Chapter 51 Genetic Disorders
INHERITED DISEASES
Inherited diseases and congenital defects are structural or functional abnormalities that may or may not be obvious at birth. There are a wide range of possible defects, from a single structural change (e.g., leg length) to the involvement of multiple organs or systems (e.g., storage diseases, chromosomal abnormalities), depending on the type and extent of toxic exposure during gestation or genetic mutation. It is important to identify animals with genetic or congenital disorders because of the economic and emotional impact on clients, but many disorders can be difficult to recognize and trace, particularly if they lead to embryonic or fetal death, abortion, dysmaturity, premature birth, or full-term stillbirth.
It is important to remember that not all congenital defects are genetic in origin and making the distinction between conditions caused by an environmental factor and a genetic cause will help clients make appropriate changes to prevent the disorder in the future. To determine the cause of a disorder, it is vital that each case be thoroughly examined. Each case should be subjected to a full, careful description of the disorder, including a complete necropsy. The environment and management conditions should be investigated for potential causative agents, and all available genetic information for the affected animal(s) and unaffected herdmates should be collected, including gender, birth date, and breed. Environmental teratogens affecting large animals include toxic plants, drugs, viruses, and physical agents (e.g., hyperthermia, irradiation). Although it may be difficult to identify inciting teratogens, analysis of affected herds will reveal patterns that follow seasonal or management changes or stressful events.
This chapter focuses on the essential genetic information that veterinarians require to inform their clients about genetic diseases and how to find information on diseases and traits with a genetic component.
GENETIC INFORMATION
The basic blueprint for life in most organisms, the genetic material, is deoxyribonucleic acid (DNA). DNA is composed of two strands of bases: adenine (A), thymine (T), guanine (G), and cytosine (C). These bases align to form complementary base pairings such that A always pairs with T and G always pairs with C. These strands can be completely separated and replicated in preparation for cellular division. Alternatively, these strands can be partially separated, transcribed into ribonucleic acid (RNA), and translated into functional proteins.
The DNA sequence of a mammalian genome is divided into autosomes and a pair of sex chromosomes, X and Y. Each chromosome contains a variety of genes as well as “filler” DNA that does not code for proteins, but may determine where and to what degree each gene is expressed. Genes on each chromosome code for the various structural and enzymatic proteins required for life. One set of chromosomes, and thus one set of genes, is inherited from each parent. As a result, a mammal generally has two copies, or “alleles,” of any given gene. Simple traits or diseases involving these genes can be inherited in dominant or recessive patterns and involve the autosomes, or sex chromosomes. Unfortunately, not all traits or inherited diseases are simple; many involve multiple genes and interactions between genes and the environment, which can make it difficult to diagnose and understand a complex disease or trait.
Recessive Inheritance
Traits or diseases with a recessive mode of inheritance require two affected copies, or alleles, of the gene to express that trait or disease. These conditions can be autosomal, involving genes on the autosomes, or sex-linked, caused by genes on either the X or Y chromosome. Generally, autosomal recessive traits or diseases are caused by mutations that render a protein unable to perform its original function. When an animal inherits the affected allele from both parents, it is homozygous for the gene and is unable to make functional copies of that specific protein. Heterozygous individuals inherit only one copy of the affected allele, and they have a “backup” copy of the normal or wild-type allele to provide the functional protein. These heterozygous individuals are carriers of the trait or disease allele and can produce affected offspring if bred to another heterozygous or homozygous affected individual.
The hallmarks of autosomal recessive traits include that all offspring of two affected parents are affected and approximately equal numbers of males and females are affected. Unaffected carrier parents can produce affected offspring, and thus the trait or disease can skip generations. Additionally, mating an affected individual to a noncarrier (homozygous normal) produces all offspring that appear normal but carry the affected allele. Breeding two carrier individuals results in approximately 25% normal, 50% carrier, and 25% affected offspring.
An example of an autosomal recessive trait is spider lamb syndrome, or hereditary chondrodysplasia. Lambs affected with this syndrome display several skeletal abnormalities, including disproportionately long “spider” legs, curvature of the spine, facial deformities, rib and sternum deformities, lack of body fat, and muscular atrophy. The causative mutation is a single base-pair substitution, from a T to an A, altering a highly conserved amino acid in the fibroblast growth factor receptor 3 (FGFR-3) gene, which prevents the receptor from functioning to limit endochondral ossification and thus bone growth.1
Dominant Inheritance
Traits or diseases with a dominant mode of inheritance only require one affected allele to express the trait or disease, also known as the affected phenotype, and there is no distinguishable difference in the affected phenotype of a heterozygote and homozygote. Again, the genes that are responsible for these traits or diseases can be on the autosomes or sex chromosomes. Dominant traits or diseases may be caused by mutations that enable the mutant protein to have an altered function or structure. Alternatively, a single, functional copy of a gene may not be enough to achieve normal levels of function (“haploinsufficient”), resulting in the need for two normal copies of the gene to achieve a normal phenotype.
The main feature of an autosomal dominant trait or disorder is that affected individuals must have at least one affected parent (unless it is a new mutation), and thus the disorder does not skip generations. There is no gender bias with an autosomal dominant trait, and approximately half the offspring of a heterozygous affected individual will be affected.
An example of an autosomal dominant disease is myotonia in goats. When startled or making sudden, forceful movements, these goats can develop severe, acute muscle stiffness causing immobility and sometimes falling over, resulting in descriptions such as “fainting,” “nervous,” “stiff-legged,” or “epileptic” goats. A single nucleotide change was identified that substituted a proline for a conserved alanine residue in a chloride channel in the muscle fibers. This alteration in the chloride channel causes a diminished channel-open probability at voltages near the resting membrane potential of skeletal muscle, resulting in decreased chloride conductance and a significantly decreased electrical threshold for firing action potentials. Ultimately, this altered chloride channel allows conduction of repetitive impulses that result in sustained muscle fiber contraction and stiffness.2
Co-dominant Inheritance
The co-dominant (or semidominant) pattern of inheritance is distinguished by the fact that homozygotes can be differentiated from heterozygotes based on clinical features. A well-known example of a co-dominant disease is hyperkalemic periodic paralysis (HYPP) in quarter horses. In HYPP the voltage-gated sodium channels in the muscle fibers have a mutation that increases sodium permeability across the skeletal muscle cell membrane, resulting in increased muscle mass, but also drooling, prolapse of the nictitating membrane (“third eyelid”), respiratory stridor, and weakness. Homozygous HYPP horses experience more frequent and severe clinical signs of disease than heterozygous horses.3 Thus, it is important to counsel breeders to avoid producing homozygous affected horses by not mating two heterozygous horses, because 25% of their offspring would be expected to be homozygous affected, and 50% of the offspring would be heterozygotes. Breeding a heterozygous horse to an unaffected horse will also result in approximately 50% of the offspring being heterozygous and no possibility of producing a homozygous affected foal.
Sex-Linked Traits
Sex-linked traits and diseases involve genes located on either the X or the Y chromosome. Males are particularly susceptible to these conditions because they have only one each of the X and Y chromosomes. Thus, if they have a mutant allele for one of the genes on either of these chromosomes, they will express the affected phenotype because they do not have a “backup” copy. They have only one copy of the X and Y chromosomes, so they are hemizygous. Females generally are less likely to demonstrate sex-linked recessive traits or diseases because they have two copies of the X chromosome. However, because of random X inactivation in each cell during fetal development, some females may express an affected phenotype. To achieve this, the X inactivation must be skewed such that the unaffected chromosome is inactivated more often than the X chromosome carrying the affected allele. In rare cases in which the inactivation is significantly skewed, females may be mildly to severely affected, depending on the level of expression for the affected chromosome.
Key characteristics of X-linked recessive traits are that the trait appears with much greater frequency in males than females, half the sons of carrier females will be affected, and half the daughters of carrier females will also be carriers. Hemophilia A, characterized by a strong tendency to bleed resulting from mutations in the clotting factor VIII gene on the X chromosome, is an example of an X-linked recessive disorder.
X-linked dominant traits are characterized by the following: affected offspring must have at least one affected parent, the disorder does not skip generations, and an affected male mated to normal females will transmit the mutation to all his daughters but not to his sons.
There are very few Y-linked diseases recognized in any mammalian species because of the small number of genes on the Y chromosome. Additionally, the majority of the genes contained on the Y chromosome are involved in male fertility; thus mutations in these genes generally render the animal sterile, resulting in no transmission of the mutations to future generations.
Polygenic Traits
In addition to the single-gene traits and diseases, numerous conditions are the result of two or more genes and may also involve genetic and environmental interactions. Some of these polygenic traits are economically important, such as milk quality and yield in dairy cows. For milk quality, a large number of genes may be contributing a small portion to the overall milk production of an animal.4 Teasing out the details of a complex polygenic system is difficult; however, research is ongoing to identify the exact genes involved and the role each gene plays for many economically important traits and diseases.
Penetrance and Expressivity
For many simple traits, there is an obvious difference between the two phenotypes that can be translated to the alleles involved, or genotype, with 100% certainty. A good example of this is albinism, which is considered to be 100% penetrant, with penetrance defined as the percentage of individuals with a given genotype who actually show the phenotype associated with that genotype. Influences such as modifying genes or environmental interactions can alter the expression of certain genes such that the exact genotype is not expressed in the outward phenotype. Alternatively, the function of a gene may be subtle, making it difficult to measure distinctions adequately between genotypes. Genotypes that may not be expressed in every individual are considered to be “incompletely penetrant.”
Another measure of genetic expression is the concept of expressivity. Modifier genes and environmental influences can affect the degree to which a genotype is expressed. For example, “variable expressivity” may be involved in pigment intensity when two individuals have the same genotype at the gene responsible for red pigmentation, but one has clearly darker red hairs than the other.
Incomplete penetrance and variable expressivity can greatly complicate analysis of genetic traits and diseases. These factors can also make breeding decisions more difficult because it may not be possible to classify an animal’s genotype based solely on a phenotype. In these more ambiguous cases, genetic testing will play a vital role in the unequivocal determination of genotypes, allowing appropriate and informed breeding decisions.
POSITIVE AND NEGATIVE SELECTION
Breeders can improve their breeding stock by selecting for or against specific traits and diseases. These processes are known as positive and negative selection, respectively. A breeder may choose to breed an animal because it has desirable genes or qualities (positive selection). Alternatively, they may choose not to breed an animal that has been shown, either through genetic or breeding tests, to carry an undesirable trait or disease (negative selection). Ultimately, both scenarios will achieve the goal of improved breeding stock.
CHROMOSOMAL ABNORMALITIES
Occasionally, problems arise during mitosis, meiosis, or fertilization, resulting in chromosomal, or karyotype, abnormalities. Many karyotypic abnormalities have been identified in all large animal species, but not every chromosomal abnormality has been associated with an overt disease phenotype.5,6 Some examples of common chromosomal abnormalities include the absence of an X chromosome, called XO Turner’s syndrome, which results in female infertility. An additional X chromosome in XXY Klinefelter’s syndrome causes underdeveloped males with underdeveloped male sexual behavior. In addition to abnormal numbers of sex chromosomes, two chromosomes can unite into a single chromosome, called a “Robertsonian translocation,” leading to reduced reproductive capacity in some species and early embryonic death in cattle. Because of possible chromosomal abnormalities, several countries have instituted mandatory kayotyping of some breeding cattle. Additionally, clinicians should consider karyotyping any animal that presents for reduced fertility or has multiple abnormalities.
BREEDING SCHEMES: TEST MATINGS
As a result of numerous advances in the field of genetics, the genetic causes of many diseases and traits are known and can be tested before using an animal in a breeding program. Additionally, more DNA-based tests are expected as research continues. However, clients may be interested in testing for a disease or trait that does not have a known genetic cause. For these situations, sires can be tested using breeding trials, as described in Table 51-1. For example, if a sire is bred to carrier females for a disease of concern, he would have to produce 10 normal offspring to ascertain that he is not a carrier with 95% confidence, 16 normal offspring to have 99% confidence, and 24 normal offspring to have 99.9% confidence that he is not a carrier for the disease of concern.
Table 51-1 Test Mating Schemes to Examine Males for Genetic Traits or Diseases at Various Confidence Levels
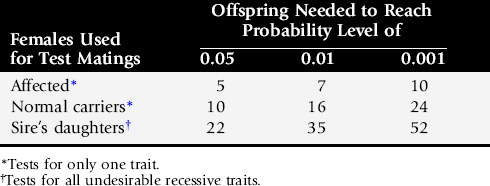
The time required to complete test matings can be significantly shortened using superovulated affected females, insemination, and embryo transfer (two per recipient) followed by early cesarean section at 60 days and examination of the offspring for the genetic defect(s) of concern.
OBTAINING GENETIC INFORMATION
Because of the ever-increasing wealth of genetic information, the most complete and current information can be found using online resources. It is recommended that clinicians use websites produced by reputable institutions to obtain the most accurate and referenced information available (Box 51-1).
Alternatively, clinicians are encouraged to contact the Veterinary Genetics services available at many veterinary schools if they are unable to locate the information they require.
RECOMMENDATIONS FOR BREEDING PROGRAMS
When a genetic disease has been identified, it is recommended that parentage be verified by DNA analysis and the extended pedigree be confirmed. Additionally, a pathologist should make appropriate examinations of tissues for an accurate diagnosis, and a certified statement detailing the condition should be written by a veterinarian or other third-party witness. The final decision with regard to the status of a particular animal should be withheld until all reasonable doubt has been removed; for bulls, this is usually at least two thoroughly documented cases or a positive genetic test, if available. The artificial insemination (AI) organization and breed registry should be informed of the status of each animal because disease information is important in future breeding decisions. Many AI organizations list any undesirable recessive alleles in their advertising, and some also remove carrier animals from service.
Depending on the value of the animal in question, a breeder may choose to remove the animal from the breeding program at the first sign of a potential genetic problem. In these cases, documentation of the abnormal conditions should still be conducted for completeness and for comparison with other cases that may be found in the future.
1 Beever JE, Smit MA, Meyers SN, et al. A single-base change in the tyrosine kinase II domain of ovine FGFR3 causes hereditary chondrodysplasia in sheep. Anim Genet. 2006;37:66.
2 Beck CL, Fahlke C, George AL. Molecular basis for decreased muscle chloride conductance in the myotonic goat. Proc Natl Acad Sci USA. 1996;93:11248.
3 Naylor JM, Nickel DD, Trimino G, et al. Hyperkalaemic periodic paralysis in homozygous and heterozygous horses: a co-dominant genetic condition. Equine Vet J. 1999;31:153.
4 Khatkar MS, Thomson PC, Tammen I, et al. Quantitative trait loci mapping in dairy cattle: review and meta-analysis. Genet Sel Evol. 2004;36:163.
5 McFeely RA. Chromosome abnormalities. Vet Clin North Am Food Anim Pract. 1993;9:11.
6 Nicholas FW. Introduction to veterinary genetics, ed 2, Blackwell: Oxford, England; 2003:93.