57 Orthomyxoviruses
On April 15, 2009, a 33-year-old woman at 35 weeks’ gestation had a 1-day history of myalgias, dry cough, and low-grade fever when examined by her obstetrician-gynecologist. The patient had not recently traveled to Mexico. Rapid influenza diagnostic testing performed in the physician’s office was positive. On April 19, she was examined in a local emergency department, with worsening shortness of breath, fever, and productive cough. She experienced severe respiratory distress, and was intubated and placed on mechanical ventilation. An emergency cesarean delivery was performed, resulting in a healthy female infant. On April 21, the patient developed acute respiratory distress syndrome (ARDS). The patient began receiving oseltamivir on April 28 and broad-spectrum antibiotics but died on May 4.*
1. How did the woman acquire the infection?
2. What is the normal presentation, and what is abnormal about this presentation of influenza?
Answers
1. Breathing contaminated aerosol.
2. Normally, there is an abrupt onset of fever, chills, severe myalgias, loss of appetite, weakness and fatigue, sore throat, and usually a nonproductive cough. The fever persists for 3 to 8 days, and unless a complication occurs, recovery is complete within 7 to 10 days. This woman suffered ARDS.
3. Cell-mediated immunity is suppressed in pregnant women. This allowed the virus to replicate and spread to a greater extent and enhanced the pathogenicity of the infection.
4. This H1N1 strain is a reassortant of viral strains from humans, pigs, and ducks generated by subsequent infections of pigs with virus from duck and then human and other pig viruses. It created a unique H1N1 virus.
Influenza A, B, and C viruses are the only members of the Orthomyxoviridae family, and only influenza A and B viruses cause significant human disease. The orthomyxoviruses are enveloped and have a segmented negative-sense ribonucleic acid (RNA) genome. The segmented genome of these viruses facilitates the development of new strains through the mutation and reassortment of the gene segments among different human and animal (influenza A) strains of virus. This genetic instability is responsible for the annual epidemics (mutation: drift) and for influenza A, periodic pandemics (reassortment: shift) of influenza infection worldwide.
Influenza is one of the most prevalent and significant viral infections. Probably the most famous influenza pandemic (worldwide) is the Spanish influenza that swept the world in 1918 to 1919, killing 20 to 40 million people. In fact, more people died of influenza during that time than in the battles of World War I. Pandemics caused by novel influenza viruses occurred in 1918, 1947, 1957, 1968, 1977, and 2009. An outbreak of avian influenza first noted in Hong Kong in 1997, as well as the pandemic in 2009, caused human disease and fatalities. Fortunately, prophylaxis in the form of vaccines and antiviral drugs is now available for people at risk for serious outcomes.
Influenza viruses cause respiratory symptoms and the classic flulike symptoms of fever, malaise, headache, and myalgias (body aches). The term flu, however, has been mistakenly used to refer to many other respiratory and viral infections (e.g., “intestinal flu”).
Structure and Replication
Influenza virions are pleomorphic, appearing spheric or tubular (Box 57-1 and Figure 57-1) and ranging in diameter from 80 to 120 nm. The envelope contains two glycoproteins, hemagglutinin (HA) and neuraminidase (NA), the membrane (M2) protein and is internally lined by the matrix (M1) protein. The genome of the influenza A and B viruses consists of eight different helical nucleocapsid segments, each of which contains a negative-sense RNA associated with the nucleoprotein (NP) and the transcriptase (RNA polymerase components: PB1, PB2, PA) (Table 57-1). Influenza C has only seven genomic segments.
Box 57-1
Unique Features of the Influenza A and B Viruses
The enveloped virion has a genome of eight unique negative-sense RNA nucleocapsid segments.
Hemagglutinin glycoprotein is the viral attachment protein and fusion protein; it elicits neutralizing, protective antibody responses.
Influenza transcribes and replicates its genome in the target cell nucleus but assembles and buds from the plasma membrane.
The antiviral drugs amantadine and rimantadine inhibit an uncoating step and target the M2 (membrane) protein for influenza A only.
The antiviral drugs zanamivir and oseltamivir inhibit the neuraminidase protein of influenza A and B.
The segmented genome promotes genetic diversity caused by mutation and reassortment of segments on infection with two different strains.
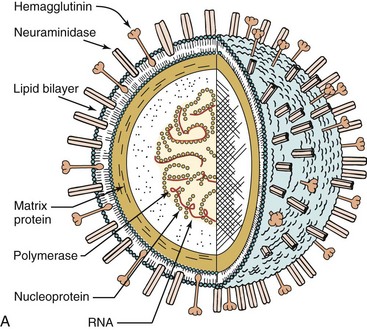
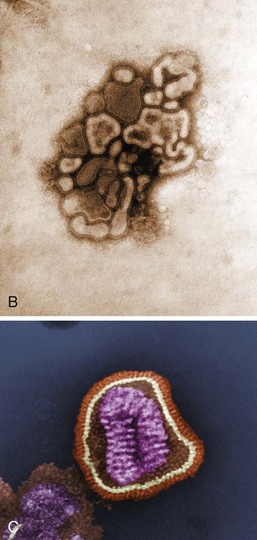
Figure 57-1 A, Model of influenza A virus. B and C, Electron micrographs of influenza A virus. RNA, Ribonucleic acid.
(A, From Kaplan MM, Webster RG: The epidemiology of influenza, Sci Am 237:88–106, 1977. B, From Balows A, et al: Laboratory diagnosis of infectious diseases: principles and practice, vol 2, New York, 1988, Springer-Verlag.)
Table 57-1 Products of Influenza Gene Segments
| Segment* | Protein | Function |
|---|---|---|
| 1 | PB2 | Polymerase component |
| 2 | PB1 | Polymerase component |
| 3 | PA | Polymerase component |
| 4 | HA | Hemagglutinin, viral attachment protein, fusion protein, target of neutralizing antibody |
| 5 | NP | Nucleocapsid |
| 6 | NA | Neuraminidase (cleaves sialic acid and promotes virus release) |
| 7† | M1 | Matrix protein: viral structural protein (interacts with nucleocapsid and envelope, promotes assembly) |
| M2 | Membrane protein (forms membrane channel and target for amantadine, facilitates uncoating and HA production) | |
| 8† | NS1 | Nonstructural protein (inhibits cellular messenger RNA translation) |
| NS2 | Nonstructural protein (promotes export of nucleocapsid from nucleus) |
* Listed in decreasing order of size.
The genomic segments in the influenza A virus range from 890 to 2340 bases. All the proteins are encoded on separate segments, with the exception of the nonstructural proteins (NS1 and NS2) and the M1 and M2 proteins, which are transcribed from one segment each.
The HA forms a spike-shaped trimer; each unit is activated by a protease and is cleaved into two subunits held together by a disulfide bond (see Chapter 44, Figure 44-8). The HA has several functions. It is the viral attachment protein, binding to sialic acid on epithelial cell surface receptors; it promotes fusion of the envelope to the cell membrane at acidic pH; it hemagglutinates (binds and aggregates) human, chicken, and guinea pig red blood cells; and it elicits the protective neutralizing antibody response. Mutation-derived changes in HA are responsible for the minor (“drift”) and major (“shift”) changes in antigenicity. Shifts occur only with influenza A virus, and the different HAs are designated H1, H2…H16.
The NA glycoprotein forms a tetramer and has enzyme activity. The NA cleaves the sialic acid on glycoproteins, including the cell receptor. Cleavage of the sialic acid on virion proteins prevents clumping and facilitates the release of virus from infected cells, making NA a target for two antiviral drugs, zanamivir (Relenza), and oseltamivir (Tamiflu). The NA of influenza A virus also undergoes antigenic shift, and major differences acquire the designations N1, N2…N9.
The M1, M2, and NP proteins are type specific and are therefore used to differentiate influenza A from B or C viruses. The M1 proteins line the inside of the virion and promote assembly. The M2 protein forms a proton channel in membranes and promotes uncoating and viral release. The M2 of influenza A is a target for the antiviral drugs amantadine and rimantadine.
Viral replication begins with the binding of HA to sialic acid on cell surface glycoproteins (Figure 57-2). The different HAs (HA1-16) bind to different sialic acid structures. The virus is then internalized into a coated vesicle and transferred to an endosome. Acidification of the endosome causes the HA to bend over and expose hydrophobic fusion-promoting regions of the protein. The viral envelope then fuses with the endosome membrane. The proton channel formed by the M2 protein promotes acidification of the envelope contents to break the interaction between the M1 protein and the NP, allowing uncoating and delivery of the nucleocapsid into the cytoplasm.
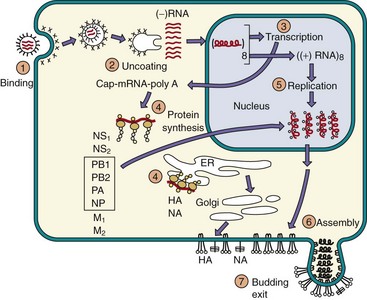
Figure 57-2 Replication of influenza A virus. After binding (1) to sialic acid–containing receptors, influenza is endocytosed and fuses (2) with the vesicle membrane. Unlike for most other ribonucleic acid (RNA) viruses, transcription (3) and replication (5) of the genome occur in the nucleus. Viral proteins are synthesized (4), helical nucleocapsid segments form and associate (6) with the M1 protein–lined membranes containing M2 and the hemagglutinin (HA) and neuraminadase (NA) glycoproteins. The virus buds (7) from the plasma membrane and eventually kills the cell. (−), Negative sense; (+), positive sense; ER, endoplasmic reticulum; NP, nucleocapsid; NS1, NS2, nonstructural proteins 1 and 2; PA, PB1, PB2, polymerase A, B1, and B2; polyA, polyadenylate.
Unlike most RNA viruses, the influenza nucleocapsid travels to the nucleus, where it is transcribed into messenger RNA (mRNA). The transcriptase (PA, PB1, and PB2) uses host cell mRNA as a primer for viral mRNA synthesis. In so doing, it steals the methylated cap region of the RNA, the sequence required for efficient binding to ribosomes. All the genomic segments are transcribed into 5′-capped, 3′-polyadenylated (polyA) mRNA for individual proteins except the segments for the M1, M2 and NS1, NS2 proteins, which are each differentially spliced (using cellular enzymes) to produce two different mRNAs. The mRNAs are translated into protein in the cytoplasm. The HA and NA glycoproteins are processed by the endoplasmic reticulum and Golgi apparatus. The M2 protein inserts into cellular membranes. Its proton channel prevents acidification of Golgi and other vesicles, thus preventing acid-induced folding and inactivation of the HA within the cell. The HA and NA are then transported to the cell surface.
Positive-sense RNA templates for each segment are produced, and the negative-sense RNA genome is replicated in the nucleus. The genomic segments associate with polymerase and NP proteins to form nucleocapsids, and the NS2 protein facilitates the transport of ribonucleocapsids into the cytoplasm, where they interact with the M1 protein–lined plasma membrane sections containing M2, HA, and NA. The virus buds selectively from the apical (airway) surface of the cell as a result of the preferential insertion of the HA in this membrane. Virus is released approximately 8 hours after infection.
Pathogenesis and Immunity
Influenza initially establishes a local upper respiratory tract infection (Box 57-2). To do so, the virus first targets and kills mucus-secreting, ciliated, and other epithelial cells, causing the loss of this primary defense system. NA facilitates the development of the infection by cleaving sialic acid (neuraminic acid) residues of the mucus, thereby providing access to tissue. Preferential release of the virus at the apical surface of epithelial cells and into the lung promotes cell-to-cell spread and transmission to other hosts. If the virus spreads to the lower respiratory tract, the infection can cause severe desquamation (shedding) of bronchial or alveolar epithelium down to a single-cell basal layer or to the basement membrane.
Box 57-2
Disease Mechanisms of Influenza A and B Viruses
Virus can establish infection of the upper and lower respiratory tract.
Systemic symptoms are caused by the interferon and cytokine response to the virus. Local symptoms result from epithelial cell damage, including ciliated and mucus-secreting cells.
Interferon and cell-mediated immune responses (natural killer and T cells) are important for immune resolution and immunopathogenesis.
Infected people are predisposed to bacterial superinfection because of the loss of natural barriers and exposure of binding sites on epithelial cells.
Antibody is important for future protection against infection and is specific for defined epitopes on hemagglutinin (HA) and neuraminidase (NA) proteins.
The HA and NA of influenza A virus can undergo major (reassortment: shift) and minor (mutation: drift) antigenic changes to ensure the presence of immunologically naïve, susceptible people.
In addition to compromising the mucociliary defenses of the respiratory tract, influenza infection promotes bacterial adhesion to the epithelial cells. Pneumonia may result from a viral pathogenesis or from a secondary bacterial infection. Influenza may also cause a transient or low-level viremia but rarely involves tissues other than the lung.
Influenza infection leads to an inflammatory cell response of the mucosal membrane, which consists primarily of monocytes and lymphocytes and few neutrophils. Submucosal edema is present. Lung tissue may reveal hyaline membrane disease, alveolar emphysema, and necrosis of the alveolar walls (Figure 57-3).
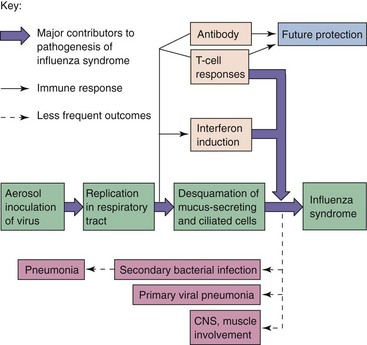
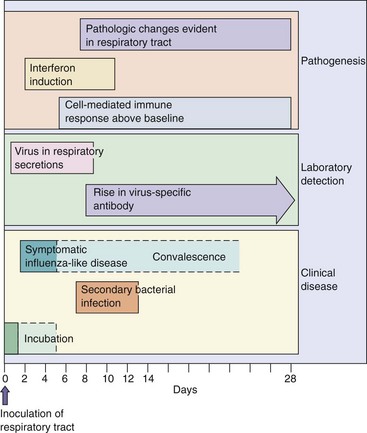
Figure 57-3 Pathogenesis of influenza A virus. The symptoms of influenza are caused by viral pathologic and immunopathologic effects, but the infection may promote secondary bacterial infection. CNS, Central nervous system.
Interferon and cytokine responses, which peak at almost the same time as virus in nasal washes, may be sufficient to control the infection, and are responsible for the systemic “flulike” symptoms. T-cell responses are important for effecting recovery and immunopathogenesis, but antibody, including vaccine-induced antibody, can prevent disease. As for measles, influenza infection depresses macrophage and T-cell function, hindering immune resolution. Of interest, recovery often precedes detection of antibody in serum or secretions.
Protection against reinfection is primarily associated with the development of antibodies to HA, but antibodies to NA are also protective. The antibody response is specific for each strain of influenza, but the cell-mediated immune response is more general and is capable of reacting to influenza strains of the same type (influenza A or B virus). Antigenic targets for T-cell responses include peptides from HA but also from the nucleocapsid proteins (NP, PB2) and M1 protein. The NP, PB2, and M1 proteins differ considerably for influenza A and B but minimally between strains of these viruses; hence T-cell memory may provide future protection against infection by different strains of either influenza A or B.
The symptoms and time course of the disease are determined by the extent of viral and immune killing of epithelial tissue and cytokine action. Influenza is normally a self-limited disease that rarely involves organs other than the lung. Many of the classic “flu” symptoms (e.g., fever, malaise, headache, and myalgia) are associated with interferon and cytokine induction. Repair of the compromised tissue is initiated within 3 to 5 days of the start of symptoms but may take as long as a month or more, especially for elderly people. The time course of influenza virus infection is illustrated in Figure 57-4.
Epidemiology
Strains of influenza A virus are classified by the following four characteristics:
For example, a current strain of influenza virus might be designated A/Bangkok/1/79 (H3N2), meaning that it is an influenza A virus that was first isolated in Bangkok in January 1979 and contains HA (H3) and NA (N2) antigens.
Strains of influenza B are designated by (1) type, (2) geography, and (3) date of isolation (e.g., B/Singapore/3/64), but without specific mention of HA or NA antigens, because influenza B does not undergo antigenic shift or pandemics as does influenza A.
Minor antigenic changes resulting from mutation of the HA and NA genes are called antigenic drift. This process occurs every 2 to 3 years, causing local outbreaks of influenza A and B infection. Major antigenic changes (antigenic shift) result from the reassortment of genomes among different strains, including animal strains. This process occurs only with the influenza A virus. Such changes are often associated with the occurrence of pandemics. In contrast to influenza A, influenza B is predominantly a human virus and does not undergo antigenic shift.
Antigenic shifts occur infrequently, but the pandemics they cause can be devastating (Table 57-2). For example, the prevalent influenza A virus in 1947 was the H1N1 subtype. In 1957, there was a shift in both antigens, resulting in an H2N2 subtype. H3N2 appeared in 1968, and H1N1 reappeared in 1977. The reappearance of H1N1 put the population younger than age 30 years at risk of disease. Prior exposure and an anamnestic antibody response protected the members of the population older than 30 years.
Table 57-2 Influenza Pandemics Resulting from Antigenic Shift
| Year of Pandemic | Influenza A Subtype |
|---|---|
| 1918 | H1N1 |
| 1947 | H1N1 |
| 1957 | H2N2; Asian flu strain |
| 1968 | H3N2; Hong Kong flu strain |
| 1977 | H1N1; Russian |
| 1997, 2003 | H5N1: China, Avian |
| 2009 | H1N1, Swine flu |
The genetic diversity of influenza A is fostered by its segmented genomic structure and ability to infect and replicate in humans and many animal species (zoonose), including birds and pigs. Hybrid viruses are created by coinfection of a cell with different strains of influenza A virus, allowing the genomic segments to randomly associate into new virions. An exchange of the HA glycoproteins may generate a new virus that can infect an immunologically naïve human population. Figure 57-5 depicts the origins of the pandemic A/California/04/2009/H1N1 virus through multiple reassortments of human, avian, and pig influenza viruses, resulting in a virus that was able to infect humans.
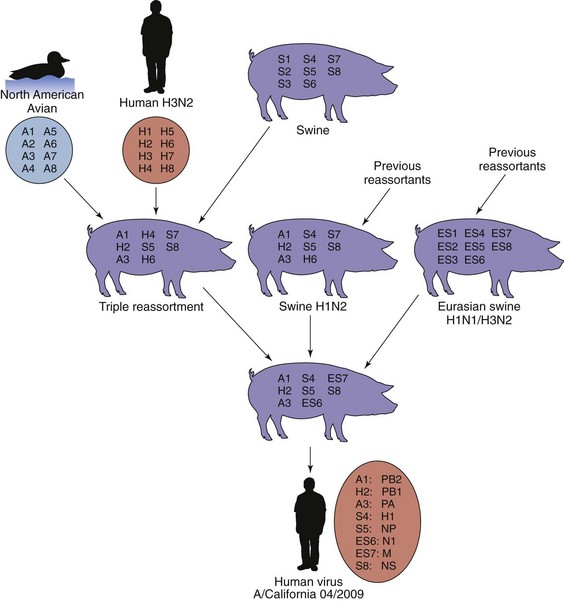
Figure 57-5 Generation of A/CALIFORNIA/04/2009(H1N1) pandemic swine flu by multiple reassortments of genomic segments of influenza A virus. The pandemic H1N1 virus arose from the mixing of a triple reassortment of avian, human and swine viruses with two other swine viruses, each of which were also generated by reassortment between swine, human, and other influenza viruses. This new virus emerged in the spring of 2009 (out of season) in Mexico but was first identified in California.
In the spring of 2009, a new amantadine- and rimantadine-resistant reassortant H1N1 virus was detected in a 10-year-old patient in California and proceeded to cause a pandemic. As indicated in Figure 57-5, the virus is a triple-triple reassortant of multiple swine, avian, and human influenza viruses. The virus originated in Mexico and rapidly spread as many cases went unrecognized because of the unseasonal nature of the outbreak. Up to 25,000 deaths have occurred worldwide, primarily in people between the ages of 22 months and 57 years. People with chronic medical conditions, especially pregnant women, were at greatest risk to complications, but unlike other outbreaks, this virus had a tendency to affect younger and healthier individuals. Of interest, many people older than 60 years had cross-reactive antibody resulting from prior exposure to an H1N1 influenza virus. Neuraminidase inhibitors were made available for prophylaxis, but detection of resistant strains became a concern. By September, a vaccine had been developed, approved, manufactured, and available for distribution on a prioritized basis and then was administered with the seasonal influenza vaccine. The pandemic was declared over by August 2010, and the H1N1 virus joined H3N2 and influenza B as a seasonal virus.
Because of its high population density and proximity of humans, pigs, chickens, and ducks, China is often the breeding ground for new reassortant viruses and the source of many of the pandemic strains of influenza. In 1997, a highly pathogenic avian influenza virus (HPAIV) (H5N1) strain was isolated from at least 18 humans and caused six deaths in Hong Kong (Clinical Case 57-1). The virus was spread by domestic and wild water fowl in their feces and directly from bird to man, with cases occurring around the globe. This avian H5N1 virus is unusual because it is not a reassortant; yet, it can infect and kill cells of the lower human lung. This requires inhalation of large amounts of virus (shared living environments). Outbreaks of avian influenza require the destruction of all potentially infected birds, such as for the 1.6 million chickens in Hong Kong, to destroy the potential source of the virus.
Clinical Case 57-1
H5N1 Avian Influenza
The first case of H5N1 avian influenza in a human was described by Ku and Chan (J Paediatr Child Health 35:207–208, 1999). After a 3-year-old boy from China developed a fever of 40° C and abdominal pain, he was given antibiotics and aspirin. On the 3rd day, he was hospitalized with sore throat, and his chest radiograph demonstrated bronchial inflammation. Blood studies showed a left shift with 9% band forms. On the 6th day, the boy was still febrile and fully conscious, but on the 7th day, his fever increased, he was hyperventilating, and his blood oxygen levels decreased. A chest radiograph indicated severe pneumonia. The patient was intubated. On the 8th day, the boy was diagnosed with fulminant sepsis and acute respiratory distress syndrome (ARDS). Therapy for ARDS and other attempts to improve oxygen uptake were unsuccessful. He was treated empirically for sepsis, herpes simplex virus infection (acyclovir), methicillin-resistant Staphylococcus aureus (vancomycin), and fungal infection (amphotericin B), but his condition deteriorated further, with disseminated intravascular coagulation and liver and renal failure. He died on the 11th day. Laboratory results indicated elevated influenza A antibody on the 8th day, and influenza A was isolated from a tracheal isolate taken on the 9th day. The isolate was sent to the Centers for Disease Control and Prevention and elsewhere, where it was typed as H5N1 avian influenza and named A/Hong Kong/156/97 (H5N1). The child may have contracted the virus while playing with pet ducklings and chickens at his kindergarten. Although the H5N1 virus still has difficulty infecting humans, this case demonstrates the speed and severity of the respiratory and systemic manifestations of avian influenza H5N1 disease.
The changing antigenic nature of influenza ensures a large proportion of immunologically naïve, susceptible people (especially children) in the population (Box 57-3). An influenza outbreak can be readily detected from the increased absenteeism in schools and work and the number of emergency department visits. During the winter, influenza outbreaks occur annually in temperate climates. Fortunately, influenza virus is usually present in a community for only a short time (4 to 6 weeks).
Box 57-3
Epidemiology of Influenza A and B Viruses
Disease/Viral Factors
Virus has a large, enveloped virion that is easily inactivated by dryness, acid, and detergents
Segmented genome facilitates major genetic changes, especially on hemagglutinin and neuraminidase proteins
Influenza A infects many vertebrate species, including other mammals and birds
Co-infection with animal and human strains of influenza can generate very different virus strains by genetic reassortment
Influenza infection is spread readily via small airborne droplets expelled during talking, breathing, and coughing. Low humidity and cool temperatures stabilize the virus, and close proximity during the winter months promotes spread of the virus. The virus can also survive on countertops for as long as a day.
The most susceptible population is children, and school-age children are most likely to spread the infection. Contagion precedes symptoms and lasts for a long time, especially in children. Children, immunosuppressed people (including pregnant women), the elderly, and people with heart and lung ailments (including smokers) are at highest risk for more serious disease, pneumonia, or other complications of infection. More than 90% of mortalities occur in patients who are older than 65 years.
Extensive surveillance of influenza A and B outbreaks is conducted to identify new strains that should be incorporated into new vaccines. The prevalence of a particular strain of influenza A or B virus changes each year and reflects the particular immunologic naïveté of the population at that time. Surveillance also extends into the animal populations because of the possible presence of recombinant animal influenza A strains that can cause human pandemics.
Clinical Syndromes (Box 57-4)
Depending on the degree of immunity to the infecting strain of virus and other factors, infection may range from asymptomatic to severe. Patients with underlying cardiorespiratory disease, people with immune deficiency (even that associated with pregnancy), the elderly, and smokers are more prone to have a severe case.
Box 57-4
Clinical Summary
Influenza A: A 70-year-old woman has rapid onset of fever with headache, myalgia, sore throat, and nonproductive cough. The disease progresses to pneumonia with bacterial involvement. There is no history of recent immunization with influenza A vaccine. Her husband is treated with amantadine or a neuraminidase inhibitor.
After an incubation period of 1 to 4 days, the “flu syndrome” begins with a brief prodrome of malaise and headache lasting a few hours. The prodrome is followed by the abrupt onset of fever, chills, severe myalgias, loss of appetite, weakness and fatigue, sore throat, and usually a nonproductive cough. The fever persists for 3 to 8 days, and unless a complication occurs, recovery is complete within 7 to 10 days. Influenza in young children (younger than 3 years) resembles other severe respiratory tract infections, causing bronchiolitis, croup, otitis media, vomiting, and abdominal pain, accompanied rarely by febrile convulsions (Table 57-3). Complications of influenza include bacterial pneumonia, myositis, and Reye syndrome. The central nervous system can also be involved. Influenza B disease is similar to influenza A disease.
Table 57-3 Diseases Associated with Influenza Virus Infection
| Disorder | Symptoms |
|---|---|
| Acute influenza infection in adults | Rapid onset of fever, malaise, myalgia, sore throat, and nonproductive cough |
| Acute influenza infection in children | Acute disease similar to that in adults but with higher fever, gastrointestinal tract symptoms (abdominal pain, vomiting), otitis media, myositis, and more frequent croup |
| Complications of influenza virus infection |
Influenza may directly cause pneumonia, but it more commonly promotes a secondary bacterial superinfection that leads to bronchitis or pneumonia. The tissue damage caused by progressive influenza virus infection of alveoli can be extensive, leading to hypoxia and bilateral pneumonia. Secondary bacterial infection usually involves Streptococcus pneumoniae, Haemophilus influenzae, or Staphylococcus aureus. In these infections, sputum usually is produced and becomes purulent.
Although the infection generally is limited to the lung, some strains of influenza can spread to other sites in certain people. For example, myositis (inflammation of muscle) may occur in children. Encephalopathy, although rare, may accompany an acute influenza illness and can be fatal. Postinfluenza encephalitis occurs 2 to 3 weeks after recovery from influenza. These diseases are thought to be autoimmune diseases triggered by influenza.
Reye syndrome is an acute encephalitis that affects children and occurs after a variety of acute febrile viral infections, including varicella and influenza B and A diseases. Children given salicylates (aspirin) are at increased risk for this syndrome. In addition to encephalopathy, hepatic dysfunction is present. The mortality rate may be as high as 40%.
Laboratory Diagnosis
The diagnosis of influenza is usually based on the characteristic symptoms, the season, and the presence of the virus in the community. Laboratory methods that distinguish influenza from other respiratory viruses and identify its type and strain confirm the diagnosis (Table 57-4).
Table 57-4 Laboratory Diagnosis of Influenza Virus Infection
| Test | Detects |
|---|---|
| Cell culture in primary monkey kidney or Madin-Darby canine kidney cells | Presence of virus; limited cytopathologic effects |
| Hemadsorption to infected cells | Presence of hemagglutinin protein on cell surface |
| Hemagglutination | Presence of virus in secretions |
| Hemagglutination inhibition | Type and strain of influenza virus or specificity of antibody |
| Antibody inhibition of hemadsorption | Identification of influenza type and strain |
| Immunofluorescence, ELISA | Influenza virus antigens in respiratory secretions or tissue culture |
| Serology: hemagglutination inhibition, hemadsorption inhibition, ELISA, immunofluorescence, complement fixation | Seroepidemiology |
| Genomics: RT-PCR | Identification of influenza type and strain |
ELISA, Enzyme-linked immunosorbent assay; RT-PCR, reverse transcriptase polymerase chain reaction.
Influenza viruses are obtained from respiratory secretions taken early in the illness. The virus is generally isolated in primary monkey kidney cell cultures or the Madin-Darby canine kidney cell line. Nonspecific cytopathologic effects are often difficult to distinguish but may be noted within as few as 2 days (average, 4 days). Before the cytopathologic effects develop, the addition of guinea pig erythrocytes may reveal hemadsorption (the adherence of these erythrocytes to HA-expressing infected cells) (see Chapter 47, Figure 47-5). The addition of influenza virus–containing media to erythrocytes promotes the formation of a gel-like aggregate resulting from hemagglutination. Hemagglutination and hemadsorption are not specific to influenza viruses because parainfluenza and other viruses also exhibit these properties.
More rapid techniques detect and identify the influenza genome or antigens of the virus. Rapid antigen assays (less than 30 minutes) can detect and distinguish influenza A and B. Reverse transcriptase polymerase chain reaction (RT-PCR), using generic influenza primers, can be used to detect and distinguish influenza A and B, and more specific primers can be used to distinguish the different strains, such as H5N1. Enzyme immunoassay or immunofluorescence can be used to detect viral antigen in exfoliated cells, respiratory secretions, or cell culture and are more sensitive assays. Immunofluorescence or inhibition of hemadsorption or hemagglutination (hemagglutination inhibition) with specific antibody (see Chapter 47, Figure 47-6) can also detect and distinguish different influenza strains. Laboratory studies are primarily used for epidemiologic purposes.
Treatment, Prevention, and Control
Hundreds of millions of dollars are spent on acetaminophen, antihistamines, and similar drugs to relieve the symptoms of influenza. The antiviral drug amantadine and its analogue rimantadine inhibit an uncoating step of the influenza A virus but do not affect the influenza B and C viruses. The target for their action is the M2 protein. Zanamivir and oseltamivir inhibit both influenza A and B as enzyme inhibitors of neuraminidase. Without neuraminidase, the hemagglutinin of the virus binds to sialic acid on other glycoproteins and viral particles to form clumps, thereby preventing virus release. Zanamivir is inhaled, whereas oseltamivir is taken orally as a pill. These drugs are effective for prophylaxis and for treatment during the first 24 to 48 hours after the onset of influenza A illness. Treatment cannot prevent the later host-induced immunopathogenic stages of the disease. Naturally resistant or mutant strains are selected when antiviral prophylaxis is used.
The airborne spread of influenza is almost impossible to limit. However, the best way to control the virus is through immunization. Natural immunization, which results from prior exposure, is protective for long periods. A killed-virus vaccine representing the “strains of the year” and antiviral drug prophylaxis can also prevent infection.
The influenza vaccine is a mixture of extracts or purified HA and NA proteins from three different strains of virus. The vaccines are prepared from virus grown in embryonated eggs and then chemically inactivated. Killed (formalin-inactivated) virion preparations have also been used. Vaccines grown in tissue culture cells or by genetic engineering are being developed. Ideally, the vaccine incorporates antigens of the A and B influenza strains that will be prevalent in the community during the upcoming winter. For instance, the trivalent influenza vaccine for the Northern Hemisphere for the 2010 to 2011 season included an A/California/7/2009 (H1N1)-like virus; an A/Perth/16/2009 (H3N2)-like virus; and a B/Brisbane/60/2008-like virus. The 2009 pandemic H1N1 strain is included in this vaccine. Vaccination is routinely recommended for all individuals and especially persons older than 50 years, health care workers, pregnant women who will be in their second or third trimester during flu season, people living in a nursing home, people with chronic pulmonary heart disease, and others at high risk. As of 2008, all children aged 5 to 18 years should also be vaccinated. Persons with allergies to eggs should not get the vaccine.
A live vaccine is also available for administration as a nasal spray instead of a “flu shot.” The trivalent vaccine consists of reassortants for the HA and NA gene segments of the desired influenza strains, with a master donor virus that is cold adapted to optimum growth at 25° C. This vaccine will elicit a more natural protection, including cell-mediated, serum antibody and mucosal-secretory IgA antibody. Currently, the vaccine is recommended for people ages 2 to 50 years.
In late December, a 22-year-old man suddenly experienced headache, myalgia, malaise, dry cough, and fever. He basically felt lousy. After a couple of days, he had a sore throat, his cough had worsened, he started to feel nauseated, and he began vomiting. Several of his family members had experienced similar symptoms during the previous 2 weeks.
1. In addition to influenza, what other agents could cause similar symptoms (differential diagnosis)?
2. How would the diagnosis of influenza be confirmed?
3. Amantadine is effective against influenza. What is its mechanism of action? Will it be effective for this patient? For uninfected family members or contacts?
4. When was the patient contagious, and how was the virus transmitted?
5. What family members were at greatest risk for serious disease and why?
6. Why is influenza so difficult to control, even when there is a national vaccination program?
1. These symptoms can be caused by the parainfluenza, metapneumovirus, or respiratory syncytial paramyxoviruses or by adenovirus.
2. The diagnosis can be confirmed by detection of hemagglutinating activity in nasal washings and confirmation by hemagglutination inhibition with virus-specific antibody or RT-PCR analysis for the influenza genome.
3. Amantadine and rimantidine inhibit the uncoating of the virus by blocking the M2 viral protein–derived channel that is inserted into the endosomal uptake vesicle. This prevents the flow of protons through the channel and the subsequent dissociation of the nucleocapsid. Antiinfluenza therapy with amantadine or neuraminidase inhibitors is effective before or within the first 48 hours of infection, when virus replication is occurring but before extensive tissue damage is caused by the virus and the host immune responds to the virus and causes immunopathogenesis. Other individuals can take amantadine as a prophylactic drug.
4. The patient was contagious approximately 1 day before and up to 5 days after the onset of disease signs. The virus is transmitted by the respiratory route.
5. Very young and very old family members are at greatest risk. The young are immunonaïve, and the elderly may be immunodeficient or may not have been exposed and thus lack a response to the current strain of influenza. Older individuals also have difficulty repairing the damage caused by the influenza virus or a bacterial superinfection of the lung (pneumonia) that often accompanies influenza infection.
6. Influenza readily undergoes mutation (drift) to produce new strains of influenza, and influenza A can undergo reassortment of its genome segments with animal (especially avian) influenza viruses to create new viruses (shift). Both shift and drift create new serotypes of virus. The composition of the influenza vaccine is reevaluated on an annual basis in an attempt to out-guess the changes in influenza that Mother Nature delivers.