12 Bacterial Classification, Structure, and Replication
Bacteria, the smallest cells, are visible only with the aid of a microscope. The smallest bacteria (Chlamydia and Rickettsia) are just 0.1 to 0.2 µm in diameter, whereas larger bacteria may be many microns in length. A newly described species is hundreds of times larger than the average bacterial cell and is visible to the naked eye. Most species, however, are approximately 1 µm in diameter and are therefore visible with the use of the light microscope, which has a resolution of 0.2 µm. In comparison, animal and plant cells are much larger, ranging from 7 µm (the diameter of a red blood cell) to several feet (the length of certain nerve cells).
Differences between Eukaryotes and Prokaryotes
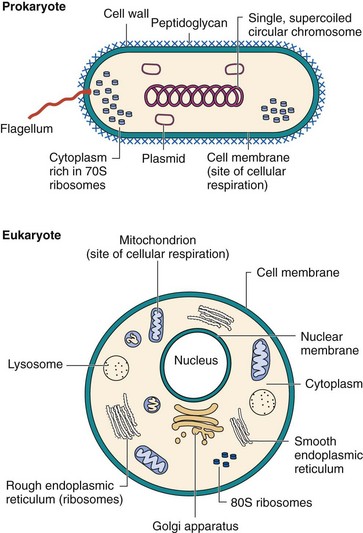
Cells from animals, plants, and fungi are eukaryotes (Greek for “true nucleus”), whereas bacteria, archae, and blue-green algae belong to the prokaryotes (Greek for “primitive nucleus”). The archae (archaebacteria) resemble bacteria in most ways but represent a domain unique from bacteria and eukaryotes. In addition to lacking a nucleus and other organelles, the bacterial chromosome differs from the human chromosome in several ways. The chromosome of a typical bacterium, such as Escherichia coli, is a single, double-stranded, circular molecule of deoxyribonucleic acid (DNA) containing approximately 5 million base pairs (or 5000 kilobase [kb] pairs), an approximate length of 1.3 mm (i.e., nearly 1000 times the diameter of the cell). The smallest bacterial chromosomes (from mycoplasmas) are approximately one fourth of this size. In comparison, humans have two copies of 23 chromosomes, which represent 2.9 × 109 base pairs 990 mm in length. Bacteria use a smaller ribosome, the 70S ribosome, and in most bacteria, a meshlike peptidoglycan cell wall surrounds the membranes to protect them against the environment. Bacteria can survive, and in some cases, grow in hostile environments in which the osmotic pressure outside the cell is so low that most eukaryotic cells would lyse, at temperature extremes (both hot and cold), with dryness, and with very dilute and diverse energy sources. Bacteria have evolved their structures and functions to adapt to these conditions. These and other distinguishing features are depicted in Figure 12-1 and outlined in Table 12-1. Several of these distinctions provide the basis for antimicrobial action.
Table 12-1 Major Characteristics of Eukaryotes and Prokaryotes
| Characteristic | Eukaryote | Prokaryote |
|---|---|---|
| Major groups | Algae, fungi, protozoa, plants, animals | Bacteria |
| Size (approximate) | >5 µm | 0.5-3.0 µm |
| Nuclear Structures | ||
| Nucleus | Classic membrane | No nuclear membrane |
| Chromosomes | Strands of DNA diploid genome | Single, circular DNA haploid genome |
| Cytoplasmic Structures | ||
| Mitochondria | Present | Absent |
| Golgi bodies | Present | Absent |
| Endoplasmic reticulum | Present | Absent |
| Ribosomes (sedimentation coefficient) | 80S (60S + 40S) | 70S (50S + 30S) |
| Cytoplasmic membrane | Contains sterols | Does not contain sterols |
| Cell wall | Present for fungi; otherwise absent | Is a complex structure containing protein, lipids, and peptidoglycans |
| Reproduction | Sexual and asexual | Asexual (binary fission) |
| Movement | Complex flagellum, if present | Simple flagellum, if present |
| Respiration | Via mitochondria | Via cytoplasmic membrane |
Modified from Holt S. In Slots J, Taubman M, editors: Contemporary oral microbiology and immunology, St Louis, 1992, Mosby.
Bacterial Classification
Bacteria can be classified by their macroscopic and microscopic appearance, by characteristic growth and metabolic properties, by their antigenicity, and finally by their genotype.
Macroscopic and Microscopic Distinction
The initial distinction between bacteria can be made by growth characteristics on different nutrient and selective media. The bacteria grow in colonies; each colony is like a city of as many as a million or more organisms. The sum of their characteristics provides the colony with distinguishing characteristics, such as color, size, shape, and smell. The ability to resist certain antibiotics, ferment specific sugars (e.g., lactose, to distinguish E. coli from Salmonella), to lyse erythrocytes (hemolytic properties), or to hydrolyze lipids (e.g., clostridial lipase) can also be determined using the appropriate growth media.
The microscopic appearance, including the size, shape, and configuration of the organisms (cocci, rods, curved, or spiral) and their ability to retain the Gram stain (gram-positive or gram-negative) are the primary means for distinguishing the bacteria. A spherical bacterium, such as Staphylococcus, is a coccus; a rod-shaped bacterium, such as E. coli, is a bacillus; and the snakelike treponeme is a spirillum. In addition, Nocardia and Actinomyces species have branched filamentous appearances similar to those of fungi. Some bacteria form aggregates, such as the grapelike clusters of Staphylococcus aureus or the diplococcus (two cells together) observed in Streptococcus or Neisseria species.
Gram stain is a rapid, powerful, easy test that allows clinicians to distinguish between the two major classes of bacteria, develop an initial diagnosis, and initiate therapy based on inherent differences in the bacteria (Figure 12-2). Bacteria are heat fixed or otherwise dried onto a slide, stained with crystal violet (Figure 12-3), a stain that is precipitated with iodine, and then the unbound and excess stain is removed by washing with the acetone-based decolorizer and water. A red counterstain, safranin, is added to stain any decolorized cells. This process takes less than 10 minutes.
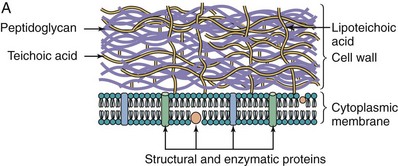
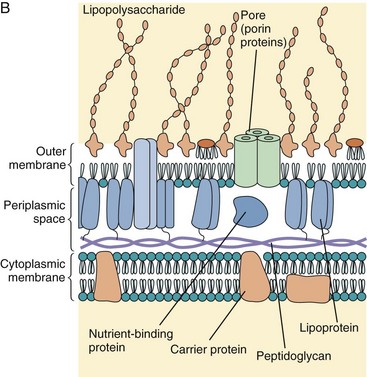
Figure 12-2 Comparison of the gram-positive and gram-negative bacterial cell walls. A, A gram-positive bacterium has a thick peptidoglycan layer that contains teichoic and lipoteichoic acids. B, A gram-negative bacterium has a thin peptidoglycan layer and an outer membrane that contains lipopolysaccharide, phospholipids, and proteins. The periplasmic space between the cytoplasmic and outer membranes contains transport, degradative, and cell wall synthetic proteins. The outer membrane is joined to the cytoplasmic membrane at adhesion points and is attached to the peptidoglycan by lipoprotein links.
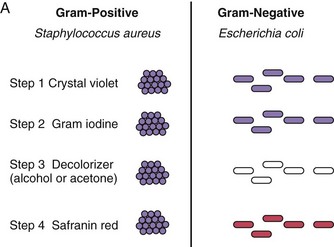
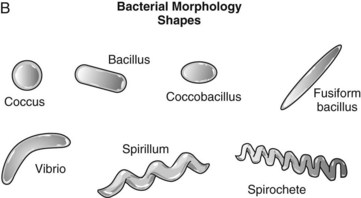
Figure 12-3 Gram-stain morphology of bacteria. A, The crystal violet of Gram stain is precipitated by Gram iodine and is trapped in the thick peptidoglycan layer in gram-positive bacteria. The decolorizer disperses the gram-negative outer membrane and washes the crystal violet from the thin layer of peptidoglycan. Gram-negative bacteria are visualized by the red counterstain. B, Bacterial morphologies.
For gram-positive bacteria, which turn purple, the stain gets trapped in a thick, cross-linked, meshlike structure, the peptidoglycan layer, which surrounds the cell. Gram-negative bacteria have a thin peptidoglycan layer that does not retain the crystal violet stain; so the cells must be counterstained with safranin and turned red (Figure 12-4). A mnemonic device that may help is “P-PURPLE-POSITIVE.”
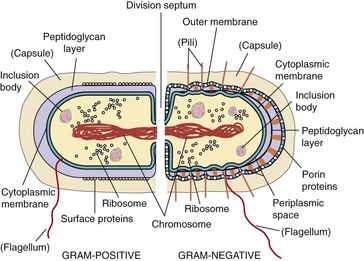
Figure 12-4 Gram-positive and gram-negative bacteria. A gram-positive bacterium has a thick layer of peptidoglycan (filling the purple space) (left). A gram-negative bacterium has a thin peptidoglycan layer (single black line) and an outer membrane (right). Structures in parentheses are not found in all bacteria. Upon cell division, the membrane and peptidoglycan grow toward each other to form a division septum to separate the daughter cells.
Due to degradation of the peptidoglycan, Gram staining is not a dependable test for bacteria that are starved (e.g., old or stationary-phase cultures) or treated with antibiotics. Bacteria that cannot be classified by Gram staining include mycobacteria, which have a waxy outer shell and are distinguished with the acid-fast stain, and mycoplasmas, which have no peptidoglycan.
Metabolic, Antigenic, and Genetic Distinction
The next level of classification is based on the metabolic signature of the bacteria, including requirement for anaerobic or aerobic environments, requirement for specific nutrients (e.g., ability to ferment specific carbohydrates or use different compounds as a source of carbon for growth), and production of characteristic metabolic products (acid, alcohols) and specific enzymes (e.g., staphylococcal catalase). Automated procedures for distinguishing enteric and other bacteria have been developed; they analyze the growth in different media and their microbial products and provide a numerical biotype for each of the bacteria.
A particular strain of bacteria can be distinguished using antibodies to detect characteristic antigens on the bacteria (serotyping). These serologic tests can also be used to identify organisms that are difficult (Treponema pallidum, the organism responsible for syphilis) or too dangerous (e.g., Francisella, the organism that causes tularemia) to grow in the laboratory, are associated with specific disease syndromes (e.g., E. coli serotype O157:H7, responsible for hemorrhagic colitis), or need to be identified rapidly (e.g., Streptococcus pyogenes, responsible for streptococcal pharyngitis). Serotyping is also used to subdivide bacteria below the species level for epidemiologic purposes.
The most precise method for classifying bacteria is by analysis of their genetic material. New methods distinguish bacteria by detection of specific characteristic DNA sequences. These techniques include DNA hybridization, polymerase chain reaction (PCR) amplification, and related techniques described in Chapter 5. These genetic techniques do not require living or growing bacteria and can be used for rapid detection and identification of slow-growing organisms, such as mycobacteria and fungi, or analysis of pathology samples of even very virulent bacteria. The technology is now available for rapid analysis of the nucleic acid sequence of specific segments or the entire bacterial chromosome. The most common application of this technique is analysis of sequences of ribosomal DNA to detect the highly conserved sequences that identify a family or genus and the highly variable sequences that distinguish a species or subspecies. It has also been used to define the evolutionary relationship among organisms and to identify organisms that are difficult or impossible to grow. Various other methods that have been used, primarily to classify organisms at the subspecies level for epidemiologic investigations, include: plasmid analysis, ribotyping, and analysis of chromosomal DNA fragments. In recent years the technical aspects of these methods have been simplified to the point that most clinical laboratories use variations of these methods in their day-to-day practice.
Bacterial Structure
Cytoplasmic Structures
The cytoplasm of the bacterial cell contains the DNA chromosome, messenger RNA (mRNA), ribosomes, proteins, and metabolites (see Figure 12-4). Unlike eukaryotes, the bacterial chromosome is a single, double-stranded circle that is contained not in a nucleus, but in a discrete area known as the nucleoid. Histones are not present to maintain the conformation of the DNA, and the DNA does not form nucleosomes. Plasmids, which are smaller, circular, extrachromosomal DNAs, may also be present. Plasmids, although not usually essential for cellular survival, often provide a selective advantage: many confer resistance to one or more antibiotics.
The lack of a nuclear membrane simplifies the requirements and control mechanisms for the synthesis of proteins. Without a nuclear membrane, transcription and translation are coupled; in other words, ribosomes can bind to the mRNA, and protein can be made as the mRNA is being synthesized and still attached to the DNA.
The bacterial ribosome consists of 30S + 50S subunits, forming a 70S ribosome. This is unlike the eukaryotic 80S (40S + 60S) ribosome. The proteins and RNA of the bacterial ribosome are significantly different from those of eukaryotic ribosomes and are major targets for antibacterial drugs.
The cytoplasmic membrane has a lipid bilayer structure similar to the structure of the eukaryotic membranes, but it contains no steroids (e.g., cholesterol); mycoplasmas are the exception to this rule. The cytoplasmic membrane is responsible for many of the functions attributable to organelles in eukaryotes. These tasks include electron transport and energy production, which are normally achieved in the mitochondria. In addition, the membrane contains transport proteins that allow the uptake of metabolites and the release of other substances, ion pumps to maintain a membrane potential, and enzymes. The inside of the membrane is lined with actin-like protein filaments, which help determine the shape of the bacteria and the site of septum formation for cell division. These filaments determine the spiral shape of the treponemes.
Cell Wall
The structure (Table 12-2), components, and functions (Table 12-3) of the cell wall distinguish gram-positive from gram-negative bacteria. Cell wall components are also unique to bacteria, and their repetitive structures bind to Toll-like receptors on human cells to elicit innate protective responses. The important differences in membrane characteristics are outlined in Table 12-4. Rigid peptidoglycan (murein) layers surround the cytoplasmic membranes of most prokaryotes. The exceptions are Archaea organisms (which contain pseudoglycans or pseudomureins related to peptidoglycan) and mycoplasmas and chlamydia (which have no peptidoglycan). Because the peptidoglycan provides rigidity, it also helps to determine the shape of the particular bacterial cell. Gram-negative bacteria are also surrounded by outer membranes.
Table 12-2 Bacterial Membrane Structures
| Structure | Chemical Constituents | Functions |
|---|---|---|
| Plasma membrane | Phospholipids, proteins, and enzymes | Containment, generation of energy, membrane potential, and transport |
| Cell Wall | ||
| Gram-Positive Bacteria | ||
| Peptidoglycan | Glycan chains of GlcNAc and MurNAc cross-linked by peptide bridge | Cell shape and structure; protection from environment and complement killing |
| Teichoic acid | Polyribitol phosphate or glycerol phosphate cross-linked to peptidoglycan | Strengthens cell wall; calcium ion sequestration; activator of innate host protections |
| Lipoteichoic acid | Lipid-linked teichoic acid | |
| Gram-Negative Bacteria | ||
| Peptidoglycan | Thinner version of that found in gram-positive bacteria | Cell shape and structure |
| Periplasmic space | Enzymes involved in transport, degradation, and synthesis | |
| Outer membrane | Cell structure; protection from host environment | |
| Proteins | Porin channel | Permeation of small, hydrophilic molecules; restricts some antibiotics |
| Secretory devices (types I, II, III, IV) | Penetrates and delivers proteins across membranes, including virulence factors | |
| Lipoprotein | Outer membrane link to peptidoglycan | |
| LPS | Lipid A, core polysaccharide, O antigen | Outer membrane structure; potent activator of innate host responses |
| Phospholipids | With saturated fatty acids | |
| Other Structures | ||
| Capsule | Polysaccharides (disaccharides and trisaccharides) and polypeptides | Antiphagocytic |
| Biofilm | Polysaccharides | Protection of colony from environment, antimicrobials and host response |
| Pili | Pilin, adhesins | Adherance, sex pili |
| Flagellum | Motor proteins, flagellin | Movement, chemotaxis |
| Proteins | M protein of streptococci (for example) | Various |
GlcNAc, N-Acetylglucosamine; LPS, lipopolysaccharide; MurNAc, N-acetylmuramic acid.
Table 12-3 Functions of the Bacterial Envelope
| Function | Component |
|---|---|
| Structure | |
| Rigidity | All |
| Packaging of internal contents | All |
| Bacterial Functions | |
| Permeability barrier | Outer membrane or plasma membrane |
| Metabolic uptake | Membranes and periplasmic transport proteins, porins, permeases |
| Energy production | Plasma membrane |
| Motility | Flagella |
| Mating | Pili |
| Host Interaction | |
| Adhesion to host cells | Pili, proteins, teichoic acid |
| Immune recognition by host | All outer structures and peptidoglycan |
| Escape from host immune protections | |
| Antibody | M protein |
| Phagocytosis | Capsule |
| Complement | Gram-positive peptidoglycan |
| Medical Relevance | |
| Antibiotic targets | Peptidoglycan synthesis, outer membrane |
| Antibiotic resistance | Outer membrane barrier |
Table 12-4 Membrane Characteristics of Gram-Positive and Gram-Negative Bacteria
| Characteristic | Gram-Positive | Gram-Negative |
|---|---|---|
| Outer membrane | − | + |
| Cell wall | Thicker | Thinner |
| Lipopolysaccharide | − | + |
| Endotoxin | − | + |
| Teichoic acid | Often present | − |
| Sporulation | Some strains | − |
| Capsule | Sometimes present | Sometimes present |
| Lysozyme | Sensitive | Resistant |
| Antibacterial activity of penicillin | More susceptible | More resistant |
| Exotoxin production | Some strains | Some strains |
Gram-Positive Bacteria
A gram-positive bacterium has a thick, multilayered cell wall consisting mainly of peptidoglycan (150 to 500 Å) surrounding the cytoplasmic membrane (Figure 12-5). The peptidoglycan is a meshlike exoskeleton similar in function to the exoskeleton of an insect. Unlike the exoskeleton of the insect, however, the peptidoglycan of the cell is sufficiently porous to allow diffusion of metabolites to the plasma membrane. A new model for peptidoglycan suggests that the glycan extends out from the plasma membrane like bristles that are cross-linked with short peptide chains. The peptidoglycan is essential for the structure, for replication, and for survival in the normally hostile conditions in which bacteria grow.
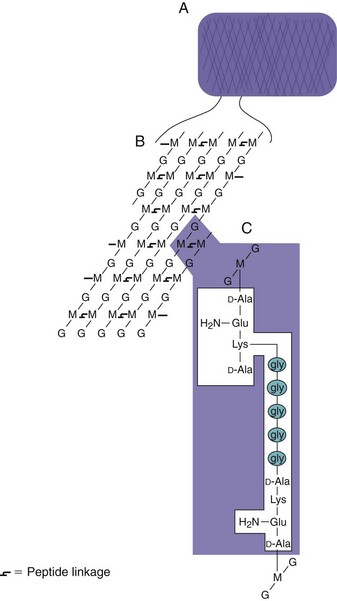
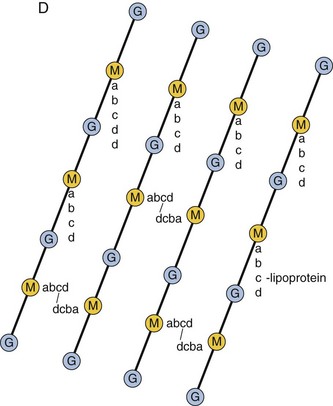
Figure 12-5 General structure of the peptidoglycan component of the cell wall. A, The peptidoglycan forms a meshlike layer around the cell. B, The peptidoglycan mesh consists of a polysaccharide polymer that is cross-linked by peptide bonds. C, Peptides are cross-linked through a peptide bond between the terminal D-alanine (D-Ala) from one chain and a lysine (Lys) (or another diamino amino acid) from the other chain. A pentaglycine bridge (gly5) expands the cross-link in Staphylococcus aureus (as shown). D, Representation of the Escherichia coli peptidoglycan structure. Diaminopimelic acid, the diamino amino acid in the third position of the peptide, is directly linked to the terminal alanine of another chain to cross-link the peptidoglycan. Lipoprotein anchors the outer membrane to the peptidoglycan. G, N-Acetylglucosamine; Glu, D-glutamic acid; gly, glycine; M, N-acetylmuramic acid.
(A to C, Modified from Talaro K, Talaro A: Foundations in microbiology, ed 2, Dubuque, Iowa, 1996, William C Brown. D, Modified from Joklik KJ, et al: Zinsser microbiology, Norwalk, Conn, 1988, Appleton & Lange.)
The peptidoglycan can be degraded by treatment with lysozyme. Lysozyme is an enzyme in human tears and mucus, but is also produced by bacteria and other organisms. Lysozyme cleaves the glycan backbone of the peptidoglycan. Without the peptidoglycan, the bacteria succumb to the large osmotic pressure differences across the cytoplasmic membrane and lyse. Removal of the cell wall produces a protoplast that lyses, unless it is osmotically stabilized.
The gram-positive cell wall may also include other components such as proteins, teichoic and lipoteichoic acids, and complex polysaccharides (usually called C polysaccharides). The M protein of streptococci and R protein of staphylococci associate with the peptidoglycan. Teichoic acids are water-soluble, anionic polymers of polyol phosphates, which are covalently linked to the peptidoglycan and essential to cell viability. Lipoteichoic acids have a fatty acid and are anchored in the cytoplasmic membrane. These molecules are common surface antigens that distinguish bacterial serotypes and promote attachment to other bacteria and to specific receptors on mammalian cell surfaces (adherence). Teichoic acids are important factors in virulence. Lipoteichoic acids are shed into the media and the host and although weaker, they can initiate innate protective host responses similar to endotoxin.
Gram-Negative Bacteria
Gram-negative cell walls are more complex than gram-positive cell walls, both structurally and chemically (see Figure 12-2). Structurally, a gram-negative cell wall contains two layers external to the cytoplasmic membrane. Immediately external to the cytoplasmic membrane is a thin peptidoglycan layer, which accounts for only 5% to 10% of the gram-negative cell wall by weight. There are no teichoic or lipoteichoic acids in the gram-negative cell wall. External to the peptidoglycan layer is the outer membrane, which is unique to gram-negative bacteria. The area between the external surface of the cytoplasmic membrane and the internal surface of the outer membrane is referred to as the periplasmic space. This space is actually a compartment containing components of transport systems for iron, proteins, sugars and other metabolites, and a variety of hydrolytic enzymes that are important to the cell for the breakdown of large macromolecules for metabolism. These enzymes typically include proteases, phosphatases, lipases, nucleases, and carbohydrate-degrading enzymes. In the case of pathogenic gram-negative species, many of the lytic virulence factors, such as collagenases, hyaluronidases, proteases, and β-lactamase, are in the periplasmic space.
The gram-negative cell wall is also traversed by different transport systems, including the type I, II, III, IV, and V secretion devices. Transport systems provide mechanisms for the uptake and release of different metabolites and other compounds. Production of the secretion devices may be induced during infection and contribute to the virulence of the microbe by transporting molecules that facilitate bacterial adhesion or intracellular growth. The type III secretion device is a major virulence factor for some bacteria, with a complex structure that traverses both the inner and outer membranes and can act as a syringe to inject proteins into other cells.
As mentioned previously, outer membranes (see Figure 12-2) are unique to gram-negative bacteria. The outer membrane is like a stiff canvas sack around the bacteria. The outer membrane maintains the bacterial structure and is a permeability barrier to large molecules (e.g., proteins such as lysozyme) and hydrophobic molecules (e.g., some antimicrobials). It also provides protection from adverse environmental conditions, such as the digestive system of the host (important for Enterobacteriaceae organisms). The outer membrane has an asymmetric bilayer structure that differs from any other biologic membrane in the structure of the outer leaflet of the membrane. The inner leaflet contains phospholipids normally found in bacterial membranes. However, the outer leaflet is composed primarily of lipopolysaccharide (LPS). Except for those LPS molecules in the process of synthesis, the outer leaflet of the outer membrane is the only location where LPS molecules are found.
LPS is also called endotoxin, a powerful stimulator of innate and immune responses. LPS is shed from the bacteria into the media and host. LPS activates B cells and induces macrophage, dendritic, and other cells to release interleukin-1, interleukin-6, tumor necrosis factor, and other factors. LPS induces fever and can cause shock. The Shwartzman reaction (disseminated intravascular coagulation) follows the release of large amounts of endotoxin into the bloodstream. Neisseria bacteria shed large amounts of a related molecule, lipooligosaccharide (LOS), resulting in fever and severe symptoms.
The variety of proteins found in gram-negative outer membranes is limited, but several of the proteins are present in high concentration, resulting in a total protein content that is higher than that of the cytoplasmic membrane. Many of the proteins traverse the entire lipid bilayer and are thus transmembrane proteins. A group of these proteins is known as porins because they form pores that allow the diffusion of hydrophilic molecules less than 700 Da in mass through the membrane. The porin channel allows passage of metabolites and small hydrophilic antimicrobials. The outer membrane also contains structural proteins, receptor molecules for bacteriophages, and other ligands and components of transport and secretory systems.
The outer membrane is connected to the cytoplasmic membrane at adhesion sites and is tied to the peptidoglycan by lipoprotein. The lipoprotein is covalently attached to the peptidoglycan and is anchored in the outer membrane. The adhesion sites provide a membranous route for the delivery of newly synthesized outer membrane components to the outer membrane.
The outer membrane is held together by divalent cation (Mg+2 and Ca+2) linkages between phosphates on LPS molecules and hydrophobic interactions between the LPS and proteins. These interactions produce a stiff, strong membrane that can be disrupted by antibiotics (e.g., polymyxin) or by the removal of Mg and Ca ions (chelation with ethylenediaminetetraacetic acid [EDTA] or tetracycline). Disruption of the outer membrane weakens the bacteria and allows the permeability of large, hydrophobic molecules. The addition of lysozyme to cells with a disrupted outer membrane produces spheroplasts, which, like protoplasts, are osmotically sensitive.
External Structures
Some bacteria (gram-positive or gram-negative) are closely surrounded by loose polysaccharide or protein layers called capsules. In cases in which it is loosely adherent and nonuniform in density or thickness, the material is referred to as a slime layer. The capsules and slime layers are also called the glycocalyx. Bacillus anthracis, the exception to this rule, produces a polypeptide capsule. The capsule is hard to see in a microscope, but its space can be visualized by the exclusion of India ink particles.
Capsules and slimes are unnecessary for the growth of bacteria, but are very important for survival in the host. The capsule is poorly antigenic and antiphagocytic and is a major virulence factor (e.g., Streptococcus pneumoniae). The capsule can also act as a barrier to toxic hydrophobic molecules, such as detergents, and can promote adherence to other bacteria or to host tissue surfaces. For Streptococcus mutans, the dextran and levan capsules are the means by which the bacteria attach and stick to the tooth enamel. Bacterial strains lacking a capsule may arise during growth under laboratory conditions, away from the selective pressures of the host, and are therefore less virulent. Some bacteria (e.g., Pseudomonas aeruginosa, S. aureus) will produce a polysaccharide biofilm when sufficient numbers of bacteria (quorum) are present and under conditions which support growth, which establishes a bacterial community and protects them from antibiotics and host defenses. Another example of a biofilm is tooth plaque produced by S. mutans.
Flagella are ropelike propellers composed of helically coiled protein subunits (flagellin) that are anchored in the bacterial membranes through hook and basal body structures and are driven by membrane potential. Bacterial species may have one or several flagella on their surfaces, and they may be anchored at different parts of the cell. Flagella are composed of an adenosine triphosphate (ATP)-driven protein motor connected to a whiplike propeller made of multiple subunits of flagellin. Flagella provide motility for bacteria, allowing the cell to swim (chemotaxis) toward food and away from poisons. Bacteria approach food by swimming straight and then tumbling in a new direction. The swimming period becomes longer as the concentration of chemoattractant increases. The direction of flagellar spinning determines whether the bacteria swim or tumble. Flagella express antigenic and strain determinants and are a ligand for Toll-like receptor 5 to activate innate host protections.
Fimbriae (pili) (Latin for “fringe”) are hairlike structures on the outside of bacteria; they are composed of protein subunits (pilin). Fimbriae can be morphologically distinguished from flagella because they are smaller in diameter (3 to 8 nm versus 15 to 20 nm) and usually are not coiled in structure. In general, several hundred fimbriae are arranged peritrichously (uniformly) over the entire surface of the bacterial cell. They may be as long as 15 to 20 µm or many times the length of the cell.
Fimbriae promote adherence to other bacteria or to the host (alternative names are adhesins, lectins, evasins, and aggressins). As an adherence factor (adhesin), fimbriae are an important virulence factor for colonization and infection of the urinary tract by E. coli, Neisseria gonorrhoeae, and other bacteria. The tips of the fimbriae may contain proteins (lectins) that bind to specific sugars (e.g., mannose). F pili (sex pili) bind to other bacteria and are a tube for transfer of large segments of bacterial chromosomes between bacteria. These pili are encoded by a plasmid (F).
Bacterial Exceptions
Mycobacteria have a peptidoglycan layer (slightly different structure), which is intertwined with and covalently attached to an arabinogalactan polymer and surrounded by a waxlike lipid coat of mycolic acid (large α-branched β-hydroxy fatty acids), cord factor (glycolipid of trehalose and two mycolic acids), wax D (glycolipid of 15 to 20 mycolic acids and sugar), and sulfolipids (see Figure 25-1). These bacteria are described as staining acid-fast. The coat is responsible for virulence and is antiphagocytic. Corynebacterium and Nocardia organisms also produce mycolic acid lipids. Chlamydia and mycoplasmas have no peptidoglycan cell wall and mycoplasmas incorporate steroids from the host into their membranes.
Structure and Biosynthesis of the Major Components of the Bacterial Cell Wall
The cell wall components are large structures made up of polymers of subunits. This type of structure facilitates their synthesis. Like astronauts building a space station, bacteria face problems assembling their cell walls. Synthesis of the peptidoglycan, LPS, teichoic acid, and capsule occurs on the outside of the bacteria, away from the synthetic machinery and energy sources of the cytoplasm and in an inhospitable environment. For both the space station and the bacteria, prefabricated precursors and subunits of the final structure are assembled in a factory-like setting on the inside, attached to a structure similar to a conveyor belt, brought to the surface, and then attached to the preexisting structure. For bacteria, the molecular conveyor beltlike structure is a large hydrophobic phospholipid called bactoprenol (undecaprenol, [C55 isoprenoid]). The prefabricated precursors must also be activated with high-energy bonds (e.g., phosphates) or other means to power the attachment reactions occurring outside the cell. For gram-negative bacteria, the outer membrane components are delivered through adhesion sites.
Peptidoglycan (Mucopeptide, Murein)
The peptidoglycan is a rigid mesh made up of bristle-like linear polysaccharide chains cross-linked by peptides. The polysaccharide is made up of repeating disaccharides of N-acetylglucosamine (GlcNAc, NAG, G) and N-acetylmuramic acid (MurNAc, NAM, M) (Figure 12-6; see Figure 12-5).
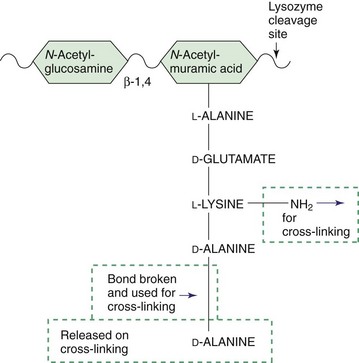
Figure 12-6 Precursor of peptidoglycan. The peptidoglycan is built from prefabricated units that contain a pentapeptide attached to the N-acetylmuramic acid. The pentapeptide contains a terminal D-alanine-D-alanine unit. This dipeptide is required for cross-linking the peptidoglycan and is the basis for the action of β-lactam and vancomycin antibiotics. The β-1,4 disaccharide link cleaved by lysozyme is indicated.
A tetrapeptide is attached to the MurNAc. The peptide is unusual because it contains both D and L amino acids (D amino acids are not normally used in nature) and the peptide is produced enzymatically rather than by a ribosome. The first two amino acids attached to the MurNAc may vary for different organisms.
The diamino amino acids in the third position are essential for the cross-linking of the peptidoglycan chain. Examples of diamino amino acids include lysine and diaminopimelic and diaminobutyric acids. The peptide cross-link is formed between the free amine of the diamino amino acid and the D-alanine in the fourth position of another chain. S. aureus and other gram-positive bacteria use an amino acid bridge (e.g., a glycine5 peptide) between these amino acids to lengthen the cross-link. The precursor form of the peptide has an extra D-alanine, which is released during the cross-linking step.
The peptidoglycan in gram-positive bacteria forms multiple layers and is often cross-linked in three dimensions, providing a very strong, rigid cell wall. In contrast, the peptidoglycan in gram-negative cell walls is usually only one molecule (layer) thick. The number of cross-links and the length of the cross-link determine the rigidity of the peptidoglycan mesh. The site where lysozyme cleaves the glycan of the peptidoglycan is shown in Figure 12-6.
Peptidoglycan Synthesis
Peptidoglycan synthesis occurs in four phases (Figure 12-7). First, the precursors are synthesized and activated inside the cell. Glucosamine is enzymatically converted into MurNAc and then energetically activated by a reaction with uridine triphosphate (UTP) to produce uridine diphosphate-N-acetylmuramic acid (UDP-MurNAc). Next, the UDP-MurNAc-pentapeptide precursor is assembled in a series of enzymatic steps.
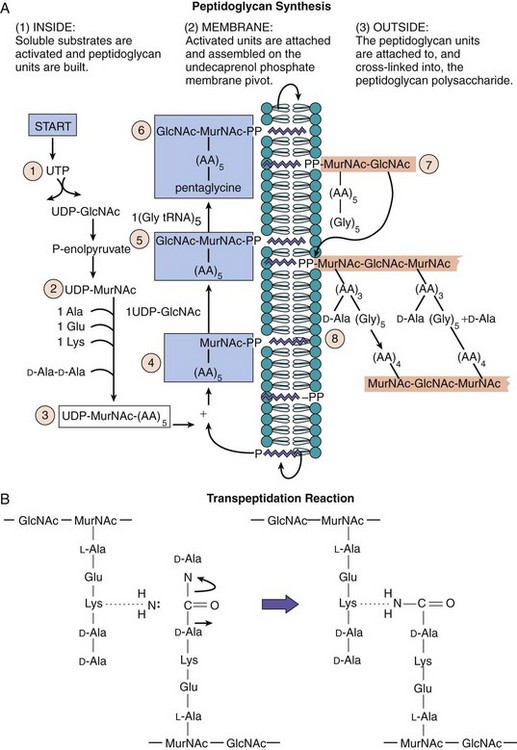
Figure 12-7 Peptidoglycan synthesis. A, Peptidoglycan synthesis occurs in the following four phases: (1) Peptidoglycan is synthesized from prefabricated units constructed and activated for assembly and transport inside the cell. (2) At the membrane the units are assembled onto the undecaprenol phosphate conveyor belt, and fabrication is completed. (3) The unit is translocated to the outside of the cell and (4) the unit is attached to the polysaccharide chain, and the peptide is cross-linked to finish the construction. Staphylococcus aureus utilizes a pentaglycine bridge in the cross-link. Such a construction can be compared with the assembly of a space station of prefabricated units. B, The cross-linking reaction is a transpeptidation. Escherichia coli uses a direct cross-link between D-alanine and lysine. One peptide bond (produced inside the cell) is traded for another (outside the cell) with the release of D-alanine. The enzymes that catalyze the reaction are called D-alanine, D-alanine transpeptidase-carboxypeptidases. These enzymes are the targets of β-lactam antibiotics and are called penicillin-binding proteins. AA5, pentapeptide with D-alanine-D-alanine; AA4, tetrapeptide with terminal D-alanine; AA3, tripeptide; Gly5, glycine pentapeptide; Glu, glutamate; Lys, lysine; MurNAc-PP, N-acetylmuramic acid diphosphate; tRNA, transfer ribonucleic acid; UDP-GlcNAc, uridine diphosphate N-acetylglucosamine; UDP-MurNAc, uridine diphosphate-N-acetylmuramic acid; UTP, uridine triphosphate.
In the second phase, the UDP-MurNAc pentapeptide is attached to the bactoprenol “conveyor belt” in the cytoplasmic membrane through a pyrophosphate link, with the release of uridine monophosphate (UMP). GlcNAc is added to make the disaccharide building block of the peptidoglycan. Some bacteria (e.g., S. aureus) add a pentaglycine or another chain to the diamino amino acid at the third position of the peptide chain to lengthen the cross-link.
In the third phase, the bactoprenol molecule translocates the disaccharide:peptide precursor to the outside of the cell.
In the last phase, the peptidoglycan is extended at the outside surface of the plasma membrane. The GlcNAc-MurNAc disaccharide is attached to a peptidoglycan chain, using the pyrophosphate link between itself and the bactoprenol as energy to drive the reaction by enzymes called transglycosylases. The pyrophosphobactoprenol is converted back to a phosphobactoprenol and recycled. Bacitracin blocks the recycling. The peptide chains from adjacent glycan chains are cross-linked to each other by a peptide bond exchange (transpeptidation) between the free amine of the amino acid in the third position of the pentapeptide (e.g., lysine), or the N-terminus of the attached pentaglycine chain, and the D-alanine at the fourth position of the other peptide chain, releasing the terminal D-alanine of the precursor. This step requires no additional energy because peptide bonds are “traded.”
The cross-linking reaction is catalyzed by membrane-bound transpeptidases. Related enzymes, D-carboxypeptidases, remove unreacted terminal D-alanines to limit the extent of cross-linking. The transpeptidases and carboxypeptidases are called penicillin-binding proteins (PBPs) because they are targets for penicillin and other β-lactam antibiotics. Penicillin and related β-lactam antibiotics resemble the “transition state” conformation of the D-Ala-D-Ala substrate when bound to these enzymes. Vancomycin binds to the D-Ala-D-Ala structure to block these reactions. Different PBPs are used for extending the peptidoglycan, creating a septum for cell division and curving the peptidoglycan mesh (cell shape). Peptidoglycan extension and cross-linking is necessary for cell growth and division.
The peptidoglycan is constantly being synthesized and degraded. Autolysins, such as lysozyme, are important for determining bacterial shape. Inhibition of synthesis or the cross-linking of the peptidoglycan does not stop the autolysins, and their action weakens the mesh and leads to cell lysis and death. New peptidoglycan synthesis does not occur during starvation, which leads to a weakening of the peptidoglycan and a loss in the dependability of the Gram stain.
An understanding of the biosynthesis of peptidoglycan is essential in medicine because these reactions are unique to bacterial cells and hence can be inhibited with little or no adverse effect on host (human) cells. As indicated above, a number of antibacterials target one or more steps in this pathway (see Chapter 17).
Teichoic Acid
Teichoic and lipoteichoic acid are polymers of chemically modified ribose or glycerol connected by phosphates (Figure 12-8). Sugars, choline, or D-alanine may be attached to the hydroxyls of the ribose or glycerol, providing antigenic determinants. These can be distinguished by antibodies and may determine the bacterial serotype. Lipoteichoic acid has a fatty acid and is anchored in the membrane. Teichoic acid is synthesized from building blocks using the bactoprenol in a manner similar to that of peptidoglycan. Teichoic acid and some surface proteins (e.g., protein A from S. aureus) are secreted from the cells and are enzymatically attached to the N-terminus of the peptide of peptidoglycan.
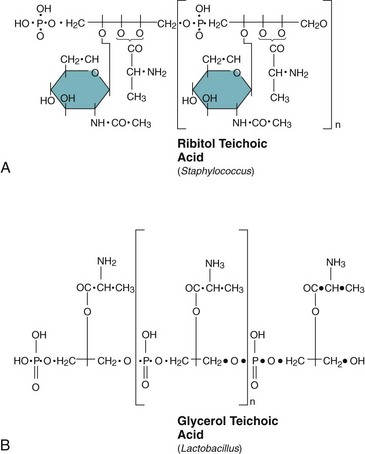
Figure 12-8 Teichoic acid. Teichoic acid is a polymer of chemically modified ribitol (A) or glycerol phosphate (B). The nature of the modification (e.g., sugars, amino acids) can define the serotype of the bacteria. Teichoic acid is covalently attached to the peptidoglycan. Lipoteichoic acid is anchored in the cytoplasmic membrane by a covalently attached fatty acid.
Lipopolysaccharide
LPS (endotoxin) consists of three structural sections: Lipid A, core polysaccharide (rough core), and O antigen (Figure 12-9). Lipid A is a basic component of LPS and is essential for bacterial viability. Lipid A is responsible for the endotoxin activity of LPS. It has a phosphorylated glucosamine disaccharide backbone with fatty acids attached to anchor the structure in the outer membrane. The phosphates connect LPS units into aggregates. One carbohydrate chain is attached to each disaccharide backbone and extends away from the bacteria. The core polysaccharide is a branched polysaccharide of 9 to 12 sugars. Most of the core region is also essential for LPS structure and bacterial viability. The core region contains an unusual sugar, 2-keto-3-deoxy-octanoate (KDO), and is phosphorylated. Divalent cations link the phosphates of the LPS and core to strengthen the outer membrane. The O antigen is attached to the core and extends away from the bacteria. It is a long, linear polysaccharide consisting of 50 to 100 repeating saccharide units of 4 to 7 sugars per unit. LOS, which is present in Neisseria species, lacks the O-antigen portion of LPS and is readily shed from the bacteria. The shorter O antigen makes Neisseria more susceptible to host-mediated complement control.
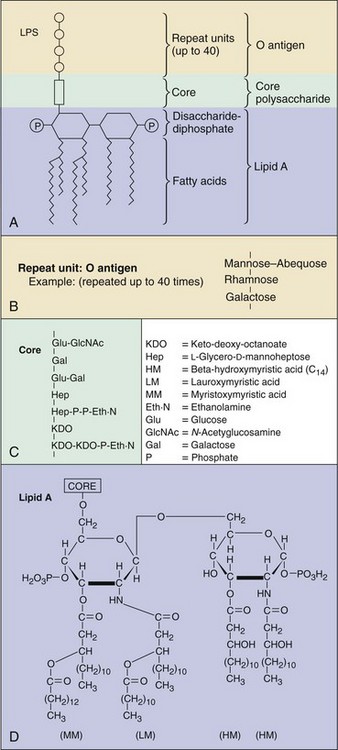
Figure 12-9 The lipopolysaccharide (LPS) of the gram-negative cell envelope. A, Segment of the molecule showing the arrangements of the major constituents. Each LPS molecule has one lipid A and one polysaccharide core unit but many repeats of O antigen. B, Typical O-antigen repeat unit (Salmonella typhimurium). C, Polysaccharide core. D, Structure of lipid A of S. typhimurium.
(Modified from Brooks GF, Butel JS, Ornston LN: Jawetz, Melnick, and Aldenberg’s medical microbiology, ed 19, Norwalk, Conn, 1991, Appleton & Lange.)
LPS structure is used to classify bacteria. The basic structure of lipid A is identical for related bacteria and is similar for all gram-negative Enterobacteriaceae. The core region is the same for a species of bacteria. The O antigen distinguishes serotypes (strains) of a bacterial species. For example, the O157:H7 (O antigen:flagellin) serotype identifies the E. coli agent of hemolytic-uremic syndrome.
The lipid A and core portions are enzymatically synthesized in a sequential manner on the inside surface of the cytoplasmic membrane. The repeat units of the O antigen are assembled on a bactoprenol molecule and then transferred to a growing O-antigen chain. The finished O-antigen chain is transferred to the core lipid A structure. The LPS molecule is translocated through adhesion sites to the outer surface of the outer membrane.
Cell Division
The replication of the bacterial chromosome also triggers the initiation of cell division (Figure 12-10). The production of two daughter bacteria requires the growth and extension of the cell wall components, followed by the production of a septum (cross wall) to divide the daughter bacteria into two cells. The septum consists of two membranes separated by two layers of peptidoglycan. Septum formation is initiated at midcell, at a site defined by protein complexes affixed to a protein filament ring that lines the inside of the cytoplasmic membrane. The septum grows from opposite sides toward the center of the cell, causing cleavage of the daughter cells. This process requires special transpeptidases (PBPs) and other enzymes. For streptococci, the growth zone is located at 180 degrees from each other, producing linear chains of bacteria. In contrast, the growth zone of staphylococci is at 90 degrees. Incomplete cleavage of the septum can cause the bacteria to remain linked, forming chains (e.g., streptococci) or clusters (e.g., staphylococci).
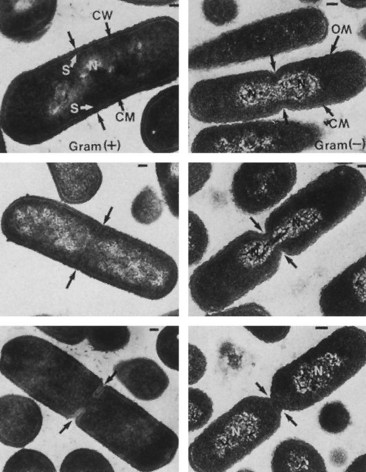
Figure 12-10 Electron photomicrographs of gram-positive cell division (Bacillus subtilis) (left) and gram-negative cell division (Escherichia coli) (right). Progression in cell division from top to bottom. CM, Cytoplasmic membrane; CW, cell wall; N, nucleoid; OM, outer membrane; S, septum. Bar = 0.2 µm.
(From Slots J, Taubman MA: Contemporary oral biology and immunology, St Louis, 1992, Mosby.)
Spores
Some gram-positive, but never gram-negative, bacteria, such as members of the genera Bacillus (e.g., Bacillus anthracis) and Clostridium (e.g., Clostridium tetani or botulinum) (soil bacteria), are spore formers. Under harsh environmental conditions, such as the loss of a nutritional requirement, these bacteria can convert from a vegetative state to a dormant state, or spore. The location of the spore within a cell is a characteristic of the bacteria and can assist in identification of the bacterium.
The spore is a dehydrated, multishelled structure that protects and allows the bacteria to exist in “suspended animation” (Figure 12-11). It contains a complete copy of the chromosome, the bare minimum concentrations of essential proteins and ribosomes, and a high concentration of calcium bound to dipicolinic acid. The spore has an inner membrane, two peptidoglycan layers, and an outer keratin-like protein coat. The spore looks refractile (bright) in the microscope. The structure of the spore protects the genomic DNA from intense heat, radiation, and attack by most enzymes and chemical agents. In fact, bacterial spores are so resistant to environmental factors that they can exist for centuries as viable spores. Spores are also difficult to decontaminate with standard disinfectants.
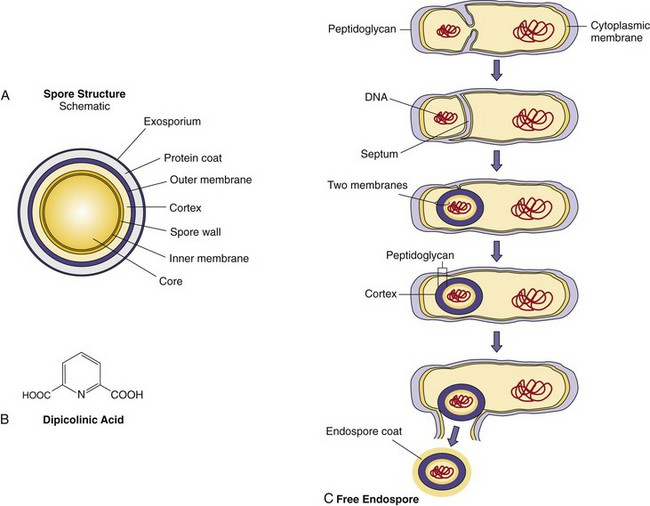
Figure 12-11 A, Structure of a spore. B, High concentrations of dipicolinic acid in the spore bind calcium and stabilize the contents. C, Sporogenesis, the process of endospore formation.
Depletion of specific nutrients (e.g., alanine) from the growth medium triggers a cascade of genetic events (comparable to differentiation) leading to the production of a spore. Spore mRNAs are transcribed, and other mRNAs are turned off. Dipicolinic acid is produced, and antibiotics and toxins are often excreted. After duplication of the chromosome, one copy of the DNA and cytoplasmic contents (core) are surrounded by the cytoplasmic membrane, the peptidoglycan, and the membrane of the septum. This wraps the DNA in the two layers of membrane and peptidoglycan that would normally divide the cell. These two layers are surrounded by the cortex, which is made up of a thin inner layer of tightly cross-linked peptidoglycan surrounding a membrane (which used to be the cytoplasmic membrane) and a loose outer peptidoglycan layer. The cortex is surrounded by the tough, keratin-like protein coat, which protects the spore. The process requires 6 to 8 hours for completion.
The germination of spores into the vegetative state is stimulated by disruption of the outer coat by mechanical stress, pH, heat, or another stressor and requires water and a triggering nutrient (e.g., alanine). The process takes approximately 90 minutes. After the germination process begins, the spore will take up water, swell, shed its coats, and produce one new vegetative cell identical to the original vegetative cell, thus completing the entire cycle. Once germination has begun and the spore coat has been compromised, the spore is weakened, vulnerable, and can be inactivated like other bacteria.
1. How does each of the differences between prokaryotes and eukaryotes influence bacterial infection and treatment? (See Figure 12-1.)
2. How do the differences between gram-positive and gram-negative cell walls influence the cells’ clinical behavior, detection, and treatment?
3. List the cell wall components that contribute to virulence by protecting the bacteria from immune responses. List those that contribute to virulence by eliciting toxic responses in the human host.
4. When peptidoglycan synthesis is inhibited, what processes kill the bacteria? List the precursors that would build up within the bacteria if recycling of bactoprenol were inhibited by penicillin, vancomycin, or bacitracin.
5. Why are spores more resistant to environmental stresses?
6. The laboratory would like to selectively eliminate gram-positive bacteria from a mixture of gram-positive and gram-negative bacteria. Which of the following procedures would be more appropriate and why or why not?
1. Size: The smaller size of prokaryotes allows them to enter smaller spaces. It also means that the cells have a smaller chromosome.
Nuclear structures: The lack of a nuclear membrane allows chromosome replication, transcription, and translation to be tied together. Inhibition of any one of them affects the others to a greater degree.
Chromosomes: The bacterial chromosome is a single, circular genome. As a circular chromosome, topoisomerases are very important to relieve stress on the structure and to maintain its function. As a result, these enzymes are excellent targets for antibacterial drugs (e.g., quinolones). Having only one copy of each gene (haploid genome) instead of a diploid genome means that a single mutation will inactivate a function because there is no “backup copy.”
Cytoplasmic structures: Prokaryotes lack organelles, but this does not have a big effect on bacterial infection and treatment.
Ribosomes: The 70S (50S + 30S) provides an excellent target for antibacterial drugs because it differs so significantly from the 80S eukaryotic ribosome.
Cytoplasmic membrane: The prokaryotic membrane contains different phospholipids, which makes it susceptible to polymyxin action. The bacterial membrane also maintains a membrane potential to drive ATP synthesis and other functions.
Cell wall: The bacterial cell wall is a complex structure containing protein, lipids, and peptidoglycan, which is unique to bacteria. The cell wall provides sufficient strength against osmotic shock to allow bacteria to exist in distilled water. It contains structures that promote interactions with tissues and target cells to promote and define the types of infections and diseases caused by bacteria; the enzymes that synthesize these structures are sufficiently unique to be excellent targets for antibacterial drugs (e.g., β-lactams, vancomycin, bacitracin). Pili are very important for promoting adhesion, which allows the bacteria to attach and maintain their location in the body (e.g., in the bladder).
2. The thickness of the gram-positive membrane facilitates its identification by the Gram stain by trapping the stain, whereas the gram-negative peptidoglycan is only a single layer thick, and the stain washes away during the procedure, requiring use of a counterstain. The LPS present in the outer membrane is the most potent activator of innate and immune host cell functions of any cell wall component and can induce fever and sepsis. Gram-negative bacteria are more likely to induce fever and sepsis. The presence of the outer membrane of gram-negative bacteria provides a unique barrier to complement, to the permeability of large and hydrophobic molecules, and prevents access to peptidoglycan and other internal bacterial structures, including antibacterial drugs.
3. Protection from immune responses:
• Peptidoglycan prevents and the O antigen of LPS limits access of the complement membrane attack complex from the membrane surface.
• Capsule protects against antibody, complement, and phagocytosis.
• Proteins may inhibit specific functions (e.g., Staphylococcus protein A binds the Fc portion of IgG; M protein of Streptococcus is antiphagocytic).
• LPS, which contains endotoxin activity and is a potent activator of Toll-like receptor and other receptors.
• Teichoic acid, peptidoglycan, and other cell wall components are weaker activators of Toll-like receptors.
4. Inhibition of peptidoglycan synthesis inhibits cell wall production and the growth of the bacteria. The peptidoglycan is constantly being degraded, rebuilt, and reshaped. Inhibition of peptidoglycan synthesis does not prevent these processes, and, therefore, the peptidoglycan IN A GROWING CELL will continue to degrade, become weakened, and promote the lysis of the cell.
Upon inhibition of peptidoglycan synthesis by β-lactams, vancomycin, or bacitracin antibacterial drugs, the NAG-NAM-pentapeptide (the precursor with a terminal D-ala-D-ala) will build up in the cytoplasm because the chain is not extended (β-lactams, vancomycin) or because the bactoprenol translocation system is inhibited.
5. Spores are more resistant because they are not growing; they are desiccated, and they are covered with multilayers of a peptidoglycan-like material and a keratin-like protein coat.
6. a. EDTA will disrupt gram-negative outer membranes but will have minimal effect on gram-positive bacteria.
b. Mild detergent will affect gram-positive bacteria to a greater extent than it affects gram-negative bacteria because the outer membrane of the latter provides some protection.
c. Lysozyme will degrade the peptidoglycan of gram-positive bacteria, causing them to lyse in water, whereas the outer membrane of gram-negative bacteria poses a barrier and is a protection from lysozyme.
d. Transpeptidase will have no effect on either bacteria.
e. Ampicillin will inhibit peptidoglycan synthesis of both gram-positive and gram-negative bacteria because it can pass through the porin channels of the gram-negative outer membrane.