6 Serologic Diagnosis
Immunologic techniques are used to detect, identify, and quantitate antigen in clinical samples, as well as to evaluate the antibody response to infection and a person’s history of exposure to infectious agents. The specificity of the antibody-antigen interaction and the sensitivity of many of the immunologic techniques make them powerful laboratory tools (Table 6-1). In most cases, the same technique can be adapted to evaluate antigen and antibody. Because many serologic assays are designed to give a positive or negative result, quantitation of the antibody strength is obtained as a titer. The titer of an antibody is defined as the lowest dilution of the sample that retains a detectable activity.
Table 6-1 Selected Immunologic Techniques
| Technique | Purpose | Clinical Examples |
|---|---|---|
| Ouchterlony immuno–double-diffusion | Detect and compare antigen and antibody | Fungal antigen and antibody |
| Immunofluorescence | Detection and localization of antigen | Viral antigen in biopsy (e.g., rabies, herpes simplex virus) |
| Enzyme immunoassay (EIA) | Same as immunofluorescence | Same as immunofluorescence |
| Immunofluorescence flow cytometry | Population analysis of antigen-positive cells | Immunophenotyping |
| ELISA | Quantitation of antigen or antibody | Viral antigen (rotavirus); viral antibody (anti-HIV) |
| Western blot | Detection of antigen-specific antibody | Confirmation of anti-HIV seropositivity |
| Radioimmunoassay (RIA) | Same as ELISA | Same as for ELISA |
| Complement fixation | Quantitate specific antibody titer | Fungal, viral antibody |
| Hemagglutination inhibition | Antiviral antibody titer; serotype of virus strain | Seroconversion to current influenza strain; identification of influenza |
| Latex agglutination | Quantitation and detection of antigen and antibody | Rheumatoid factor; fungal antigens; streptococcal antigens |
ELISA, Enzyme-linked immunosorbent assay; HIV, human immunodeficiency virus.
Antibodies
Antibodies can be used as sensitive and specific tools to detect, identify, and quantitate the antigens from a virus, bacterium, fungus, or parasite. Specific antibodies may be obtained from convalescent patients (e.g., antiviral antibodies) or prepared in animals. These antibodies are polyclonal; that is, they are heterogeneous antibody preparations that can recognize many epitopes on a single antigen. Monoclonal antibodies recognize individual epitopes on an antigen. Monoclonal antibodies for many antigens are commercially available, especially for lymphocyte cell surface antigens.
The development of monoclonal antibody technology revolutionized the science of immunology. For example, because of the specificity of these antibodies, lymphocyte subsets (e.g., CD4 and CD8 T cells) and lymphocyte cell surface antigens were identified. Monoclonal antibodies are the products of hybrid cells generated by the fusion and cloning of a spleen cell from an immunized mouse and a myeloma cell, which produces a hybridoma. The myeloma provides immortalization to the antibody-producing B cells of the spleen. Each hybridoma clone is a factory for one antibody molecule, yielding a monoclonal antibody that recognizes only one epitope. Monoclonal antibodies can also be prepared and manipulated through genetic engineering and “humanized” for therapeutic usage.
The advantages of monoclonal antibodies are (1) that their specificity can be confined to a single epitope on an antigen and (2) that they can be prepared in “industrial-sized” tissue culture preparations. A major disadvantage of monoclonal antibodies is that they are often too specific, such that a monoclonal antibody specific for one epitope on a viral antigen of one strain may not be able to detect different strains of the same virus.
Methods of Detection
Antibody-antigen complexes can be detected directly, by precipitation techniques, or by labeling the antibody with a radioactive, fluorescent, or enzyme probe, or they can be detected indirectly through measurement of an antibody-directed reaction, such as complement fixation.
Precipitation and Immunodiffusion Techniques
Specific antigen-antibody complexes and cross-reactivity can be distinguished by immunoprecipitation techniques. Within a limited concentration range for both antigen and antibody, termed the equivalence zone, the antibody cross-links the antigen into a complex that is too large to stay in solution and therefore precipitates. This technique is based on the multivalent nature of antibody molecules (e.g., immunoglobulin [Ig] G has two antigen-binding domains). The antigen-antibody complexes are soluble at concentration ratios of antigen to antibody that are above and below the equivalence concentration.
Various immunodiffusion techniques make use of the equivalence concept to determine the identity of an antigen or the presence of antibody. Single radial immunodiffusion can be used to detect and quantify an antigen. In this technique, antigen is placed into a well and allowed to diffuse into antibody-containing agar. The higher the concentration of antigen, the farther it diffuses before it reaches equivalence with the antibody in the agar and precipitates as a ring around the well.
The Ouchterlony immuno–double-diffusion technique is used to determine the relatedness of different antigens, as shown in Figure 6-1. In this technique, solutions of antibody and antigen are placed in separate wells cut into agar, and the antigen and antibody are allowed to diffuse toward each other to establish concentration gradients of each substance. A visible precipitin line occurs where the concentrations of antigen and antibody reach equivalence. On the basis of the pattern of the precipitin lines, this technique can also be used to determine whether samples are identical, share some but not all epitopes (partial identity), or are distinct. This technique is used to detect antibody to fungal antigens (e.g., Histoplasma species, Blastomyces species, and coccidioidomycoses).
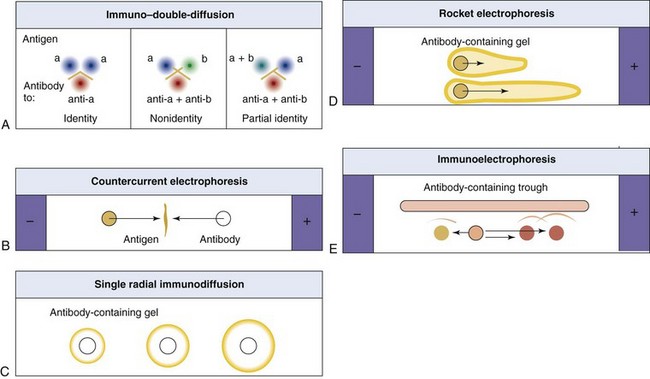
Figure 6-1 Analysis of antigens and antibodies by immunoprecipitation. The precipitation of protein occurs at the equivalence point, at which multivalent antibody forms large complexes with antigen. A, Ouchterlony immuno–double-diffusion. Antigen and antibody diffuse from wells, meet, and form a precipitin line. If identical antigens are placed in adjacent wells, the concentration of antigen between them is doubled, and precipitation does not occur in this region. If different antigens are used, two different precipitin lines are produced. If one sample shares antigen but is not identical, then a single spur results for the complete antigen. B, Countercurrent electrophoresis. This technique is similar to the Ouchterlony method, but antigen movement is facilitated by electrophoresis. C, Single radial immunodiffusion. This technique involves the diffusion of antigen into an antibody-containing gel. Precipitin rings indicate an immune reaction, and the area of the ring is proportional to the concentration of antigen. D, Rocket electrophoresis. Antigens are separated by electrophoresis into an agar gel that contains antibody. The length of the “rocket” indicates concentration of antigen. E, Immunoelectrophoresis. Antigen is placed in a well and separated by electrophoresis. Antibody is then placed in the trough, and precipitin lines form as antigen and antibody diffuse toward each other.
In other immunodiffusion techniques, the antigen may be separated by electrophoresis in agar and then reacted with antibody (immunoelectrophoresis); it may be pushed into agar that contains antibody by means of electrophoresis (rocket electrophoresis), or antigen and antibody may be placed in separate wells and allowed to move electrophoretically toward each other (countercurrent immunoelectrophoresis).
Immunoassays for Cell-Associated Antigen (Immunohistology)
Antigens on the cell surface or within the cell can be detected by immunofluorescence and enzyme immunoassay (EIA). In direct immunofluorescence, a fluorescent molecule is covalently attached to the antibody (e.g., fluorescein-isothiocyanate (FITC)–labeled rabbit antiviral antibody). In indirect immunofluorescence, a second fluorescent antibody specific for the primary antibody (e.g., FITC–labeled goat anti–rabbit antibody) is used to detect the primary antiviral antibody and locate the antigen (Figures 6-2 and 6-3). In EIA, an enzyme such as horseradish peroxidase or alkaline phosphatase is conjugated to the antibody and converts a substrate into a chromophore to mark the antigen. Alternatively, an antibody modified by the attachment of a biotin (the vitamin) molecule can be localized by the very high affinity binding of avidin or streptavidin molecules. A fluorescent molecule or an enzyme attached to the avidin and streptavidin allows detection. These techniques are useful for the analysis of tissue biopsy specimens, blood cells, and tissue culture cells.
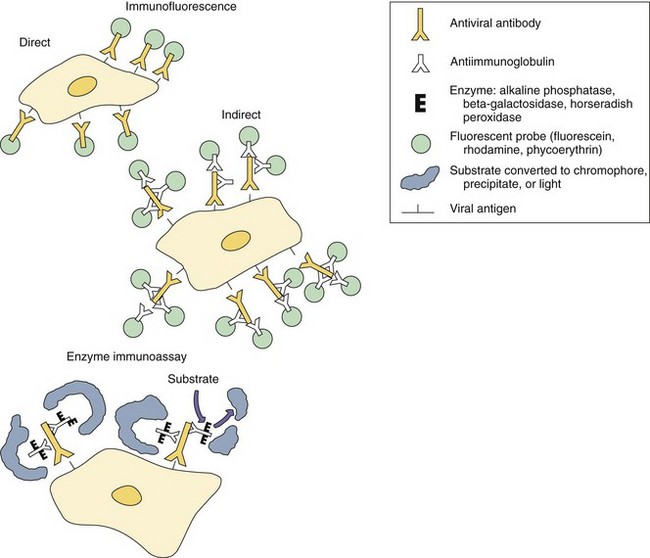
Figure 6-2 Immunofluorescence and enzyme immunoassays for antigen localization in cells. Antigen can be detected by direct assay with antiviral antibody modified covalently with a fluorescent or enzyme probe, or by indirect assay using antiviral antibody and chemically modified antiimmunoglobulin. The enzyme converts substrate to a precipitate, chromophore, or light.
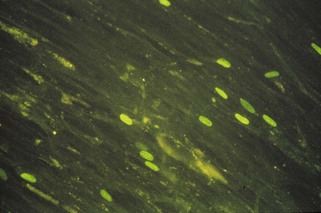
Figure 6-3 Immunofluorescence localization of herpes simplex virus–infected nerve cells in a brain section from a patient with herpes encephalitis.
(From Emond RT, Rowland HAK: A color atlas of infectious diseases, ed 2, London, 1987, Wolfe.)
The flow cytometer can be used to analyze the immunofluorescence of cells in suspension and is especially useful for identifying and quantitating lymphocytes (immunophenotyping). A laser is used in the flow cytometer to excite the fluorescent antibody attached to the cell and to determine the size of the cell by means of light-scattering measurements. The cells flow past the laser at rates of more than 5000 cells per second, and analysis is performed electronically. The fluorescence-activated cell sorter (FACS) is a flow cytometer that can also isolate specific subpopulations of cells for tissue culture growth on the basis of their size and immunofluorescence.
The data obtained from a flow cytometer are usually presented in the form of a histogram, with the fluorescence intensity on the x-axis and the number of cells on the y-axis, or in the form of a dot plot, in which more than one parameter is compared for each cell. The flow cytometer can perform a differential analysis of white blood cells and compare CD4 and CD8 T-cell populations simultaneously (Figure 6-4). Flow cytometry is also useful for analyzing cell growth after the fluorescent labeling of deoxyribonucleic acid (DNA) and other fluorescent applications.
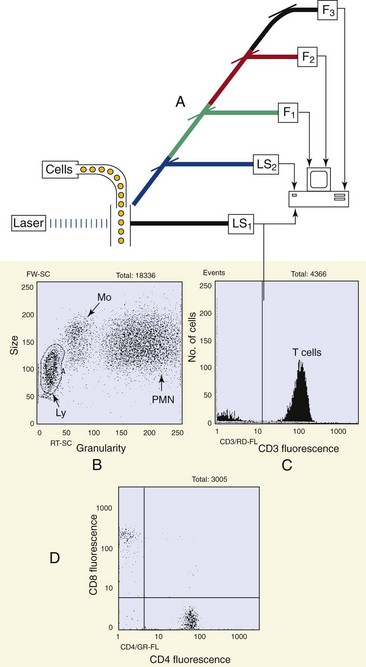
Figure 6-4 Flow cytometry. A, The flow cytometer evaluates individual cell parameters as the cells flow past a laser beam at rates of more than 5000 per second. Cell size and granularity are determined by light scattering (LS), and antigen expression is evaluated by immunofluorescence (F), using antibodies labeled with different fluorescent probes. Graphs B to D depict T-cell analysis of a normal patient. B, Light-scatter analysis was used to define the lymphocytes (Ly), monocytes (Mo), and polymorphonuclear (neutrophil) leukocytes (PMN). C, The lymphocytes were analyzed for CD3 expression to identify T cells (presented in a histogram). D, CD4 and CD8 T cells were identified. Each dot represents one T cell.
(Data courtesy Dr. Tom Alexander, Akron, Ohio.)
Immunoassays for Antibody and Soluble Antigen
The enzyme-linked immunosorbent assay (ELISA) uses antigen immobilized on a plastic surface, bead, or filter to capture and separate the specific antibody from other antibodies in a patient’s serum (Figure 6-5). An antihuman antibody with a covalently linked enzyme (e.g., horseradish peroxidase, alkaline phosphatase, β-galactosidase) then detects the affixed patient antibody. It is quantitated spectrophotometrically according to the intensity of the color produced in response to the enzyme conversion of an appropriate substrate. The actual concentration of specific antibody can be determined by comparison with the reactivity of standard human antibody solutions. The many variations of ELISAs differ in the way in which they capture or detect antibody or antigen.
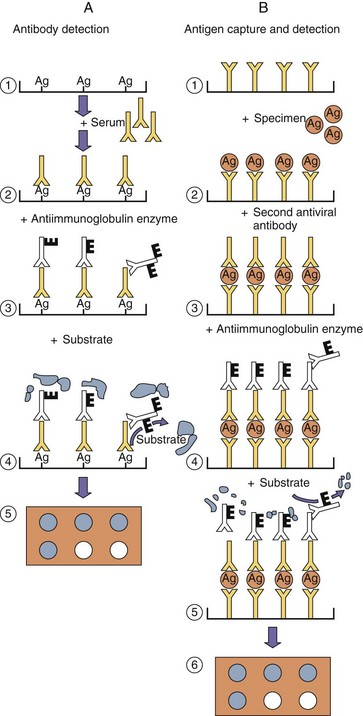
Figure 6-5 Enzyme immunoassays for quantitation of antibody or antigen. A, Antibody detection. 1, Viral antigen, obtained from infected cells, virions, or genetic engineering, is affixed to a surface. 2, Patient serum is added and allowed to bind to the antigen. Unbound antibody is washed away. 3, Enzyme-conjugated antihuman antibody (E) is added, and unbound antibody is washed away. 4, Substrate is added and converted (5) into chromophore, precipitate, or light. B, Antigen capture and detection. 1, Antiviral antibody is affixed to a surface. 2, A specimen that contains antigen is added, and unbound antigen is washed away. 3, A second antiviral antibody is added to detect the captured antigen. 4, Enzyme-conjugated antiantibody is added, washed, and followed by substrate (5), which is converted (6) into a chromophore, precipitate, or light.
ELISAs can also be used to quantitate the soluble antigen in a patient’s sample. In these assays, soluble antigen is captured and concentrated by an immobilized antibody and then detected with a different antibody labeled with the enzyme. An example of a commonly used ELISA is the home pregnancy test for the human chorionic gonadotropin hormone.
Western blot analysis is a variation of an ELISA. In this technique, viral proteins separated by electrophoresis according to their molecular weight or charge are transferred (blotted) onto a filter paper (e.g., nitrocellulose, nylon). When exposed to a patient’s serum, the immobilized proteins capture virus-specific antibody and are visualized with an enzyme-conjugated antihuman antibody. This technique shows the proteins recognized by the patient serum. Western blot analysis is used to confirm ELISA results in patients suspected to be infected with the human immunodeficiency virus (HIV) (Figure 6-6; also see Figure 47-7).
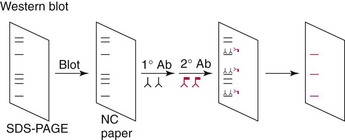
Figure 6-6 Western blot analysis. Proteins are separated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE), electroblotted onto nitrocellulose (NC) paper, and incubated with antigen-specific or patient’s antisera (1° Ab) and then enzyme-conjugated antihuman serum (2° Ab). Enzyme conversion of substrate identifies the antigen.
In radioimmunoassay (RIA), radiolabeled (e.g., with iodine-125) antibody or antigen is used to quantitate antigen-antibody complexes. RIA can be performed as a capture assay, as described previously for ELISA, or as a competition assay. In a competition assay, antibody in a patient’s serum is quantitated according to its ability to compete with and replace a laboratory-prepared, radiolabeled antibody from antigen-antibody complexes. The antigen-antibody complexes are precipitated and separated from free antibody, and the radioactivity is measured for both fractions. The amount of the patient’s antibody is then quantitated from standard curves prepared with use of known quantities of competing antibody. The radioallergosorbent assay is a variation of an RIA capture assay, in which radiolabeled anti-IgE is used to detect allergen-specific responses.
Complement fixation is a standard but technically difficult serologic test (Box 6-1). In this test, the patient’s serum sample is reacted with laboratory-derived antigen and extra complement. Antibody-antigen complexes bind, activate, and fix (use up) the complement. The residual complement is then assayed through the lysis of red blood cells coated with antibody. Antibodies measured by this system generally develop slightly later in an illness than those measured by other techniques.
Box 6-1
Serologic Assays
Hemagglutination inhibition*
Neutralization*
Immunofluorescence (direct and indirect)
In situ enzyme immunoassay (EIA)
Enzyme-linked immunosorbent assay (ELISA)
* For detection of antibody or serotyping of virus.
Antibody inhibition assays make use of the specificity of an antibody to prevent infection (neutralization) or other activity (hemagglutination inhibition) to identify the strain of the infecting agent, usually a virus, or to quantitate antibody responses to a specific strain of virus. For example, hemagglutination inhibition is used to distinguish different strains of influenza A. These tests are discussed further in Chapter 57.
Latex agglutination is a rapid, technically simple assay for detecting antibody or soluble antigen. Virus-specific antibody causes latex particles coated with viral antigens to clump. Conversely, antibody-coated latex particles are used to detect soluble viral antigen. In passive hemagglutination, antigen-modified erythrocytes are used as indicators instead of latex particles.
Serology
The humoral immune response provides a history of a patient’s infections. Serology can be used to identify the infecting agent, evaluate the course of an infection, or determine the nature of the infection—whether it is a primary infection or a reinfection, and whether it is acute or chronic. The antibody type and titer and the identity of the antigenic targets provide serologic data about an infection. Serologic testing is used to identify viruses and other agents that are difficult to isolate and grow in the laboratory or that cause diseases that progress slowly (Box 6-2).
Box 6-2
Viruses Diagnosed by Serology*
* Serologic testing is also used to determine a person’s immune status with regard to other viruses.
The relative antibody concentration is reported as a titer. A titer is the inverse of the greatest dilution, or lowest concentration (e.g., dilution of 1 : 64 = titer of 64), of a patient’s serum that retains activity in one of the immunoassays just described. The amount of IgM, IgG, IgA, or IgE reactive with antigen can also be evaluated through the use of a labeled second antihuman antibody that is specific for the antibody isotype.
Serology is used to determine the time course of an infection. Seroconversion occurs when antibody is produced in response to a primary infection. Specific IgM antibody, found during the first 2 to 3 weeks of a primary infection, is a good indicator of a recent primary infection. Reinfection or recurrence later in life causes an anamnestic (secondary or booster) response. Antibody titers may remain high, however, in patients whose disease recurs frequently (e.g., herpesviruses). Seroconversion or reinfection is indicated by the finding of at least a fourfold increase in the antibody titer between serum obtained during the acute phase of disease and that obtained at least 2 to 3 weeks later during the convalescent phase. A twofold serial dilution will not distinguish between samples with 512 and 1023 units of antibody, both of which would give a reaction on a 512-fold dilution but not on a 1024-fold dilution, and both results would be reported as titers of 512. On the other hand, samples with 1020 and 1030 units are not significantly different but would be reported as titers of 512 and 1024, respectively.
Serology can also be used to determine the stage of a slower or chronic infection (e.g., hepatitis B or infectious mononucleosis caused by Epstein-Barr virus), based on the presence of antibody to specific microbial antigens. The first antibodies to be detected are those directed against antigens most available to the immune system (e.g., on the virion, on surfaces of infected cells, secreted). Later in the infection, when cells have been lysed by the infecting virus or the cellular immune response, antibodies directed against the intracellular proteins and enzymes are detected.
Describe the diagnostic procedure or procedures (molecular or immunologic) that would be appropriate for each of the following applications:
1. Determination of the apparent molecular weights of the HIV proteins
2. Detection of human papillomavirus 16 (a nonreplicating virus) in a Papanicolaou (Pap) smear
3. Detection of herpes simplex virus (HSV) (a replicating virus) in a Pap smear
4. Presence of Histoplasma fungal antigens in a patient’s serum
5. CD4 and CD8 T-cell concentrations in blood from a patient infected with HIV
6. The presence of antibody and the titer of anti-HIV antibody
7. Genetic differences between two HSVs (DNA virus)
8. Genetic differences between two parainfluenza viruses (ribonucleic acid virus)
9. Amount of rotavirus antigen in stool
10. Detection of group A streptococci and their distinction from other streptococci
1. SDS-polyacrylamide gel electrophoresis to separate the proteins and Western blot to identify the HIV proteins are appropriate.
2. Genome detection methods, such as in situ hybridization on the Pap smear or a polymerase chain reaction (PCR) of the cells obtained during the procedure, can be used because virus proteins would be undetectable.
3. Cytopathologic effects, such as syncytia or Cowdry type A inclusion bodies, can be seen in Pap smears. Genome detection methods, such as in situ hybridization on the Pap smear or a PCR of DNA obtained from the cells or immunologic methods to detect virus antigen, can be used to detect evidence of the virus.
4. An Ouchterlony antibody diffusion or ELISA method can be used to detect fungal antigens.
5. Flow cytometry using immunofluorescence is probably the best method for identifying and quantitating CD4 and CD8 T cells.
6. ELISA is used to detect the presence and titer of anti-HIV antibody as a screening procedure for the blood supply. Western blot analysis with patient serum is used as a qualitative means to confirm ELISA results.
7. Restriction fragment length polymorphism or PCR can be used to detect genetic differences between strains or types of HSV.
8. Reverse transcriptase PCR can be used to distinguish two parainfluenza viruses.
9. Rotavirus in stool can be quantitated by ELISA. Immune electron microscopy is a qualitative method.
10. Group A Streptococcus can be detected by ELISA techniques, including rapid methods (similar to the over-the-counter pregnancy tests) for detecting streptolysin A and S. Fancier techniques, such as pulsed field gel electrophoresis of restriction fragments of the chromosome and PCR, can be used to distinguish different strains. Technology is also available to sequence portions of the genome of the different strains for comparison.
Forbes BA, Sahm DF, Weissfeld AS. Bailey and Scott’s diagnostic microbiology, ed 12. St Louis: Mosby; 2007.
Murray PR. ASM pocket guide to clinical microbiology, ed 3. Washington, DC: American Society for Microbiology Press; 2004.
Murray PR, et al. Manual of clinical microbiology, ed 9. Washington, DC: American Society for Microbiology Press; 2007.
Rosenthal KS, Wilkinson JG. Flow cytometry and immunospeak. Infect Dis Clin Pract. 2007;15:183–191.
Specter S, Hodinka RL, Young SA. Clinical virology manual, ed 3. Washington, DC: American Society for Microbiology Press; 2000.
Strauss JM, Strauss EG. Viruses and human disease, ed 2. San Diego: Academic; 2007.