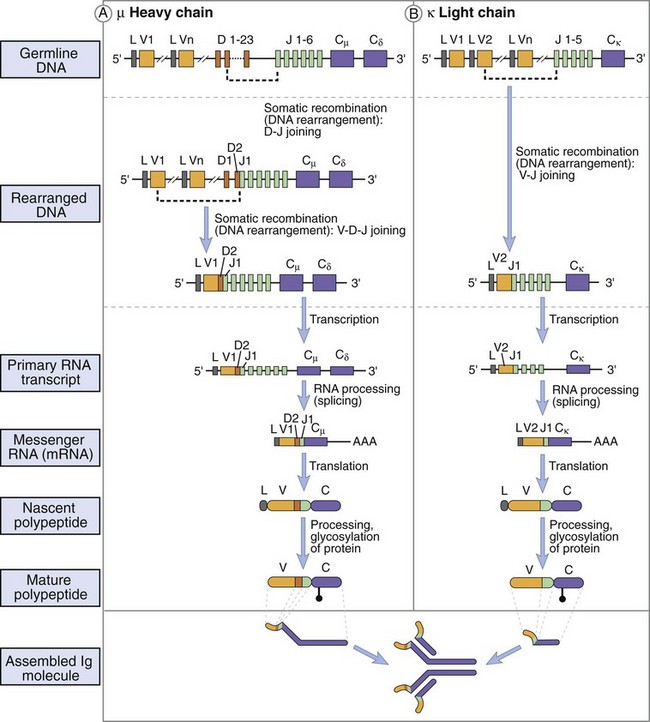
Figure 9-13 Immunoglobulin gene rearrangement to produce IgM. The germline immunoglobulin gene contains multiple V, D, and J genes that recombine and delete intervening sequences and juxtaposes the variable region sequences to the µ-δ heavy chain sequences during the development of the B cell in the bone marrow. T-cell help induces differentiation of the B cell and promotes genetic recombination and Ig class switching. Switch regions in front of the constant-region genes (including IgG subclasses) allow attachment of the preformed VDJ region with other heavy-chain constant-region genes, genetically removing the µ, δ, and other intervening genes. This produces an immunoglobulin gene with the same VDJ region (except for somatic mutation) but different heavy-chain genes. Splicing of messenger RNA (mRNA) produces the final IgM and IgD mRNA.
Human chromosomes 2, 22, and 14 contain immunoglobulin genes for κ, λ, and H chains, respectively. The germline forms of these genes consist of different and separate sets of genetic building blocks for the light (V and J gene segments) and heavy chains (V, D, and J gene segments), which are genetically recombined to produce the immunoglobulin variable regions. These variable regions are then recombined with the constant-region C gene segments. For the κ light chain, there are 300 V gene segments, 5 J gene segments, and 1 C gene segment. The number of λ gene segments for V and J is more limited. For the heavy chain, there are 300 to 1000 V genes, 12 D genes, and 6 (heavy-chain) J genes but only 9 C genes (one for each class and subclass of antibody [µ; δ; γ3, γ1, γ2, and γ4; ε; α1 and α2]). In addition, gene segments for membrane-spanning peptides can be attached to the heavy-chain genes to allow the antibody molecule to insert into the B-cell membrane as an antigen-activation receptor.
Production of the final antibody molecule in the pre-B and B cell requires genetic recombination at the deoxyribonucleic acid (DNA) level and posttranscriptional processing at the ribonucleic acid (RNA) level to assemble the immunoglobulin gene and produce the functional messenger RNA (mRNA) (see Figure 9-13). Each of the V, D, and J segments is surrounded by DNA sequences that promote directional recombination and loss of the intervening DNA sequences. Each of the recombination sites are then joined by randomly inserted nucleotides, which can enhance the diversity of sequences or disrupt the gene depending upon the number of inserted nucleotides. Juxtaposition of randomly chosen V and J gene segments of the light chains and the V, D, and J gene segments of the heavy chains produce the variable region of the immunoglobulin chains. These recombination reactions are analogous to matching and sewing together similar patterns from a long swatch of cloth, then cutting out the intervening loops of extra cloth. Somatic mutation of the immunoglobulin gene can also occur later in activated, growing B cells to add to the enormous number of possible coding sequences for the variable region and to fine-tune a specific immune response. The variable-region sequences (VDJ) are attached by recombination to the µ; δ; γ3, γ1, γ2, and γ4; ε; or α1 and α2 sequences of the C gene segments to produce a heavy-chain gene. In the pre-B and immature B cells, mRNAs are produced and contain the variable-region gene segments connected to the C gene sequences for µ and δ. Processing of the mRNA removes either the µ or δ, as if it were an intron, to produce the final immunoglobulin. The pre-B cell expresses cytoplasmic IgM, whereas the B cell expresses cytoplasmic and cell surface IgM and cell surface IgD. IgM and IgD are the only pair of isotypes that can be expressed on the same cell.
Class switching (IgM to IgG, IgE, or IgA) occurs in mature B cells in response to different cytokines produced by TH1 or TH2 CD4 helper T cells (see Figure 9-13). Each of the C gene segments, except δ, is preceded by a DNA sequence called the switch site. After the appropriate cytokine signal, the switch in front of the µ sequence recombines with the switch in front of the γ3, γ1, γ2, or γ4; ε; or α1, or α2 sequences, creating a DNA loop that is subsequently removed. Processing of the RNA transcript yields the final mRNA for the immunoglobulin heavy-chain protein. For example, IgG1 production would result from excision of DNA containing the C gene segments Cµ, Cδ, and Cγ3 to attach the variable region to the γ1 C gene segment. Class switching changes the function of the antibody molecule (Fc region) but does not change its specificity (variable region).
The final steps in B-cell differentiation to memory cells or plasma cells do not change the antibody gene. Memory cells are long-lived, antigen-responsive B cells expressing the CD45RO surface marker. Memory cells can be activated in response to antigen later in life to divide and then produce its specific antibody. Plasma cells are terminally differentiated B cells with a small nucleus but a large cytoplasm filled with endoplasmic reticulum. Plasma cells are antibody factories.
Antibody Response
An initial repertoire of IgM and IgD immunoglobulins is generated in pre-B cells by the genetic events previously described. Expression of cell surface IgM and IgD accompany differentiation of the pre-B cell to the B cell. The cell surface antibody acts as an antigen receptor to trigger activation of the B cell through its associated signal transduction receptors, Ig-α (CD79a) and Ig-β (CD79b). A cascade of protein tyrosine kinases, phospholipase C, and calcium fluxes activate transcription and cell growth to mediate the activation signal. Other surface molecules, including the CR2 (CD21) complement (C3d) receptor, amplify the activation signal. The combination of these signals triggers the growth and increases the number of cells making antibodies to that antigen. In this manner, the B cells that best recognize the different epitopes of the antigen are selected to increase in number in a process termed clonal expansion.
Clonal expansion of the antigen-specific B cells increases the number of antibody factories making the relevant antibody, and the strength of the antibody response is thus increased. Activation of the B cells also promotes somatic mutation of the variable region, increasing the diversity of antibody molecules directed at the specific antigen. The B-cell clones that express antibody with the strongest antigen binding are preferentially stimulated. This selects a better antibody response.
T-independent antigens have repetitive structures that can cross-link sufficient numbers of surface antibody to stimulate growth of the antigen-specific B cells. Binding of the C3d component of complement to its receptor (CR2, CD21) facilitates the activation of the antibody response. In contrast, production of antibody to T-dependent antigens requires receptor interactions of the B cell with the helper T cell through CD40 (on the B cell), CD40L (T cell), and the action of cytokines. Different combinations of cytokines produced by helper T cells induce class switching. TH1-helper responses (IFN-γ) promote production of IgG. TH2-helper responses (IL-4, IL-5, IL-6) promote production of IgG, IgE, and IgA. IgA production is especially promoted by IL-5 and TGF-β (Figure 9-14). Memory cells are developed with T-cell help. Terminal differentiation produces the ultimate antibody factory, the plasma cell.
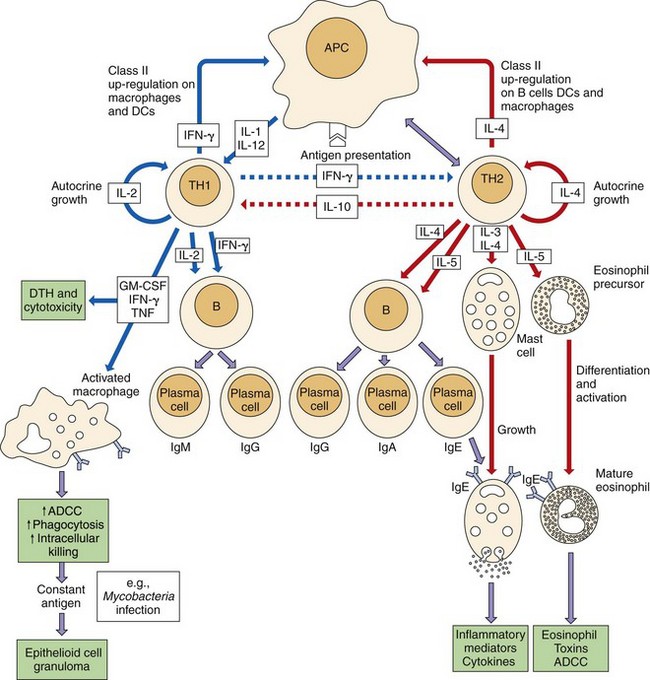
Figure 9-14 T-cell help determines the nature of the humoral immune response. Receptor-ligand interactions between T cells and B cells and cytokines associated with TH1 or TH2 determine the subsequent response. TH1 responses are initiated by interleukin (IL)-12 and delivered by interferon-γ (IFN-γ) and promote cell-mediated and IgG production (solid blue lines) and inhibit TH2 responses (dotted blue lines). IL-4 and IL-5 from TH2 cells promote humoral responses (solid red lines) and inhibit TH1 responses (dotted red lines). Mucosal epithelium promotes secretory IgA production. Colored boxes denote end results. ↑, Increase; ↓, decrease; ADCC, antibody-dependent cellular cytotoxicity; APC, antigen-presenting cell; CTL, cytotoxic T lymphocyte; DCs, dendritic cells; DTH, delayed-type hypersensitivity; GM-CSF, granulocyte-macrophage colony-stimulating factor; TNF, tumor necrosis factor.
During an immune response, antibodies are made against different epitopes of the foreign object, protein, or infectious agent. Specific antibody is a mixture of many different immunoglobulin molecules made by many different B cells (polyclonal antibody), each immunoglobulin molecule differing in the epitope it recognizes and the strength of the interaction. Antibody molecules that recognize the same antigen may bind with different strengths (affinity, monovalent binding to an epitope; avidity, multivalent binding of antibody to antigen).
Monoclonal antibodies are identical antibodies produced by a single clone of cells or by myelomas (cancers of plasma cells) or hybridomas. Hybridomas are cloned, laboratory-derived cells obtained by the fusion of antibody-producing spleen cells and a myeloma cell. In 1975, Kohler and Millstein developed the technique for producing monoclonal antibodies from B-cell hybridomas. The hybridoma is immortal and produces a single (monoclonal) antibody. This technique has revolutionized the study of immunology because it allows selection (cloning) of individual antibody-producing cells and their development into cellular factories for production of large quantities of that antibody. Monoclonal antibodies have been commercially produced for both diagnostic reagents and therapeutic purposes.
Time Course of the Antibody Response
The primary antibody response is characterized by the initial production of IgM. IgM antibodies appear in the blood within 3 days to 2 weeks after exposure to a novel immunogen. This is the only type of antibody elicited towards carbohydrates (bacterial capsule). Production of IgG, IgA, or IgE requires the development of a sufficient helper T-cell response to promote the class switch and requires approximately 8 days. The predominant serum antibody will be IgG antibodies (Figure 9-15). The first antibodies that are produced react with residual antigen and therefore are rapidly cleared. After the initial lag phase, however, the antibody titer increases logarithmically to reach a plateau.
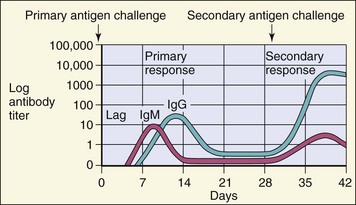
Figure 9-15 Time course of immune responses. The primary response occurs after a lag period. The IgM response is the earliest response. The secondary immune response (anamnestic response) reaches a higher titer, lasts longer, and consists predominantly of IgG.
Reexposure to an immunogen, a secondary response, induces a heightened antibody response (also termed anamnestic response). Activation of preformed memory cells yields a much more rapid production of antibody, which lasts longer and reaches a higher titer. The antibodies in a secondary response are principally of the IgG class.
What is wrong with each of the following statements, and why?
1. The laboratory tested a baby for IgM maternal antibodies.
2. An investigator attempted to use fluorescent-labeled F(ab′)2 fragments to locate class II MHC molecules on the cell surface of antigen-presenting cells without cross-linking (binding two molecules together) these cell surface molecules.
3. A patient is diagnosed as having been infected with a specific strain of influenza A (A/Bangkok/1/79/H3N2) on the basis of the presence of antiinfluenza IgG in serum taken from the patient at the initial visit (within 2 days of symptoms).
4. A patient was considered unable to use the complement systems because of a T-cell deficiency, which precluded the ability to promote class switching of B cells.
5. Analysis of immunoglobulin genes from B cells taken from the patient described in statement 4 did not contain recombined VDJ variable-region gene sequences.
6. A patient was considered to have a B-cell deficiency because serum levels of IgE and IgD were undetectable despite proper concentrations of IgG and IgM.
1. IgM molecules are too large to leave the plasma and cannot cross the placenta.
2. Native immunoglobulin and F(ab′)2 molecules are divalent or multivalent and can bind to more than one cell surface molecule, which will cross-link the cell surface.
3. IgG is only produced at approximately 6 days after a first-time infection and requires T-cell help. IgG could be present from a previous infection. IgM is produced early in an infection as part of a primary response and is a good indication of a first-time infection.
4. Although perforin is made by T cells and resembles C9, the complement components are synthesized by the liver and other cells and not by T cells such that a deficiency in T cells will not affect complement levels. Also, IgM fixes complement very well and will be produced in the absence of T cells.
5. Differentiation to a B cell requires recombination of the VDJ variable region, but this occurs without T-cell help.
6. The Fc portion of the immunoglobulin gene produces immunoglobulins in the order of IgM, IgD, IgG, IgE, and IgA. It would be unlikely that a lack of expression in IgD would occur without a lack in all the rest of the genes.