49 Orthomyxoviruses
Influenza
Key points
• Influenza viruses cause respiratory infections worldwide; there are three types: A, B and C.
• Influenza A is found in aquatic birds, poultry and pigs, and these play important roles in the epidemiology of human infections.
• Influenza viruses are segmented RNA viruses that regularly undergo genetic change.
• The surface proteins, the haemagglutinin and the neuraminidase, can show gradual change (antigenic drift) and sudden major change (antigenic shift).
• In humans, influenza A results in high morbidity and mortality rates in the winter months in temperate zones, but throughout the year in more tropical climates.
• The spread pattern may be pandemic, epidemic or sporadic/zoonotic.
• Pandemic influenza typically arises when a new virus from an animal host emerges into a susceptible human population. The impact of pandemics may be mild, moderate or severe, depending on the virulence properties of the virus.
• Zoonotic influenza A infections in humans may have a high case fatality and different clinical manifestations compared with seasonal influenza A.
• Since 1997, H5 avian influenza has been a major problem in the Far East, with strains from domestic poultry occasionally infecting humans directly, resulting in high mortality rates; this virus could become adapted to humans.
• Vaccines have been available for many years, recommended for annual boosting of high-risk groups in interpandemic periods to reduce mortality, morbidity and hospital admission rates.
• Antiviral drugs act to inhibit viral replication, and are clinically useful in treatment and prophylaxis.
The Orthomyxoviridae family comprises four genera: influenza A, B and C and thogotoviruses, within the negative sense RNA virus order Mononegavirales. All of these are small enveloped viruses with a segmented genome. The thogotoviruses are the most recently discovered genus and are found in mosquitoes, ticks and the banded mongoose, but are not so far associated with human disease and are not discussed further.
Influenza A, B and C host range
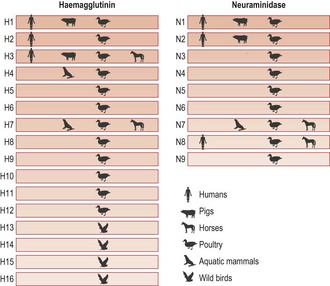
Influenza B and C have limited genetic diversity and occur almost exclusively in humans, which are the natural animal reservoir. Occasional transmission of influenza B to mammalian species such as seals and influenza C to pigs has been described. In contrast, there are many different subtypes of influenza A viruses, all of which naturally infect water-based wild birds, usually with very little disease. Influenza A viruses have a broad host range with the potential to infect a wide range of animal species. Viral subtypes are distinguished according to their surface proteins, the haemagglutinin (HA) and neuraminidase (NA). Sixteen HA subtypes and 9 NA subtypes are found in varying combinations circulating in wild birds. Only a limited number of influenza A subtypes have adapted to circulation in mammalian species, including humans, horses and swine (Fig. 49.1).
Avian influenza A subtypes are transmitted through a faeco–oral route and virus can be shed in high quantity in the environment following replication in the gastrointestinal tract. There is frequent spillover of avian influenza A subtypes into domestic poultry reservoirs, as a result of their shared habitat with wild birds. Replication in poultry may be associated with disease of varying severity or may be asymptomatic, depending on the viral subtype and the poultry species.
Nomenclature
The World Health Organization (WHO) system of nomenclature for influenza A includes the host of origin, geographical origin, strain number and year of isolation; then, in parentheses, the antigenic description of the haemagglutinin and neuraminidase is given, e.g. A/swine/Iowa/3/70 (H1N1). If isolated from a human host the origin is not given: e.g. A/Scotland/42/89 (H3N2).
Physical characteristics
In common with many enveloped viruses, influenza A, B and C are relatively labile and easily destroyed by common household cleaning agents and detergents. In the environment, influenza A virus can survive in cold seawater for several days, and is detectable in dust beyond one week although the infectivity of such material is not well understood.
The viruses
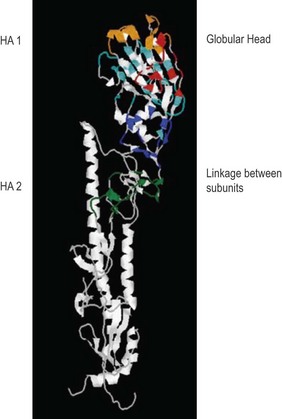
Influenza virions are spherical, 80–120 nm in diameter, but may be filamentous and up to one hundred fold longer (Fig. 49.2A). They have a helical nucleocapsid comprising, in A and B, eight segments of single-stranded RNA and, in C, seven segments. Also present within the virion particle is the viral RNA-dependent RNA polymerase enzyme; this is essential for infectivity as the viral RNA is of negative sense and therefore has to be transcribed to produce viral messenger RNA. The nucleocapsid is surrounded by the M1 protein shell, immediately exterior to which is a lipid envelope derived from the host cell. The viral M2 protein projects through the envelope to form ion channels, which assist virus entry through an endosomal route which involves pH changes. Two types of spike project from the lipid envelope, the haemagglutinin (HA) and the neuraminidase (NA) enzyme (Fig. 49.2B). The HA, so-called because the virus agglutinates certain species of erythrocyte, is about 10 nm in length and consists of trimers of identical glycoprotein subunits, each consisting of two polypeptide chains, HA1 and HA2 joined by a linkage site that may be a single basic amino acid, usually arginine, or a string of basic amino acids (Fig. 49.3).
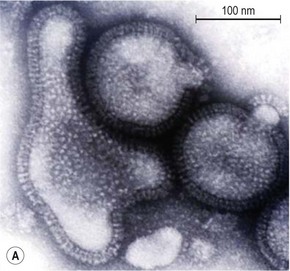
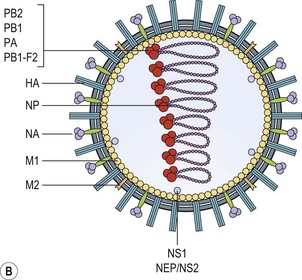
Fig. 49.2 (A) EM of influenza virus. (B) Schematic representation of virus particle and internal composition.
Influenza viruses bind to cells by the HA interacting with cell membrane receptors containing N-acetylneuraminic acid (sialic acid). The amino acid residues involved in receptor binding show variability according to host of origin. Differences in viral receptor binding characteristics have important biological significance for the transmission properties of the virus. Human influenza viruses recognize receptors that contain sialic acid attached to the penultimate sugar (usually lactose) via an α(2,6)-linkage, whereas avian strains prefer receptors with an α(2,3)-linkage (Fig. 49.4).
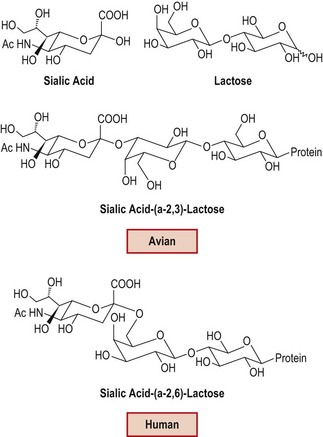
Fig. 49.4 Host virus receptor configuration in avian and human viruses, leading to differences in receptor binding preferences.
Between the HA spikes on the virus surface are the mushroom-shaped NA spikes. The NA protein is assembled from four subunits attached by a stalk containing a hydrophobic region which is anchored in the viral lipid envelope. The NA enzyme catalyses the cleavage of sialic acid and an adjacent sugar residue from glycoproteins found in mucus. This action allows the virus to permeate through the mucin overlying host epithelial surfaces. Neuraminidase activity is also important in the release of new virus particles from infected cells. Sialic acid is always present in newly synthesized virions, and its removal by NA prevents the new virus particles clumping through the binding to viral HA, thus assisting the spread of the virus from the original site of infection.
Virus variability
All influenza viruses replicate via virally encoded RNA dependent RNA polymerase enzymes. Such enzymes lack a proof reading (error correction) function, leading to high rates of mutation during replication, particularly for influenza A. Viruses carrying mutations which do not provide a severe replication disadvantage therefore evolve rapidly, a feature described as genetic drift. Genetic recombination between diverse influenza A viral subtypes occurs when more than one subtype infects a single host, creating new combinations of viral gene segments (reassortment) and altering the replication and virulence properties of the progeny virus. Together, the combination of genetic drift and segment reassortment provide the mechanism for generation of genetic diversity amongst influenza A viruses. These genetic mechanisms generate adaptive flexibility, which is reflected in the wide variety of animal species (host range) that can be infected. Different genetic lineages of influenza B and C also show some evidence of segment reassortment, but this is very much more limited than for influenza A and is not associated with significant biological diversity.
Epidemic and pandemic influenza
Major epidemics that may have been caused by influenza viruses have been described for more than 2000 years. Historically, it was rationalized that such unexpected events occurred under the influence of the stars, hence the term influenza (Italian for influence). It is well recognized that epidemics vary in severity. Occasionally infection arising from a new strain spreads throughout the world and causes very significant marked waves of illness; such pandemics have been recognized at irregular intervals.
Pandemics occurred in 1918, 1957 and 1968 with the emergence of H1N1 Spanish influenza, H2N2 and H3N2 respectively, and most recently in 2009, with the emergence of H1N1 from swine (H1N1 2009pdm) into the human population. The great pandemic of 1918–1919 was particularly severe, killing 20–40 million people as it spread over a few years. In 1957, the emergence of H2N2 as a pandemic virus displaced circulating H1N1, which disappeared until 1977, when it re-emerged after 20 years (Fig. 49.5). The emergence of new influenza A viruses in humans which are able to transmit person to person and cause a global pandemic, is likely to have arisen from an initial rare event involving a novel influenza A virus with the capacity of replicating in humans and transmitting. Virus isolation studies from 1933 onwards, and analysis of isolates, have given an understanding of how the epidemic and pandemic behaviour relates to changes in the virus.
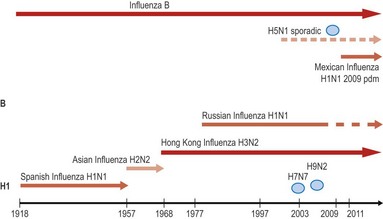
Fig. 49.5 Influenza viruses first isolated in 1933. Co-circulation of different viruses during the 20th and 21st centuries. Oval symbol denotes sporadic incidents and outbreaks with avian influenza viruses.
Major pandemics are associated with antigenic shifts – when the viral HA or NA (or both) is changed. Antigenic shift results from the acquisition of a complete new RNA segment 4 and/or 6, either as a result of reassortment or infection of humans with an animal virus. Until 1977, when H1N1 reappeared, it was considered that when a ‘new’ pandemic virus appeared the ‘old’ one disappeared, but since that time two influenza A subtypes have been circulating concurrently, namely H3N2 and H1N1 (Fig. 49.5). H1N1 viruses in 1977 were antigenically very similar to H1N1 viruses from before the 1957 pandemic, and may have been reintroduced from frozen laboratory sources, as there is no evidence of latent or persistent infection of humans. The most recent pandemic of 2009 exemplifies the emergence of an influenza strain from an animal reservoir (H1N1 from swine reservoir), to cause widespread disease in humans of all age groups. Although H1N1 has been circulating since 1918, H1N1 swine viruses were sufficiently different to H1N1 viruses circulating in humans, so that pre-existing immunity in the population was insufficient to prevent widespread transmission, especially in younger age groups. The impact of H1N1 in the population was mild to moderate overall, partly due to the lower intrinsic virulence of the strain, and partly due to the pre-existing immunity in the adult and elderly population, although susceptible groups such as pregnant women had an increased risk of serious outcome. At the time of writing, H1N1 2009pdm has almost completely displaced the previously circulating H1N1 strain.
Epidemics occurring regularly in winter months between pandemics are associated with genetic drift in the HA antigen. Amino acid changes arising as a result of genetic mutation provide a selection advantage for the virus if they occur on the globular head of the HA protein, where host antibody binds (Fig. 49.3). Mutation at these sites allows the virus to infect despite the presence of antibody to previous strains, a phenomenon known as antigenic drift.
The wild bird reservoir for influenza A is globally dispersed and mobile, creating a natural milieu for relentless evolution of genetic diversity. Consequently, there is a continuous threat to humans of the emergence of a new influenza A virus, capable of adapting to human transmission, causing a pandemic of disease in a globally susceptible population. The criteria for the establishment of a new pandemic of influenza in humans include:
Sporadic zoonotic infections of influenza A are therefore closely monitored, in view of the potential they have to create a new pandemic influenza strain. All of the HA and NA subtypes are found in aquatic birds (both seabirds and ducks), but in these animals the viruses vary little. Viral factors govern the ability to transmit and replicate in humans. These include receptor characteristics, replication competence and ability of virus to be shed directly into bodily secretions, which can be transmitted. In the laboratory it is easy to show that, if a cell is infected with two different strains, viruses arise with RNA segments derived from each parent through reassortment of segments.
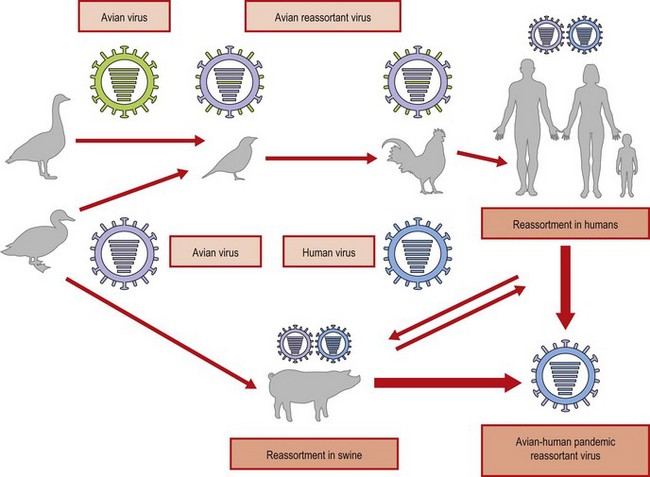
Pigs have receptors for both human and avian strains, and this mammalian host has long been considered a key ‘mixing vessel’ for reassortment following simultaneous infection with human and avian viruses. The conditions in South East Asia, where there are high densities of people, poultry and pigs, favour such an outcome (Fig. 49.6). The importance of the pig host in the generation of diversity is emphasized by the 2009 pandemic.
Cross species influenza A infection
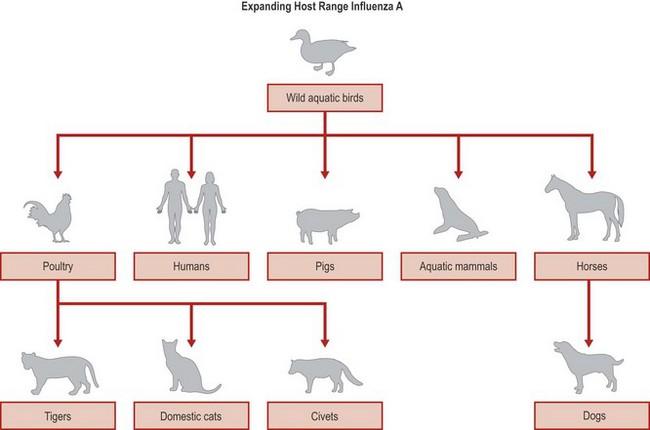
In recent years, influenza A viruses have been detected in a variety of mammalian species including dogs, mink, tigers, whales, cats and civets, usually arising from occasional transmission events from avians which may cause a range of disease symptoms in the new host, depending on the infecting virus subtype. Influenza A infection in mammalian species is usually associated with viral replication in the respiratory tract leading to respiratory illnesses with some systemic features, and occasionally extra respiratory replication. Sporadic infection events have usually involved only limited transmission in the new host. Diseased poultry and domestic ducks usually serve as the source of infection for avian influenza in mammalian hosts (Fig. 49.6). Infection arises through a variety of routes including contact with contaminated water, avian faecal material or contact with carcasses of infected birds. However, adaptation to dogs in North America has occurred, as a result of transmission of H7 from horses, creating a new animal reservoir of endemic influenza A (Fig. 49.7).
Zoonotic transmission to humans
Sporadic zoonotic infection of humans with influenza A viruses should be distinguished from epidemic human influenza A seasonal infection, which occurs during the winter months and is attributable to subtypes of influenza A (H3N2 and H1N1) which normally transmit between humans causing seasonal respiratory illness along with influenza B or C.
Human cases of infection with avian influenza viruses involving H5, H7, H10 or H9 subtypes have usually been acquired as a result of exposure to domestic poultry, material containing avian influenza virus or other mammalian species infected as an intermediate host with an avian influenza virus. The outcome of sporadic human infection ranges from severe disease of the respiratory system, which may also lead to multi-organ failure and death, to milder respiratory infection or simple conjunctivitis. The spectrum of disease is dependant on the nature of the infecting avian influenza A virus, e.g. H5 infections are associated with severe respiratory infections with a high case fatality rate; H7 infections are associated with conjunctivitis and milder respiratory infections. Risk factors for the acquisition of zoonotic avian or non human influenza A include:
• close or direct contact with diseased poultry or other domestic fowl
• close or direct contact with other mammalian species infected with unusual influenza A viruses (swine, seals)
• inhalation, ingestion or mucosal contact with infectious material.
Risk factors for severe human disease outcome following zoonotic infection of influenza A include:
Human seasonal influenza A, B and C infections
As can be seen from the foregoing discussion, influenza A virus subtypes circulating in humans (H1N1 and H3N2) were ultimately derived from an animal reservoir. Several influenza A subtypes co-circulate globally in humans in varying proportion, causing respiratory disease during the winter months in temperate climates, and during months of highest humidity in dry climates. In tropical climates, distinct seasonality is less evident, with patterns of influenza transmission throughout the year. Influenza B viruses co-circulate globally with influenza A viruses and contribute to the burden of seasonal influenza (Fig. 49.5), but are not usually associated with such severe morbidity and mortality. Influenza B viruses do not undergo antigenic shift as there is no animal reservoir and, although epidemics do occur at 3–6-year intervals, they never reach pandemic proportions. The antigenic changes in influenza B result from genetic drift, as seen in influenza A after the appearance of ‘new’ virus strains. Influenza C viruses are associated with much milder illness, with limited generation of genetic diversity and sporadic pattern of circulation.
Clinical features in human seasonal influenza A infection
Influenza is an acute infection of the human respiratory tract. Disease is characterized by the sudden onset of fever, chills, headache, muscle pain and extreme fatigue. Other common symptoms include a dry cough, sore throat and stuffy nose. For otherwise healthy individuals, influenza is an unpleasant but usually self-limiting disease with recovery usually within two to seven days. The illness may be complicated by (and may present as) bronchitis, secondary bacterial pneumonia or with otis media (ear infection) in children.
Serious illness and mortality are highest among newborns, older people and those with underlying disease, particularly chronic respiratory or cardiac disease, or those who are immunosuppressed.
In classical seasonal influenza A infection:
• the incubation period is short, 2 days, but may vary from 1–4 days
• the illness is characterized by a sudden onset of systemic symptoms such as chills, fever, headache, myalgia and anorexia
• respiratory symptoms are also common but take second place to the systemic effects, especially early in the illness.
Many patients have both upper and lower respiratory tract infection, often with a troublesome, dry cough. The main physical finding is pyrexia, which rises rapidly to a peak of 38–41°C within 12 h of onset. Fever usually lasts for 3 days, but may be present for 1–5 days. During the second and third days of the illness the systemic effects diminish, and by the fourth day, the respiratory symptoms and signs are predominant. In adults, systemic illness without respiratory symptoms is common. Some symptoms are age-specific, for example febrile convulsions and otitis media in children and dyspnoea in the elderly. About one-third of patients suffer only a common cold-like illness, and as many as 20–30% of cases are subclinical (asymptomatic). A long convalescence is common, and cough, lassitude and malaise may last for 1–2 weeks after the disappearance of other manifestations. Many other respiratory viruses can cause typical influenza-like illnesses, although the severity of the systemic symptoms is usually greatest with influenza virus.
Complications of human seasonal influenza
• Primary influenza pneumonia is an unusual complication that may occur at any age and carries a case fatality rate of 1–5%, depending on age and underlying conditions. This can be fatal, especially in young adults during an outbreak, after a very short illness of sometimes less than 1 day. A similar rapid illness can occur in the elderly.
• More commonly a bacterial pneumonia caused by Staphylococcus aureus, Streptococcus pneumoniae or Haemophilus influenzae occurs late in the course of the illness, often after a period of improvement, resulting in a classical biphasic fever pattern.
• Very rare complications include myocarditis, encephalitis or meningoencephalitis.
The incidence of chest complications is related to the age of the patient, increasing progressively after the age of 60 years. Severe infections and sudden death can occur, especially if there is some underlying disease, such as cerebrovascular, cardiovascular or chronic respiratory disease.
In the immunocompromised, symptoms may last longer and viral excretion may go on for weeks to months. Excess mortality was reported in pregnant women during the 1918 and 1957 pandemics, and even in non-pandemic outbreaks an increase in hospital admission due to cardiorespiratory disease is seen in the second and third trimesters. Susceptibility of pregnant women to severe influenza was also evident in the 2009 pandemic.
Clinical features in zoonotic influenza A infections
There is a spectrum of illness associated with the presentation of zoonotic avian influenza A virus infection, which is associated with the infecting subtype. Conjunctivitis was a feature of H7N7 outbreak cases in the Netherlands. H5N1 infections are associated with illness onset up to seven days post exposure, in contrast to the shorter incubation time of seasonal influenza, with associated delayed viral shedding. The early stages of infection may be difficult to distinguish from seasonal influenza or other respiratory infections which share many common presenting symptoms. The predominant clinical features include high temperature, cough and shortness of breath. Although the majority of cases have been respiratory in presentation, a handful of cases present atypically with fever, gastrointestinal disturbance and diarrhoea. Early clinical signs include alteration in liver function tests, particularly elevated aminotransferase enzymes, lymphopenia and evidence of interstitial pneumonia on chest X-ray. Progression of disease predominantly involves the respiratory tract and can lead to acute respiratory distress syndrome, with death most frequently being due to respiratory failure, often associated with multi organ failure. Limited autopsy evidence from fatal H5 cases is consistent with respiratory failure and primary viral interstitial pneumonia, but with little evidence of extra respiratory spread of virus to account for the multisystem dysfunction. The latter may be attributable to immune dysregulation leading to a ‘cytokine storm’ (uncontrolled release of tissue damaging immune response molecules); there is some autopsy evidence for haemophagocytic syndrome, but the overall understanding of pathogenesis is limited. Recovery from H5 infection occurs from about 7–10 days post illness onset, and is associated with a rise in neutralizing antibody titres.
Influenza C
Clinically, influenza C causes an afebrile upper respiratory tract infection usually confined to young children; outbreaks are not recognized.
Seasonal influenza A epidemics
An individual throughout life can expect to experience multiple influenza infections. Viruses bearing mutations accumulated during error prone replication (genetic drift) may have a replication advantage because they can evade the existing human immune response, giving a selection advantage due to antigenic drift in circulating strains. Over a period of several years, antigenic variants gradually predominate, displacing older strains, and cause epidemic disease.
Influenza virus is transmitted is by aerosol, droplets or direct contact with respiratory secretions of someone with the infection, for example sneezing and coughing. Influenza spreads rapidly, especially in closed communities. Most influenza cases in temperate countries tend to occur during a twelve to sixteen-week period during the winter. The timing, extent and severity of ‘seasonal’ influenza can all vary according to country. Winter activity is therefore unpredictable. Influenza A viruses cause outbreaks most years in most countries, and it is these viruses that are the usual cause of seasonal epidemics. More severe epidemics occur intermittently, often associated with the emergence of an antigenic drift variant. Influenza B tends to cause less severe disease and smaller outbreaks, although in children the severity of illness may be similar to that associated with influenza A.
High rates of infection are generally found in pre-school children; thereafter the rates are lower, even in the elderly. However, elderly patients in residential homes, if not protected, are at particular risk from acute illness and sudden death. Thus, even if the attack rate is low, the case fatality rate is high. Staff, visitors or other patients may introduce the virus, and this is an important consideration when planning intervention measures.
Pathogenesis
Inhaled virus is deposited on the mucous membrane lining the respiratory tract or directly into the alveoli, the anatomical location depending on the size of the droplets inhaled. The virus is exposed to mucoproteins containing sialic acid that can bind to the virus, thus blocking virus attachment to respiratory tract epithelial cells. However, the action of viral neuraminidase allows the virus to break any bonds formed. Specific local secretory immunoglobulin (Ig) A antibodies, if present from a previous infection, may neutralize the virus before attachment occurs, provided the antibody corresponds to the infecting virus type. If not prevented by one of these immune defence mechanisms, virus attaches to the surface of a respiratory epithelial cell and the intracellular replication cycle is initiated.
The major site of infection for seasonal influenza in humans is the ciliated columnar epithelial cell. New viruses bud from the apical membrane, the cilia are lost and viruses spread to other areas of the respiratory tract. The cell damage initiates an acute inflammatory response with oedema and the attraction of phagocytic cells. The earliest response is the synthesis and release of interferons from the infected cells: these can diffuse to and protect both adjacent and more distant cells before the virus arrives. It appears that interferons released in this way cause many of the systemic features of the ‘flu-like’ syndrome. Although viral components are absorbed and trigger the immune system, the virus itself is confined to the epithelium of the respiratory tract. Specific antibody helps to limit the extracellular spread of the virus, whereas T cell responses are directed against the viral glycoproteins on the surface of infected cells, leading to their destruction by cytotoxic T cells and also by antibody-dependent cell cytotoxicity. The pathogenesis of severe sporadic influenza A zoonotic infections in humans can involve multisystem failure, but evidence of extra respiratory virus replication is rare. It is considered likely that aberrant innate immune responses (cytokine storm arising from massive unregulated production of cytokines and chemokines) contribute to poor outcome in severe cases, possibly arising from different respiratory cell tropism.
The pathogenicity of influenza viruses is multifactorial and may involve viral, host and environmental factors. Pathogenicity is best understood with influenza A in birds. Host cell receptors are determinants of tropism in birds. Virus infectivity is dependent on host protease activity, cleaving HA into HA1 and HA2. There is a clear molecular correlate of pathogenicity associated with the presence of a string of polybasic amino acids at the HA1–HA2 cleavage site in certain H5 or H7 avian strains, as the HA protein containing such an insert can now be cleaved by a much wider range of host protease enzymes. In turn, virus dissemination within the avian host is enhanced, and replication occurs in a much wider range of tissues outside the gastrointestinal tract.
Understanding of pathogenesis in mammals is much less complete, but is recognized to be multifactorial involving several genes, including viral HA, NA, polymerase proteins and NS1. Receptor binding preference is an important determinant of tissue and cell tropism which may influence innate immune responses. Polymerase replication proteins may influence the ability of virus to replicate efficiently, and the NS1 non-structural protein may also influence pathogenicity by its role in antagonism of innate immune responses.
Immunity
After an attack of influenza, immunity to the particular strain of infecting virus is of long duration. It is related to the amount of local antibody (IgA) in the mucous secretions of the respiratory tract together with specific IgG serum antibody concentration. Immunity to infection, especially with influenza A, is subtype specific, giving little or no protection against subtypes possessing immunologically distinct H or N proteins. Once recovered from an initial influenza infection, exposure to more recent related strains will boost IgG levels to the earlier strains, the so-called original antigenic sin phenomenon.
Laboratory diagnosis
Prior to the 21st century, laboratory diagnosis of influenza depended on virus isolation through culture methods, or through detection of viral proteins in respiratory epithelial cells by immunofluoresence (IF) or enzyme linked immunosorbent assays (ELISA). The application of reverse transcription PCR to respiratory clinical material to detect viral genomic material has transformed the time taken to provide accurate and reliable diagnosis from days to within a matter of hours. Rapid diagnosis of respiratory infections has increased in importance, particularly in hospital or care facilities, such as homes for the elderly. Antiviral drugs given early in infection can be used to control disease and limit transmission. The best specimens for rapid diagnosis are nasal aspirates or nasal washes, but nasal or throat swabs containing epithelial cells are satisfactory if taken in the first few days of illness (Fig. 49.8).
In zoonotic influenza A infection (H5 or H7), virus shedding peaks several days after illness onset, and slightly later than in seasonal influenza, but may continue for up to 10 days, and declines in the recovery phase. In fatal cases, viral shedding may continue at very high levels. H5 virus replication occurs predominantly in the lower respiratory tract. Detection of virus genomic material by RT-PCR is optimal if secretions from the lower respiratory tract are obtained. Detection of infected airway epithelial cells using direct IF is an insensitive technique, and if undertaken, should be recognized as a suboptimal diagnostic test.
Whilst it is now possible to make a diagnosis of influenza without a virus isolate, it is important in the early stages of an outbreak or in sporadic cases that viruses should be isolated and analyzed antigenically to provide the best possible information for vaccine production. For primary isolation the most suitable cells are Madin–Darby canine kidney (MDCK) cells.
Serology
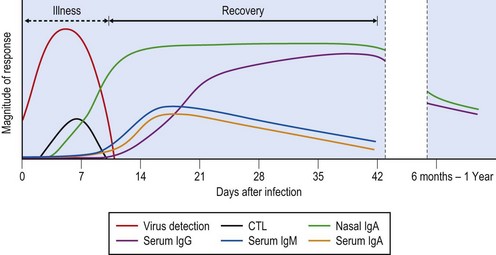
Serological confirmation of a clinical diagnosis is by demonstration of a four-fold or greater rise in functional strain specific antibody titre. Strain differences can be demonstrated by means of haemagglutination inhibition (HI) and neutralization antibody assays. Specific neutralizing antibody can be detected from about 10–14 days post infection, and reaches a plateau at around 28 days (Fig. 49.8). Complement fixation tests are still occasionally used. This test uses nucleocapsid antigens that are type-specific and can distinguish A from B and C infections, but cannot distinguish between different influenza A infections.
Treatment
Oral amantadine hydrochloride was introduced in the early 1980s, followed by a derivative, rimantadine. These drugs work by blocking the M2 ion channels in the envelope, thus preventing the pH changes that precede the membrane fusion step essential for nucleocapsid release. Unfortunately, these compounds have activity only against influenza virus type A. Amantadine is effective when given prophylactically, and also therapeutically in patients treated within 24 h of onset of illness. Viruses resistant to amantadine and rimantadine may appear within a few days of drug administration; however, resistant strains show no increased pathogenicity or transmissibility. In recent years, most circulating H3N2 and approximately 10% of H1N1 viruses have been naturally resistant to amantadine. Therefore, amantadine is not the drug of first choice, and it is essential to know which virus subtype is circulating prior to prescribing amantadine.
More recently two neuraminidase inhibitors (NIs), zanamivir and oseltamivir, have been licensed for therapeutic use in both influenza A and B infections, and other drugs with a similar mode of action are in development (e.g. peramivir). They act to prevent the release of viral particles through the action of viral neuraminidase enzymes which are conserved across all viral NA subtypes. They can reduce the duration of symptoms by 1–3 days if given within 36 h of the onset of illness. Zanamivir has poor bio-availability and is administered by inhalation of a dry powder twice daily for 5 days. Oseltamivir is given by mouth as a pro-drug and has excellent bio-availability, although nausea and vomiting may occur in some patients. Twice-daily dosage for 5–7 days has been used in those with normal renal function; once-daily dosage is recommended when renal function is impaired.
Both drugs have been licensed since 1999–2000 and a decade of global use indicates that mutations in viral NA are the major source of drug resistant variants. Resistant viruses have usually had a fitness deficit which compromises their transmissibility. Oseltamivir has been used much more extensively in this time, and as a consequence most available data on the emergence of antiviral resistance is related to oseltamivir use. Some key principles have become evident about the emergence of NI drug resistance.
Influenza A virus NAs are classified into group I and group II NAs, which differ in configuration of the enzyme active site. Group I NAs e.g. N1, have an active site cleft for substrate binding which is broader, so that the most common oseltamivir resistance mutation in N1 NA at position 275 is readily tolerated and inhibits drug action. Such mutations in H1N1 viruses arise during treatment, most commonly in younger or immunocompromised individuals where viral shedding is higher, and in H5N1 viruses where viral replication is not controlled by human immune responses. The spontaneous emergence and transmission of resistant H1N1 virus due to H275Y mutation during the northern hemisphere winter of 2007–2008 was a demonstration of the genetic evolution of influenza viruses. Compensatory mutations in the viral NA or other parts of the viral genome, arising from genetic drift have overcome the fitness deficit normally associated with mutation at position 275, enabling an otherwise disadvantaged virus to outcompete sensitive strains and predominate globally. Over a period of approximately 12 months, virtually all seasonal H1N1 viruses have become naturally oseltamivir resistant.
The patterns of mutations conferring drug resistance are subtype and drug specific. Although H275Y resistance is most common in N1 subtype containing viruses, such viruses remain susceptible to zanamivir (but not peramivir). N2 NAs fall into group 2 NA, for which the active site is narrower, and mutation at position 275 does not have the same effect. In H3N2, the most common mutation conferring NI resistance is at position 119 or position 272. As a consequence of the emergence of significant antiviral resistance, it is now important to ensure global surveillance of drug resistance and a rapid analysis of circulating subtypes so as to direct appropriate antiviral therapy.
During a pandemic there is a major role for the prophylactic use of anti-influenza drugs such as oseltamivir and zanamivir in those at high risk, in healthcare staff and in staff in long-term care facilities who have not been protected by an appropriate vaccine. Early studies have identified protection rates of over 80% with such chemoprophylaxis.
Control measures
The presence of a large global mobile animal reservoir of influenza A virus suggests that eradication of avian influenza A as a zoonotic infection of humans will be impossible. Control strategies focus on limiting the opportunities for cross species transmission of novel subtypes. For example:
• housing domestic poultry in shelters to avoid contact with over-flying migrating birds
• eliminating wild bird markets
• segregating different species of birds in markets
• housing aquatic birds and domestic poultry separately
• slaughtering domestic flocks infected with highly pathogenic influenza A viruses.
These measures may achieve some success in preventing zoonotic transmission of influenza A to humans, and have certainly reduced the circulation of H5 in domestic poultry in Hong Kong since 1997. However, they have little impact on the annual cycle of avian influenza in the wild bird population. Transmission of avian influenza by migrating birds is responsible for extending the geographical range of evolving avian influenza strains, as has been seen in 2005 and 2006 with the detection of H5N1 in birds and poultry in parts of Europe and Africa and the Middle East, whereas hitherto it was confined to South East Asia.
Infection control
Exposure to heat for 30 min at 56°C is sufficient to inactivate most strains. The viruses are inactivated by a variety of substances, detergents, soaps, and other household compounds as well as 20% ethanol, halogens and phenolic compounds. These biological properties are the basis of most infection control advice and practices surrounding the management of noscomial influenza infection and exposure of individuals to influenza viruses.
Immunization
A key control strategy for reduction of morbidity and mortality due to influenza is immunization. The aim is to produce haemagglutination inhibiting or neutralizing antibody in all vaccinees. This protects against infection, but only with strains closely related to those in the vaccine, and limits transmission. Whole virus, split and subunit inactivated influenza vaccines for intramuscular injection have been widely available for many years (Fig. 49.9), and live attenuated vaccines have been available more recently for use in children in the USA. Inactivated vaccines are trivalent and contain the H and N subunits from two type A strains and one type B strain. The strains are updated annually on the recommendation of the WHO and, in the UK, are recommended for use in people aged over 65 years, and in those of any age who suffer from chronic cardiorespiratory problems, diabetes, renal or liver failure or an immunosuppressive illness. In interpandemic years, with the elderly given annual boosting, the mortality rate can be reduced by 75% and the rate of hospital admission with complications by about 50%.
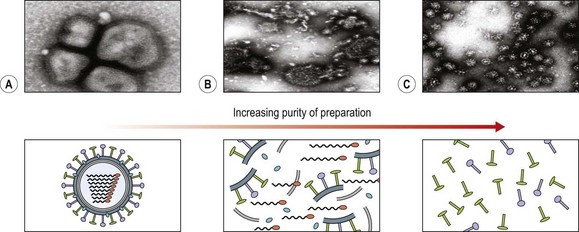
Fig. 49.9 Inactivated vaccine preparations. (A) Whole virus. (B) Split vaccine preparation, with some internal components. (C) Subunit vaccine, highly purified surface proteins.
Intervention during a pandemic is more difficult, often for logistic and vaccine supply reasons. Vaccine delivery may be delayed, and the risk groups for vaccination may differ from those normally vaccinated during seasonal influenza, as was noted during the pandemic of 2009, where the key risk groups were children and pregnant women. The interval from isolation of a potential pandemic strain to release of an inactivated vaccine made by current methods using eggs is 4–6 months. Such vaccines rely on adequate quantities of virus with the appropriate H and N antigens being produced in cells or eggs inoculated with seed virus. Reassortment of two strains, one a high-yielding laboratory-adapted strain and the other containing the required H and N antigens, may be designed for growth in eggs so that vaccine is prepared as quickly as possible. More recently using the newer process of reverse genetics, vaccine seed strains can be generated without using traditional reassortment methods. Appropriate vaccine strains are developed by cloning of relevant genes and this is a successful innovation to speed up vaccine production, which is gradually being introduced in the vaccine industry.
Vaccine seed strains need to be grown in cell or egg substrates to make inactivated vaccines. Separated whole virus particles are inactivated by either formalin or β-propiolactone, and may be used at this stage as ‘whole virus vaccines’. Whole virus vaccine should not be given to those who are allergic to egg protein. The H and N antigens may be separated from the whole virus by treatment with detergent, and such subunit or split-virus vaccines are better tolerated, especially by young children. Other types of vaccines have been tried, such as cold-adapted live-attenuated vaccines given intranasally. These have been generally effective in provoking a good local (IgA) antibody response and are particularly good at developing protective efficacy in children, though less so in adults, but are not widely used at present.
Clinical trials as part of pandemic vaccine development programmes have demonstrated an important role for vaccine adjuvants such as MF59 and ASO3 in antigen sparing, enabling reduced quantities of vaccine to go further and providing excellent immune responses, particularly in young children. Such vaccines were used for the first time as part of the response to the 2009 pandemic. Future vaccine development will concentrate on ensuring better understanding of human immune responses, improving immunogenicity of existing vaccines and developing processes and substrates for a more rapid response.
Global surveillance
The WHO Global Influenza Programme with its network of reference laboratories plays a very important role in monitoring the evolution of influenza viruses, selecting and developing prototype pandemic vaccine strains, and developing and updating WHO diagnostic reagents. Recent changes in International Health Regulations have increased the obligation in countries worldwide to have the capacity for preventive measures and to be able to detect and respond to infections of international concern. The emergence of the 2009 pandemic demonstrated the unpredictability of influenza virus evolution and the continuing threat to human health posed by this viral infection. Responses to the pandemic were the most sophisticated ever known following the emergence of a new virus, which despite its mildness still presented formidable challenges to health care systems in developed and developing world countries. As a result of the global experiences during the first waves of the pandemic, incremental improvements in surveillance and vaccine production may be expected. An enhanced search for antiviral combination therapies and new antiviral targets to provide better counter measures for future epidemic and pandemic influenza A strains, which will inevitably emerge, are required.
Bright RA, Medina MJ, Xu X, et al. Incidence of amantadine resistance among influenza A (H3N2) viruses isolated worldwide from 1994–2005: a cause for concern. The Lancet. 2005;366:1175–1181.
Capua I, Alexander DJ. The challenge of avian influenza to the veterinary community. Avian pathology. 2006;35:189–205.
de Jong M, Thanh TT, Khank TH, et al. Oseltamivir resistance during treatment of influenza H5N1. The New England Journal of Medicine. 2005;353:2667–2672.
Fiore AE, Bridges CB, Cox NJ. Seasonal influenza vaccines. Current Topics in Microbiology and Immunology. 2009;333:43–82.
Garten RJ, Davis CT, Russell CA, et al. Antigenic and genetic characteristics of swine-origin 2009 A(H1N1) influenza viruses circulating in humans. Science. 2009;325:197–201.
Hardelid P, Andrews NJ, Hoschler K, et al. Assessment of baseline age-specific antibody prevalence and incidence of infection to novel influenza AH1N1 2009. Health Technology Assessment. 2010;14:115–192.
Koopmans M, Wilbrink B, Conyn M, et al. Transmission of H7N7 avian influenza A virus to human beings during a large outbreak in commercial poultry farms in the Netherlands. The Lancet. 2004;363:587–593.
Russell CA, Jones TC, Barr IG, et al. The global circulation of seasonal influenza A (H3N2) viruses. Science. 2008;320:340–346.
Stephenson I, Nicholson KG, Wood JD, et al. Confronting the avian influenza threat: vaccine development for a potential pandemic. The Lancet Infectious Diseases. 2004;4:499–509.
Wright PF, Neumann G, Kawaoka Y. Orthomyxoviruses. In: Fields BN, Knipe DM, Howley PM. Virology. ed 5. Philadelphia: Lippincott-Raven; 2007:1691–1740.
Writing Committee of the World Health Organisation (WHO) Consultation of Human Influenza. Avian influenza A (H5N1) infection in humans. The New England Journal of Medicine. 2005;353:1374–1385.
Writing Committee of the World Health Organisation (WHO) Consultation on Clinical Aspects of Pandemic (H1N1). Influenza 2010 Clinical Aspects of Pandemic 2009 Influenza A (H1N1) virus infection. The New England Journal of Medicine. 2009;362:1708–1719.
Zambon M, Potter CW. Influenza. Zuckerman AJ, Banatvala JE, Schoub BD, et al. Principles and Practice of Clinical Virology, ed 6, Chichester: Wiley, 2009.
World Health Organization. Influenza. http://www.who.int/csr/disease/influenza/en/.
Health Protection Agency (UK). Influenza. http://www.hpa.org.uk/Topics/InfectiousDiseases/InfectionsAZ/Influenza/.
Neuraminidase Inhibitor Susceptibility Network. http://www.nisn.org/l_influenza_resources.html.
International Society for Influenza and Respiratory Viruses. http://www.isirv.org/.