47 Laboratory Diagnosis of Viral Diseases
There have been many new developments in laboratory viral diagnosis that provide more rapid and sensitive viral identification from clinical samples. These include better antibody reagents for direct analysis of samples, molecular genetics techniques and genomic sequencing for direct identification of the virus, and assays that can identify multiple viruses (multiplex) and be automated. Often, isolation of the organism is unnecessary and avoided to minimize the risk to laboratory and other personnel. The quicker turnaround allows a more rapid choice of the appropriate antiviral therapy.
Viral laboratory studies are performed to (1) confirm the diagnosis by identifying the viral agent of infection, (2) determine appropriate antiviral therapy, (3) check on the compliance of the patient taking antiviral drugs, (4) define the course of the disease, (5) monitor the disease epidemiologically, and (6) educate physicians and patients.
The laboratory methods accomplish the following results:
1. Description of virus-induced cytopathologic effects (CPEs) on cells
2. Detection of viral particles
3. Isolation and growth of the virus
4. Detection and analysis of viral components (e.g., proteins (antigens), enzymes, genomes)
5. Evaluation of the patient’s immune response to the virus (serology)
The molecular and immunologic techniques used for many of these procedures are described in Chapters 5 and 6. Viruses, viral antigens, viral genomes, and CPEs can be detected by means of direct analysis of clinical specimens and, for some viruses, after growth of the virus on tissue culture cells in the laboratory (Box 47-1).
Specimen Collection
The patient’s symptoms and history, including recent travel, the season of the year, and a presumptive diagnosis, help determine the appropriate procedures to be used to identify a viral agent (Table 47-1). The selection of the appropriate specimen for analysis is often complicated because several viruses may cause the same clinical disease. For example, the development of meningitis symptoms during the summer suggests an arbovirus, in which case cerebrospinal fluid (CSF) and blood should be collected, or an enterovirus, in which case CSF, a throat swab, and stool specimens should be collected for genome analysis and possible virus isolation. A focal encephalitis with a temporal lobe localization preceded by headaches and disorientation suggests herpes simplex virus (HSV) infection, for which CSF can be relatively quickly analyzed for viral deoxyribonucleic acid (DNA) sequences by polymerase chain reaction (PCR) amplification.
Table 47-1 Specimens for Viral Diagnosis
| Common Pathogenic Viruses | Specimens for Culture | Procedures and Comments |
|---|---|---|
| Respiratory Tract | ||
| Influenza virus; paramyxoviruses; coronavirus; rhinovirus; enterovirus (picornavirus) | Nasal washing, throat swab, nasal swab, sputum | RT-PCR, ELISA, multiplex assays detect several agents; cell culture |
| Gastrointestinal Tract | ||
| Reovirus; rotavirus; adenovirus; Norwalk virus, other calicivirus | Stool, rectal swab | RT-PCR, ELISA; viruses are not cultured |
| Maculopapular Rash | ||
| Adenovirus; enterovirus (picornavirus) | Throat swab, rectal swab | PCR, RT-PCR |
| Rubella virus; measles virus | Urine | RT-PCR, ELISA |
| Vesicular Rash | ||
| Coxsackievirus; echovirus; HSV; VZV | Vesicle fluid, scraping, or swab, enterovirus in stool | HSV and VZV: vesicle scraping (Tzanck smear), cell culture; HSV typing by PCR, IF |
| Central Nervous System (Aseptic Meningitis, Encephalitis) | ||
| Enterovirus (picornavirus) | Stool, CSF | RT-PCR |
| Arboviruses (e.g., togaviruses, bunyavirus) | Blood, CSF; rarely cultured | RT-PCR, serology; multiplex assays detect several agents |
| Rabies virus | Tissue, saliva, brain biopsy, CSF | IF of biopsy, RT-PCR |
| HSV; CMV; mumps virus; measles virus | Cerebrospinal fluid | PCR or RT-PCR, virus isolation, and antigen are assayed |
| Urinary Tract | ||
| Adenovirus; CMV | Urine | PCR; CMV may be shed without apparent disease |
| Blood | ||
| HIV; human T-cell leukemia virus; hepatitis B, C, and D viruses, EBV, CMV, HHV-6 | Blood | ELISA for antigen or antibody, PCR, and RT-PCR; multiplex assays detect several agents |
CMV, Cytomegalovirus; EBV, Epstein-Barr virus; ELISA, enzyme-linked immunosorbent assay; HIV, human immunodeficiency virus; HHV-6, human herpes virus 6; HSV, herpes simplex virus; IF, immunofluorescence; PCR, polymerase chain reaction; RT-PCR, reverse transcriptase polymerase chain reaction; VZV, varicella-zoster virus.
Specimens should be collected early in the acute phase of infection, before the virus ceases to be shed. For example, respiratory viruses may be shed for only 3 to 7 days, and shedding may lapse before the symptoms cease. HSV and varicella-zoster virus (VZV) may not be recoverable from lesions more than 5 days after the onset of symptoms. It may be possible to isolate an enterovirus from the CSF for only 2 to 3 days after the onset of the central nervous system manifestations. In addition, antibody produced in response to the infection may block the detection of virus.
The shorter the interval between the collection of a specimen and its delivery to the laboratory, the greater the potential for isolating a virus. The reasons are that many viruses are labile, and the samples are susceptible to bacterial and fungal overgrowth. Viruses are best transported and stored on ice and in special media that contain antibiotics and proteins, such as serum albumin or gelatin. Significant losses in infectious titers occur when enveloped viruses (e.g., HSV, VZV, influenza virus) are kept at room temperature or frozen at −20° C. This is not a risk for nonenveloped viruses (e.g., adenoviruses, enteroviruses).
Cytology
Many viruses produce a characteristic CPE. Characteristic CPEs in the tissue sample or in cell culture include changes in cell morphology, cell lysis, vacuolation, syncytia (Figure 47-1), and inclusion bodies. Syncytia are multinucleated giant cells formed by viral fusion of individual cells. Paramyxoviruses, HSV, VZV, and human immunodeficiency virus (HIV) promote syncytia formation. Inclusion bodies are either histologic changes in the cells caused by viral components or virus-induced changes in cell structures. For example, intranuclear basophilic (owl’s-eye) inclusion bodies found in large cells of tissues with cytomegalovirus (CMV) (see Chapter 51, Figure 51-17) or in the sediment of urine from patients with the infection are readily identifiable. Cowdry type A nuclear inclusions in single cells or in large syncytia (multiple cells fused together) are a characteristic finding in cells infected with HSV or VZV (Figure 47-2). Rabies may be detected through the finding of cytoplasmic Negri bodies (rabies virus inclusions) in brain tissue (Figure 47-3).
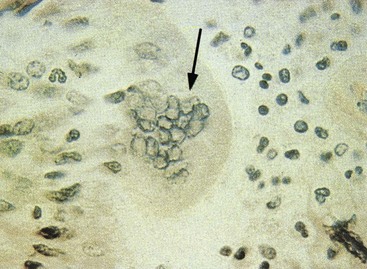
Figure 47-1 Syncytium formation by measles virus. Multinucleated giant cell (arrow) visible in a histologic section of lung biopsy tissue from a measles virus-induced giant cell pneumonia in an immunocompromised child.
(From Hart C, Broadhead RL: A color atlas of pediatric infectious diseases, London, 1992, Wolfe.)
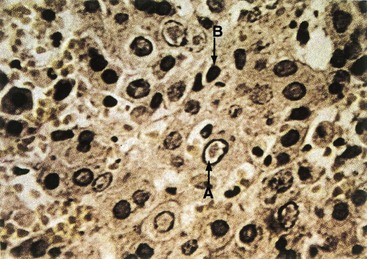
Figure 47-2 Herpes simplex virus (HSV)-induced cytopathologic effect. A biopsy specimen of an HSV-infected liver shows an eosinophilic Cowdry type A intranuclear inclusion body (A) surrounded by a halo and a ring of marginated chromatin at the nuclear membrane. An infected cell (B) exhibits a smaller condensed nucleus (pyknotic).
(Courtesy Dr JI Pugh, St Albans City Hospital, Hertfordshire, England; from Emond RT, Rowland HAK: A color atlas of infectious diseases, ed 3, London, 1995, Mosby.)
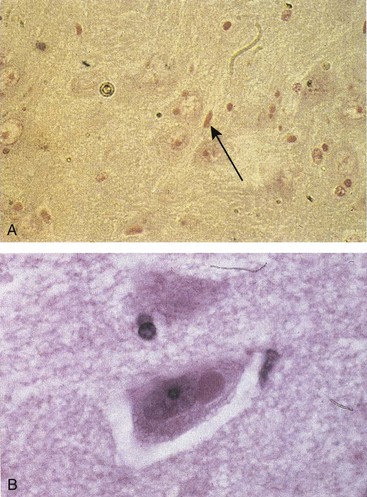
Figure 47-3 Negri bodies caused by rabies. A, A section of brain from a patient with rabies shows Negri bodies (arrow). B, Higher magnification from another biopsy specimen.
(A, From Hart C, Broadhead RL: A color atlas of pediatric infectious diseases, London, 1992, Wolfe.)
Often, the cytologic specimens will be examined for the presence of specific viral antigens by immunofluorescence or viral genomes by in situ hybridization or PCR for a rapid, definitive identification. These tests are specific for individual viruses and must be chosen based on the differential diagnosis. These methods are discussed in the following paragraphs.
Electron Microscopy
Electron microscopy is not a standard clinical laboratory technique, but it can be used to detect and identify some viruses if sufficient viral particles are present. The addition of virus-specific antibody to a sample can cause viral particles to clump, thereby facilitating the detection and simultaneous identification of the virus (immunoelectron microscopy). Enteric viruses, such as rotavirus, that are produced in abundance and have a characteristic morphology can be detected in stool by these methods. Appropriately processed tissue from a biopsy or clinical specimen can also be examined for the presence of viral structures.
Viral Isolation and Growth
A virus can be grown in tissue culture, embryonated eggs, and experimental animals (Box 47-2). Although embryonated eggs are still used for the growth of virus for some vaccines (e.g., influenza), they have been replaced by cell cultures for routine virus isolation in clinical laboratories. Experimental animals are rarely used in clinical laboratories for the purpose of isolating viruses.
Cell Culture
Specific types of tissue culture cells are used to grow viruses. Primary cell cultures are obtained by dissociating specific animal organs with trypsin or collagenase. The cells obtained by this method are then grown as monolayers (fibroblast or epithelial) or in suspension (lymphocyte) in artificial media supplemented with bovine serum or another source of growth factors. Primary cells can be dissociated with trypsin, diluted, and allowed to grow into new monolayers (passed) to become secondary cell cultures. Diploid cell lines are cultures of a single cell type that are capable of being passed a large but finite number of times before they senesce or undergo a significant change in their characteristics. Tumor cell lines and immortalized cell lines, usually initiated from human or animal tumors or by treatment of primary cells with oncogenic viruses or chemicals, consist of single cell types that can be passed continuously without senescing.
Primary monkey kidney cells are excellent for the recovery of influenza viruses, paramyxoviruses, many enteroviruses, and some adenoviruses. Human fetal diploid cells, which are generally fibroblastic cells, support the growth of a broad spectrum of viruses (e.g., HSV, VZV, CMV, adenoviruses, picornaviruses). HeLa cells, a continuous line of epithelial cells derived from a human cancer, are excellent for the recovery of respiratory syncytial virus, adenoviruses, and HSV. Many clinically significant viruses can be recovered in at least one of these cell cultures.
Viral Detection
A virus can be detected and initially identified through observation of the virus-induced CPE in the cell monolayer (Box 47-3; Figure 47-4), by immunofluorescence, or genome analysis of the infected cell culture. For example, a single virus infects, spreads, and kills surrounding cells (plaque). The type of cell culture, the characteristics of the CPE, and the rapidity of viral growth can be used to initially identify many clinically important viruses. This approach to identifying viruses is similar to that used in the identification of bacteria, which is based on the growth and morphology of colonies on selective differential media.
Box 47-3
Viral Cytopathologic Effects
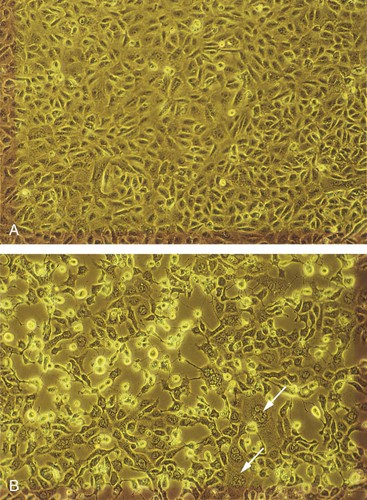
Figure 47-4 Cytopathologic effect of herpes simplex virus (HSV) infection. A, Uninfected Vero cells, an African green monkey kidney cell line. B, HSV-1–infected Vero cells showing rounded cells, multinucleated cells, and loss of the monolayer. Arrows point to syncytia.
Some viruses grow slowly or not at all or do not readily cause a CPE in cell lines typically used in clinical virology laboratories. Some cause diseases that are hazardous to personnel. These viruses are most frequently diagnosed on the basis of serologic findings or through the detection of viral genomes or proteins.
Characteristic viral properties can also be used to identify viruses that do not have a classic CPE. For example, the rubella virus may not cause a CPE, but it does prevent (interfere with) the replication of picornaviruses in a process known as heterologous interference, which can be used to identify the rubella virus. Cells infected with the influenza virus, parainfluenza virus, mumps virus, and togavirus express a viral glycoprotein (hemagglutinin) that binds erythrocytes of defined animal species to the infected cell surface (hemadsorption) (Figure 47-5). When released into the cell culture medium, such viruses can be detected from the agglutination of erythrocytes, a process termed hemagglutination. The virus can then be identified from the specific antibody that blocks the hemagglutination, a process called hemagglutination inhibition (HI). An innovative approach to detection of HSV infection uses genetically modified tissue culture cells that express the β-galactosidase gene and can be stained blue when infected with HSV (enzyme-linked virus-inducible system [ELVIS]).
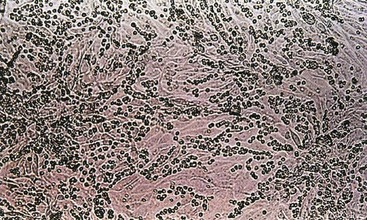
Figure 47-5 Hemadsorption of erythrocytes to cells infected with influenza viruses, mumps virus, parainfluenza viruses, or togaviruses. These viruses express a hemagglutinin on their surfaces, which binds to erythrocytes of selected animal species.
One can quantitate a virus by determining the greatest dilution that retains the following properties (titer):
1. Tissue culture dose (TCD50): titer of virus that causes cytopathologic effects in half the tissue culture cells
2. Lethal dose (LD50): titer of virus that kills 50% of a set of test animals
3. Infectious dose (ID50): titer of virus that initiates a detectable symptom, antibody, or other response in 50% of a set of test animals
The number of infectious viruses can also be evaluated with a count of the plaques produced by 10-fold dilutions of sample (plaque-forming units). The ratio of viral particles (from electron microscopy) to plaque-forming units is always much greater than one because numerous defective viral particles are produced during viral replication.
Interpretation of Culture Results
In general, the detection of any virus in host tissues, CSF, blood, or vesicular fluid can be considered a highly significant finding. However, viral shedding may occur and be unrelated to the disease symptoms. Certain viruses can be intermittently shed without causing symptoms in the affected person for periods ranging from weeks (enteroviruses in feces) to many months or years (HSV or CMV in the oropharynx and vagina; adenoviruses in the oropharynx and intestinal tract). Similarly, a negative result cannot be conclusive because the sample may have been improperly handled, contain neutralizing antibody, or be acquired before or after viral shedding.
Detection of Viral Proteins
Enzymes and other proteins are produced during viral replication and can be detected by biochemical, immunologic, and molecular biologic means (Box 47-4). The viral proteins can be separated by electrophoresis and their patterns used to identify and distinguish different viruses. For example, the electrophoretically separated HSV-infected cell proteins and virion proteins exhibit different patterns for different types and strains of HSV-1 and HSV-2.
Box 47-4
Assays for Viral Proteins and Nucleic Acids
Nucleic Acids
Restriction endonuclease cleavage patterns
Size of RNA for segmented RNA viruses (electrophoresis)
DNA genome hybridization in situ (cytochemistry)
Southern, Northern, and dot blots
Reverse transcriptase polymerase chain reaction (RNA)
DNA, Deoxyribonucleic acid; PCR, polymerase chain reaction; RNA, ribonucleic acid.
The detection and assay of characteristic enzymes or activities can identify and quantitate specific viruses. For example, the presence of reverse transcriptase in serum or cell culture indicates the presence of a retrovirus or hepadnavirus. Similarly, hemagglutination or hemadsorption can be used to easily assay the hemagglutinin produced by the influenza virus.
Antibodies can be used as sensitive and specific tools to detect, identify, and quantitate the virus and viral antigen in clinical specimens or cell cultures (immunohistochemistry). Specifically, monoclonal or monospecific antibodies are useful for distinguishing viruses. Viral antigens on the cell surface or within the cell can be detected by immunofluorescence and enzyme immunoassay (EIA) (see Chapter 6, Figures 6-2 and 6-3). Virus or antigen released from infected cells can be detected by enzyme-linked immunosorbent assay (ELISA), radioimmunoassay (RIA), and latex agglutination (LA) (see Chapter 6 for definitions). Test kits for single and multiple (multiplex) viral agents are commercially available. Multiplex kits for respiratory viruses can help to determine whether antiviral therapy is available.
The detection of CMV and other viruses can be enhanced through the use of a combination of cell culture and immunologic means. In this method, the clinical sample is centrifuged onto cells grown on a coverslip on the bottom of a shell vial (glass tube). This step increases the efficiency and accelerates the progression of infection of the cells on the coverslip. The cells can then be analyzed with immunofluorescence (direct fluorescence) or EIA for early viral antigens, which are detectable within 24 hours, instead of the 7 to 14 days it takes for a CPE to become evident.
Detection of Viral Genetic Material
The genetic sequence of a virus is a major distinguishing characteristic of the family, type, and strain of virus (see Box 47-4). The electrophoretic patterns of ribonucleic acid (RNA) (influenza, reovirus) or restriction endonuclease fragment lengths from DNA viral genomes are like genetic fingerprints for these viruses. Different strains of HSV-1 and HSV-2 can be distinguished in this way by restriction fragment length polymorphism. Newer methods for viral genome detection use sequence-specific genetic probes and PCR-like DNA and RNA amplification approaches, which allow more rapid analysis and quantitation with a minimum of risk from infectious virus. Methods for sequencing the genomes of viruses are becoming rapid and inexpensive enough to become a routine viral identification method.
DNA probes, with sequences complementary to specific regions of a viral genome, can be used like antibodies as sensitive and specific tools for detecting a virus. These probes can detect the virus even in the absence of viral replication. DNA probe analysis is especially useful for detecting slowly replicating or nonproductive viruses, such as CMV and human papillomavirus, or the viral antigen cannot be detected using immunologic tests (see Chapter 5, Figure 5-3). Specific viral genetic sequences in fixed, permeabilized tissue biopsy specimens can be detected by in situ hybridization (e.g., fluorescence in situ hybridization [FISH]).
Viral genomes can also be detected in clinical samples with the use of dot blot or Southern blot analysis. For the latter method, the viral genome or electrophoretically separated restriction endonuclease cleavage fragments of the genome are blotted onto nitrocellulose filters and then detected on the filter by their hybridization to DNA probes. Electrophoretically separated viral RNA (Northern blot–RNA : DNA probe hybridization) blotted onto a nitrocellulose filter can be detected in a similar manner. The DNA probes are detected with autoradiography or with fluorescent or EIA-like methods. Many viral probes and kits for detecting viruses are now commercially available.
For many laboratories, genome amplification techniques, including PCR for DNA genomes and reverse transcriptase PCR (RT-PCR) for RNA genomes, are the method of choice for detection and identification of viruses. Use of the appropriate primers for PCR can promote a millionfold amplification of a target sequence in a few hours. This technique is especially useful for detecting latent and integrated sequences of viruses, such as retroviruses, herpesviruses, papillomaviruses, and other papovaviruses, as well as evidence of viruses present in low concentrations and viruses that are difficult or too dangerous to isolate in cell culture. RT-PCR uses the retroviral reverse transcriptase to convert viral RNA to DNA and allow PCR amplification of the viral nucleic acid sequences. This approach was very useful for identifying and distinguishing the Hantaviruses that caused the outbreak in New Mexico in 1993. These techniques are readily automated to analyze multiple samples and for different viruses (multiplex).
Quantification of the amount of genomes within a patient (virus load) can be determined by real-time PCR. For example, the concentration of viral genome (RNA genomes would be converted to DNA) is proportional to the initial rate of PCR amplification of the genomic DNA. This test is especially important for following the course of HIV infection.
PCR is the prototype for several other HIV genome amplification techniques. Transcription-based amplification uses reverse transcriptase and viral sequence specific primers to make a complementary DNA (cDNA) and attaches a sequence recognized by the DNA-dependent RNA polymerase from the T7 bacteriophage. The DNA is transcribed to RNA by the T7 RNA polymerase, and the new RNA sequences are then cycled back into the reaction to amplify the relevant sequence. Unlike PCR, these reactions do not require special equipment.
Some other genome amplification and detection approaches are similar in concept to ELISA. These approaches use immobilized DNA sequences complementary to the relevant viral genomic sequence to capture the viral genome. This is followed by the binding of another complementary sequence that contains a detection system. The genome probe sequence may be attached to an extensively branched chain of DNA in which each of the branches elicits a reaction that amplifies the signal to detectable levels. Another variation of the theme uses an antibody that recognizes DNA-RNA complexes to capture viral DNA-RNA probe hybrids in the well of a plate, followed by an enzyme-labeled antibody and ELISA methods to detect the presence of the genome. Like ELISA, these methods can be automated and set up to analyze a panel of viruses.
Viral Serology
The humoral immune response provides a history of a patient’s infections. Serologic studies are used for the identification of viruses that are difficult to isolate and grow in cell culture, as well as viruses that cause diseases of long duration (see Box 6-2). Serology can be used to identify the virus and its strain or serotype, whether it is an acute or chronic disease, and determine whether it is a primary infection or a reinfection. The detection of virus-specific immunoglobulin M (IgM) antibody, which is present during the first 2 or 3 weeks of a primary infection, generally indicates a recent primary infection. Seroconversion is indicated by at least a fourfold increase in the antibody titer between the serum obtained during the acute phase of disease and that obtained at least 2 to 3 weeks later during the convalescent phase. Reinfection or recurrence later in life causes an anamnestic (secondary or booster) response. Antibody titers may remain high in patients who suffer frequent recurrence of a disease (e.g., herpesviruses).
Because of the inherent imprecision of serologic assays based on twofold serial dilutions, a fourfold increase in the antibody titer between acute and convalescent sera is required to indicate seroconversion. For example, samples with 512 and 1023 units of antibody would both give a signal on a 512-fold dilution but not on a 1024-fold dilution, and the titers of both would be reported as 512. On the other hand, samples with 1020 and 1030 units are not significantly different but would be reported as titers of 512 and 1024, respectively.
The course of a chronic infection can also be evaluated by a serologic profile. Specifically, the presence of antibodies to several key viral antigens and their titers can be used to identify the stage of disease caused by certain viruses. This approach is especially useful for the diagnosis of viral diseases with slow courses (e.g., hepatitis B, infectious mononucleosis caused by Epstein-Barr virus). In general, the first antibodies to be detected are directed against the antigens most available to the immune system (e.g., expressed on the virion or infected-cell surfaces). Later in the infection, when the infecting virus or the cellular immune response has lysed the cells, antibodies directed against the intracellular viral proteins and enzymes are detected. For example, antibodies to the envelope and capsid antigens of Epstein-Barr virus are detected first. Then during convalescence, antibodies to nuclear antigens, such as the Epstein-Barr virus nuclear antigen, are detected.
A serologic battery or panel consisting of assays for several viruses may be used for the diagnosis of certain diseases. Local epidemiologic factors, the time of year, and patient factors such as immunocompetence, travel history, and age influence the choice of virus assays to be included in a panel. For example, HSV and the viruses of mumps, western and eastern equine encephalitides, and St. Louis, West Nile, and California encephalitides might be included in a panel of tests for central nervous system diseases.
Serologic Test Methods
The serologic tests used in virology are listed in Chapter 6, Box 6-1. Neutralization and HI tests assay antibody on the basis of its recognition of and binding to virus. The antibody coating of the virus blocks its binding to indicator cells (Figure 47-6). Antibody neutralization of virus inhibits infection and subsequent cytopathologic effects in tissue culture cells. HI is used for the identification of viruses that can selectively agglutinate erythrocytes of various animal species (e.g., chicken, guinea pig, human). Antibody in serum prevents a standardized amount of virus from binding to and agglutinating erythrocytes.
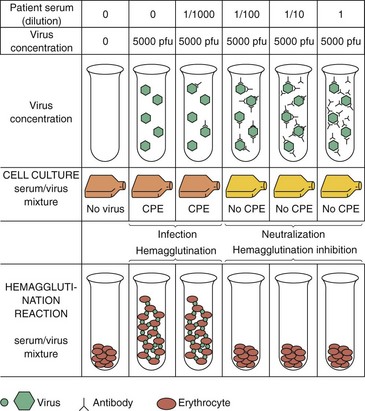
Figure 47-6 Neutralization, hemagglutination, and hemagglutination inhibition assays. In the assay shown, 10-fold dilutions of serum were incubated with virus. Aliquots of the mixture were then added to cell cultures or erythrocytes. In the absence of antibody, the virus infected the monolayer (indicated by cytopathologic effect [CPE]) or caused hemagglutination (i.e., formed a gel-like suspension of erythrocytes). In the presence of the antibody, infection was blocked, preventing CPE (neutralization), or hemagglutination was inhibited, allowing the erythrocytes to pellet. The titer of antibody of this serum would be 100.
The indirect fluorescent antibody test and solid-phase immunoassays, such as latex agglutination and ELISA, are commonly used to detect and quantitate viral antigen and antiviral antibody. The ELISA test is used to screen the blood supply to exclude individuals who are seropositive for hepatitis B and C viruses and HIV. Western blot analysis has become very important to confirm seroconversion and hence infection with HIV. The ability of the patient’s antibody to recognize specific viral proteins separated by electrophoresis, transferred (blotted) onto a filter paper (e.g., nitrocellulose, nylon), and visualized with an enzyme-conjugated antihuman antibody confirms the ELISA-indicated diagnosis of HIV infection (Figure 47-7).
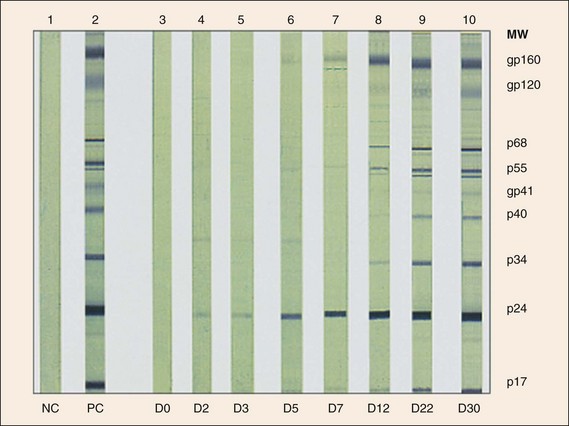
Figure 47-7 Western blot analysis of human immunodeficiency virus (HIV) antigens and antibody. HIV protein antigens are separated by electrophoresis and blotted onto nitrocellulose paper strips. The strip is incubated with patient antibody, washed to remove the unbound antibody, and then reacted with enzyme-conjugated antihuman antibody and chromophoric substrate. Serum from an HIV-infected person binds and identifies the major antigenic proteins of HIV. These data demonstrate the seroconversion of one HIV-infected individual with sera collected on day 0 (D0) to day 30 (D30) compared to a known positive control (PC) and negative control (NC). MW, molecular weight.
(From Kuritzkes DR: Diagnostic tests for HIV infection and resistance assays. In Cohen J, Powderly WG: Infectious diseases, ed 2, St Louis, 2004, Mosby.)
Limitations of Serologic Methods
The presence of an antiviral antibody indicates previous infection but is not sufficient to indicate when the infection occurred. The finding of virus-specific IgM, a fourfold increase in the antibody titer between acute and convalescent sera, or specific antibody profiles is indicative of recent infection. False-positive or false-negative test results may confuse the diagnosis. In addition, patient antibody may be bound with viral antigen (as occurs in patients with hepatitis B) in immune complexes, thereby preventing antibody detection. Serologic cross-reactions between different viruses may also confuse the identity of the infecting agent (e.g., parainfluenza and mumps express related antigens). Conversely, the antibody used in the assay may be too specific (many monoclonal antibodies) and may not recognize strains of virus from the same family, giving a false-negative result (e.g., rhinovirus). A good understanding of the clinical symptoms and knowledge of the limitations and potential problems with serologic assays aid the diagnosis.
1. Brain tissue is obtained at autopsy from a person who died of rabies. What procedures could be used to confirm the presence of rabies virus-infected cells in the brain tissue?
2. A cervical Papanicolaou smear is taken from a woman with a vaginal papilloma (wart). Certain types of papilloma have been associated with cervical carcinoma. What method or methods would be used to detect and identify the type of papilloma in the cervical smear?
3. A legal case would be settled by identification of the source of an HSV infection. Serum and viral isolates are obtained from the infected person and two contacts. What methods could be used to determine whether the person is infected with HSV-1 or HSV-2? What methods could be used to compare the type and strain of HSV obtained from each of the three people?
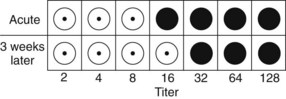
4. A 50-year-old man experiences flulike symptoms. The figure below shows results of hemagglutination inhibition tests on serum specimens collected when the disease manifested (acute) and 3 weeks later. The HI data for the current strain of influenza A (H3N2) are presented at top right. Filled circles indicate hemagglutination. Is the patient infected with the current strain of influenza A?
5. A policeman accidentally sticks his finger with a drug addict’s syringe needle. He is concerned that he may be infected with HIV. Samples are taken from the policeman a month later for analysis. What assays would be appropriate to determine whether the man is infected with the virus? In this case, it may be too early to detect an antibody response to the virus. What procedures would be appropriate to assay for virus or viral components?
1. Rabies virus infection can be identified by observation of Negri inclusion bodies and the presence of viral proteins by immunofluorescence. A tissue extract can also be analyzed by RT-PCR for viral genome.
2. The papilloma virus genome can be detected and typed by in situ hybridization and by PCR analysis using strain-specific DNA probes and primers. Immunofluorescence is not used because viral proteins may only be expressed in rare cells.
3. HSV-1 and HSV-2 can be distinguished with antibody specific for each virus type. The antibody can be used in a virus neutralization test, but a better approach is by immunofluorescence or ELISA test of cells infected by either virus using type-specific antibody. PCR tests are also available to distinguish HSV-1 and HSV-2. Genome sequencing can also be used.
Different strains of virus can be distinguished by PCR of variable regions of the genome or by restriction fragment length polymorphism. These techniques will also distinguish HSV-1 from HSV-2.
4. The figure shows that the titer of the convalescent serum taken 3 weeks after the acute serum is only different by one dilution tube (twofold). A significant difference in the titer of the antibody requires at least a fourfold difference. Therefore the patient was not infected by H3N2 virus.
5. Recent infection is indicated by detection of the presence of the HIV genome as performed by RT-PCR or a related technique. These techniques amplify the genome that may be present in the sample. The presence of the viral protein p24 would also be an indication of recent infection. It is too early for the person to provide a dependable indication of infection by the presence of antibodies to HIV.