13 Bacterial Metabolism and Genetics
Bacterial Metabolism
Metabolic Requirements
Bacterial growth requires a source of energy and the raw materials to build the proteins, structures, and membranes that make up and power the cell. Bacteria must obtain or synthesize the amino acids, carbohydrates, and lipids used as building blocks of the cell.
The minimum requirements for growth are a source of carbon and nitrogen, an energy source, water, and various ions. The essential elements include the components of proteins, lipids and nucleic acids (C, O, H, N, S, P), important ions (K, Na, Mg, Ca, Cl), and components of enzymes (Fe, Zn, Mn, Mo, Se, Co, Cu, Ni). Iron is so important that many bacteria secrete special proteins (siderophores) to concentrate iron from dilute solutions, and our bodies will sequester iron to reduce its availability as a means of protection.
Oxygen (O2 gas), although essential for the human host, is actually a poison for many bacteria. Some organisms, such as Clostridium perfringens, which causes gas gangrene, cannot grow in the presence of oxygen. Such bacteria are referred to as obligate anaerobes. Other organisms, such as Mycobacterium tuberculosis, which causes tuberculosis, require the presence of molecular oxygen for metabolism and growth and are therefore referred to as obligate aerobes. Most bacteria, however, grow in either the presence or the absence of oxygen. These bacteria are referred to as facultative anaerobes. Aerobic bacteria produce superoxide dismutase and catalase enzymes, which can detoxify hydrogen peroxide and superoxide radicals that are the toxic byproducts of aerobic metabolism.
Growth requirements and metabolic byproducts may be used as a convenient means of classifying different bacteria. Some bacteria, such as certain strains of Escherichia coli (a member of the intestinal flora), can synthesize all the amino acids, nucleotides, lipids, and carbohydrates necessary for growth and division, whereas the growth requirements of the causative agent of syphilis, Treponema pallidum, are so complex that a defined laboratory medium capable of supporting its growth has yet to be developed. Bacteria that can rely entirely on inorganic chemicals for their energy and source of carbon (carbon dioxide [CO2]) are referred to as autotrophs (lithotrophs), whereas many bacteria and animal cells that require organic carbon sources are known as heterotrophs (organotrophs). Clinical microbiology laboratories distinguish bacteria by their ability to grow on specific carbon sources (e.g., lactose) and the end products of metabolism (e.g., ethanol, lactic acid, succinic acid).
Metabolism, Energy, and Biosynthesis
All cells require a constant supply of energy to survive. This energy, typically in the form of adenosine triphosphate (ATP), is derived from the controlled breakdown of various organic substrates (carbohydrates, lipids, and proteins). This process of substrate breakdown and conversion into usable energy is known as catabolism. The energy produced may then be used in the synthesis of cellular constituents (cell walls, proteins, fatty acids, and nucleic acids), a process known as anabolism. Together these two processes, which are interrelated and tightly integrated, are referred to as intermediary metabolism.
The metabolic process generally begins with hydrolysis of large macromolecules in the external cellular environment by specific enzymes (Figure 13-1). The smaller molecules that are produced (e.g., monosaccharides, short peptides, and fatty acids) are transported across the cell membranes into the cytoplasm by active or passive transport mechanisms specific for the metabolite. These mechanisms may use specific carrier or membrane transport proteins to help concentrate metabolites from the medium. The metabolites are converted via one or more pathways to one common, universal intermediate, pyruvic acid. From pyruvic acid, the carbons may be channeled toward energy production or the synthesis of new carbohydrates, amino acids, lipids, and nucleic acids.
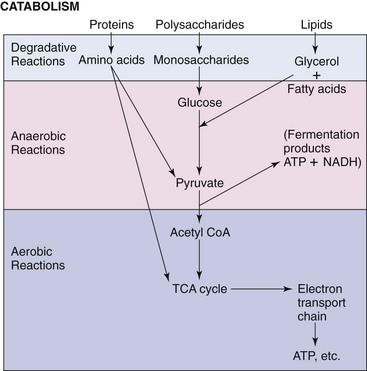
Figure 13-1 Catabolism of proteins, polysaccharides, and lipids produces glucose, pyruvate, or intermediates of the tricarboxylic acid (TCA) cycle and, ultimately, energy in the form of adenosine triphosphate (ATP) or the reduced form of nicotinamide adenine dinucleotide (NADH). CoA, Coenzyme A.
Metabolism of Glucose
For the sake of simplicity, this section presents an overview of the pathways by which glucose is metabolized to produce energy or other usable substrates. Instead of releasing all of glucose’s energy as heat (as for burning), the bacteria break down glucose in discrete steps to allow the energy to be captured in usable forms. Bacteria can produce energy from glucose by—in order of increasing efficiency—fermentation, anaerobic respiration (both of which occur in the absence of oxygen), or aerobic respiration. Aerobic respiration can completely convert the six carbons of glucose to CO2 and water (H2O) plus energy, whereas two- and three-carbon compounds are the end products of fermentation. For a more complete discussion of metabolism, please refer to a textbook on biochemistry.
Embden-Meyerhof-Parnas Pathway
Bacteria use three major metabolic pathways in the catabolism of glucose. Most common among these is the glycolytic, or Embden-Meyerhof-Parnas (EMP), pathway (Figure 13-2) for the conversion of glucose to pyruvate. These reactions, which occur under both aerobic and anaerobic conditions, begin with activation of glucose to form glucose-6-phosphate. This reaction, as well as the third reaction in the series, in which fructose-6-phosphate is converted to fructose-1,6-diphosphate, requires 1 mole of ATP per mole of glucose and represents an initial investment of cellular energy stores.
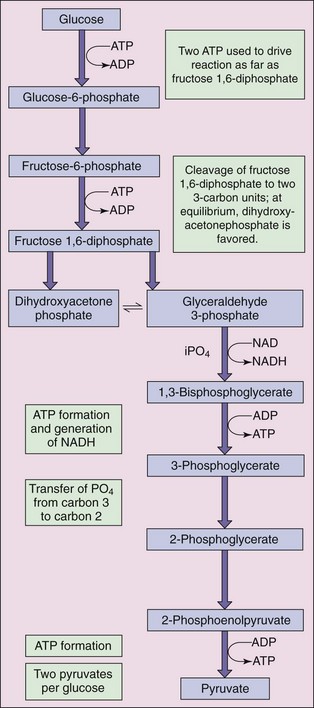
Figure 13-2 Embden-Meyerhof-Parnas glycolytic pathway results in conversion of glucose to pyruvate. ADP, Adenosine diphosphate; ATP, adenosine triphosphate; iPO4, inorganic phosphate; NAD, nicotinamide adenine dinucleotide; NADH, reduced form of NAD.
Energy is produced during glycolysis in two different forms, chemical and electrochemical. In the first, the high-energy phosphate group of one of the intermediates in the pathway is used under the direction of the appropriate enzyme (a kinase) to generate ATP from adenosine diphosphate (ADP). This type of reaction, termed substrate-level phosphorylation, occurs at two different points in the glycolytic pathway (i.e., conversion of 3-phosphoglycerol phosphate to 3-phosphoglycerate and 2-phosphoenolpyruvic acid to pyruvate).
This pathway yields four ATP molecules per molecule of glucose, but two ATP molecules were used in the initial glycolytic conversion of glucose to two molecules of pyruvic acid, resulting in a net production of two molecules of ATP, two molecules of reduced nicotinamide adenine dinucleotide (NADH) and two pyruvate molecules. NADH may then be converted to ATP by a series of oxidation reactions.
In the absence of oxygen, substrate-level phosphorylation represents the primary means of energy production. The pyruvic acid produced from glycolysis is then converted to various end products, depending on the bacterial species, in a process known as fermentation. Many bacteria are identified on the basis of their fermentative end products (Figure 13-3). These organic molecules, rather than oxygen, are used as electron acceptors to recycle the NADH, which was produced during glycolysis, to NAD. In yeast, fermentative metabolism results in the conversion of pyruvate to ethanol plus CO2. Alcoholic fermentation is uncommon in bacteria, which most commonly use the one-step conversion of pyruvic acid to lactic acid. This process is responsible for making milk into yogurt and cabbage into sauerkraut. Other bacteria use more complex fermentative pathways, producing various acids, alcohols, and often gases (many of which have vile odors). These products lend flavors to various cheeses and wines and odors to wound and other infections.
Tricarboxylic Acid Cycle
In the presence of oxygen, the pyruvic acid produced from glycolysis and from the metabolism of other substrates may be completely oxidized (controlled burning) to H2O and CO2 using the tricarboxylic acid (TCA) cycle (Figure 13-4), which results in production of additional energy. The process begins with the oxidative decarboxylation (release of CO2) of pyruvate to the high-energy intermediate, acetyl coenzyme A (acetyl CoA); this reaction also produces two NADH molecules. The two remaining carbons derived from pyruvate then enter the TCA cycle in the form of acetyl CoA by condensation with oxaloacetate, with the formation of the six-carbon citrate molecule. In a stepwise series of oxidative reactions, the citrate is converted back to oxaloacetate. The theoretical yield from each pyruvate is 2 moles of CO2, 3 moles of NADH, 1 mole of flavin adenine dinucleotide (FADH2), and 1 mole of guanosine triphosphate (GTP).
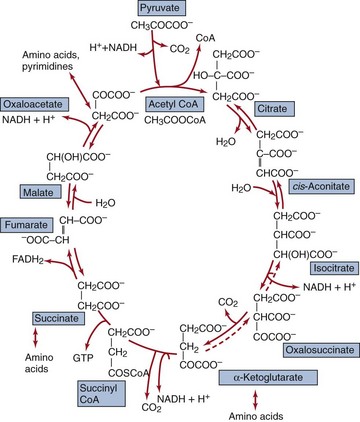
Figure 13-4 Tricarboxylic acid cycle is an amphibolic cycle. Precursors for the synthesis of amino acids and nucleotides are also shown. CoA, Coenzyme A; FADH2, reduced form of flavin adenine dinucleotide; GTP, guanosine triphosphate; NADH, reduced form of nicotinamide adenine dinucleotide.
The TCA cycle allows the organism to generate substantially more energy per mole of glucose than is possible from glycolysis alone. In addition to the GTP (an ATP equivalent) produced by substrate-level phosphorylation, the NADH and FADH2 yield ATP from the electron transport chain. In this chain the electrons carried by NADH (or FADH2) are passed in a stepwise fashion through a series of donor-acceptor pairs and ultimately to oxygen (aerobic respiration) or other terminal electron acceptor (nitrate, sulfate, CO2, ferric iron) (anaerobic respiration).
Anaerobic organisms are less efficient at energy production than aerobic organisms. Fermentation produces only two ATP molecules per glucose, whereas aerobic metabolism with electron transport and a complete TCA cycle can generate as much as 19 times more energy (38 ATP molecules) from the same starting material (and it is much less smelly) (Figure 13-5). Anaerobic respiration uses organic molecules as electron acceptors, which produces less ATP for each NADH than aerobic respiration.
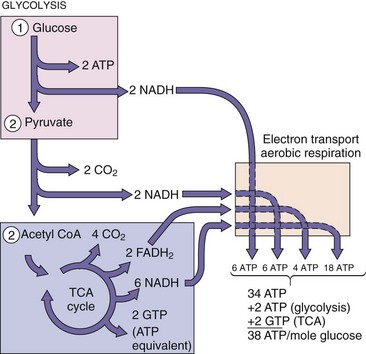
Figure 13-5 Aerobic glucose metabolism. The theoretical maximum amount of adenosine triphosphate (ATP) obtained from one glucose molecule is 38, but the actual yield depends on the organism and other conditions. FADH2, Reduced form of flavin adenine dinucleotide; GTP, guanosine triphosphate; NADH, reduced form of nicotinamide adenine dinucleotide; TCA, tricarboxylic acid.
In addition to the efficient generation of ATP from glucose (and other carbohydrates), the TCA cycle provides a means by which carbons derived from lipids (in the form of acetyl CoA) may be shunted toward either energy production or the generation of biosynthetic precursors. Similarly, the cycle includes several points at which deaminated amino acids may enter (see Figure 13-4). For example, deamination of glutamic acid yields α-ketoglutarate, whereas deamination of aspartic acid yields oxaloacetate, both of which are TCA cycle intermediates. The TCA cycle therefore serves the following functions:
1. It is the most efficient mechanism for the generation of ATP.
2. It serves as the final common pathway for the complete oxidation of amino acids, fatty acids, and carbohydrates.
3. It supplies key intermediates (i.e., α-ketoglutarate, pyruvate, oxaloacetate) for the ultimate synthesis of amino acids, lipids, purines, and pyrimidines.
The last two functions make the TCA cycle a so-called amphibolic cycle (i.e., it may function to break down and synthesize molecules).
Pentose Phosphate Pathway
The final pathway of glucose metabolism considered here is known as the pentose phosphate pathway, or the hexose monophosphate shunt. The function of this pathway is to provide nucleic acid precursors and reducing power in the form of nicotinamide adenine dinucleotide phosphate (reduced form) (NADPH) for use in biosynthesis. In the first half of the pathway, glucose is converted to ribulose-5-phosphate, with consumption of 1 mole of ATP and generation of 2 moles of NADPH per mole of glucose. The ribulose-5-phosphate may then be converted to ribose-5-phosphate (a precursor in nucleotide biosynthesis) or alternatively to xylulose-5-phosphate. The remaining reactions in the pathway use enzymes known as transketolases and transaldolases to generate various sugars, which may function as biosynthetic precursors or may be shunted back to the glycolytic pathway for use in energy generation.
Bacterial Genes and Expression
The bacterial genome is the total collection of genes carried by a bacterium, both on its chromosome and on its extrachromosomal genetic elements, if any. Genes are sequences of nucleotides that have a biologic function; examples are protein-structural genes (cistrons, which are coding genes), ribosomal ribonucleic acid (RNA) genes, and recognition and binding sites for other molecules (promoters and operators). Each genome contains many operons, which are made up of genes. Genes may also be grouped in islands, such as the pathogenicity islands, which share function or to coordinate their control.
Bacteria usually have only one copy of their chromosomes (they are therefore haploid), whereas eukaryotes usually have two distinct copies of each chromosome (they are therefore diploid). With only one chromosome, alteration of a bacterial gene (mutation) will have a more obvious effect on the cell. In addition, the structure of the bacterial chromosome is maintained by polyamines, such as spermine and spermidine, rather than by histones.
Bacteria may also contain extrachromosomal genetic elements such as plasmids or bacteriophages (bacterial viruses). These elements are independent of the bacterial chromosome and, in most cases, can be transmitted from one cell to another.
Transcription
The information carried in the genetic memory of the deoxyribonucleic acid (DNA) is transcribed into a useful messenger RNA (mRNA) for subsequent translation into protein. RNA synthesis is performed by a DNA-dependent RNA polymerase. The process begins when the sigma factor recognizes a particular sequence of nucleotides in the DNA (the promoter) and binds tightly to this site. Promoter sequences occur just before the start of the DNA that actually encodes a protein. Sigma factors bind to these promoters to provide a docking site for the RNA polymerase. Some bacteria encode several sigma factors to allow transcription of a group of genes under special conditions, such as heat shock, starvation, special nitrogen metabolism, or sporulation.
Once the polymerase has bound to the appropriate site on the DNA, RNA synthesis proceeds with the sequential addition of ribonucleotides complementary to the sequence in the DNA. Once an entire gene or group of genes (operon) has been transcribed, the RNA polymerase dissociates from the DNA, a process mediated by signals within the DNA. The bacterial DNA-dependent RNA polymerase is inhibited by rifampin, an antibiotic often used in the treatment of tuberculosis. The transfer RNA (tRNA), which is used in protein synthesis, and ribosomal RNA (rRNA), a component of the ribosomes, are also transcribed from the DNA.
Promoters and operators control the expression of a gene by influencing which sequences will be transcribed into mRNA. Operons are groups of one or more structural genes expressed from a particular promoter and ending at a transcriptional terminator. Thus all the genes coding for the enzymes of a particular pathway can be coordinately regulated. Operons with many structural genes are polycistronic. The E. coli lac operon includes all the genes necessary for lactose metabolism, as well as the control mechanisms for turning off (in the presence of glucose) or turning on (in the presence of galactose or an inducer) these genes only when they are needed. The lac operon includes a repressor sequence, a promoter sequence, and structural genes for the β-galactosidase enzyme, a permease, and an acetylase (Figure 13-6). The lac operon is discussed later in this chapter.
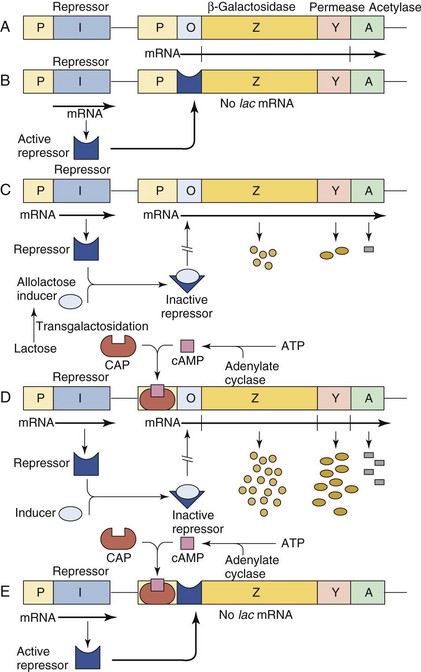
Figure 13-6 A, The lactose operon is transcribed as a polycistronic messenger RNA (mRNA) from the promoter (P) and translated into three proteins: β-galactosidase (Z), permease (Y), and acetylase (A). The (I) gene encodes the repressor protein. B, The lactose operon is not transcribed in the absence of an allolactose inducer because the repressor competes with the RNA polymerase at the operator site (O). C, The repressor, complexed with the inducer, does not recognize the operator because of a conformation change in the repressor. The lac operon is thus transcribed at a low level. D, Escherichia coli is grown in a poor medium in the presence of lactose as the carbon source. Both the inducer and the CAP-cAMP complex are bound to the promoter, which is fully “turned on,” and a high level of lac mRNA is transcribed and translated. E, Growth of E. coli in a poor medium without lactose results in the binding of the CAP-cAMP complex to the promoter region and binding of the active repressor to the operator sequence because no inducer is available. The result will be that the lac operon will not be transcribed. ATP, Adenosine triphosphate; CAP, catabolite gene-activator protein; cAMP, cyclic adenosine monophosphate.
Translation
Translation is the process by which the language of the genetic code, in the form of mRNA, is converted (translated) into a sequence of amino acids, the protein product. Each amino acid word and the punctuation of the genetic code is written as sets of three nucleotides, known as codons. There are 64 different codon combinations encoding the 20 amino acids, plus start and termination codons. Some of the amino acids are encoded by more than one triplet codon. This feature is known as the degeneracy of the genetic code and may function in protecting the cell from the effects of minor mutations in the DNA or mRNA. Each tRNA molecule contains a three-nucleotide sequence complementary to one of the codon sequences. This tRNA sequence is known as the anticodon; it allows base pairing and binds to the codon sequence on the mRNA. Attached to the opposite end of the tRNA is the amino acid that corresponds to the particular codon-anticodon pair.
The process of protein synthesis (Figure 13-7) begins with the binding of the 30S ribosomal subunit and a special initiator tRNA for formyl methionine (fMet) at the methionine codon (AUG) start codon to form the initiation complex. The 50S ribosomal subunit binds to the complex to initiate mRNA synthesis. The ribosome contains two tRNA binding sites, the A (aminoacyl) site and the P (peptidyl) site, each of which allows base pairing between the bound tRNA and the codon sequence in the mRNA. The tRNA corresponding to the second codon occupies the A site. The amino group of the amino acid attached to the A site forms a peptide bond with the carboxyl group of the amino acid in the P site in a reaction known as transpeptidation, and the empty tRNA in the P site (uncharged tRNA) is released from the ribosome. The ribosome then moves down the mRNA exactly three nucleotides, thereby transferring the tRNA with attached nascent peptide to the P site and bringing the next codon into the A site. The appropriate charged tRNA is brought into the A site, and the process is then repeated. Translation continues until the new codon in the A site is one of the three termination codons, for which there is no corresponding tRNA. At that point the new protein is released to the cytoplasm and the translation complex may be disassembled, or the ribosome shuffles to the next start codon and initiates a new protein. The ability to shuffle along the mRNA to start a new protein is a characteristic of the 70S bacterial but not of the 80S eukaryotic ribosome. The eukaryotic constraint has implications for the synthesis of proteins for some viruses.
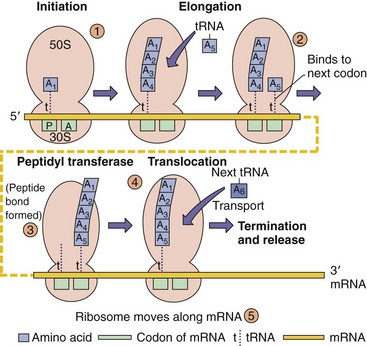
Figure 13-7 Bacterial protein synthesis. 1, Binding of the 30S subunit to the messenger RNA (mRNA) with the formyl methionine transfer RNA (fMet-tRNA) at the AUG start codon allows assembly of the 70S ribosome. The fMet-tRNA binds to the peptidyl site (P). 2, The next tRNA binds to its codon at the A site and “accepts” the growing peptide chain. 3 and 4, Before translocation to the peptidyl site. 5, The process is repeated until a stop codon and the protein are released.
The process of protein synthesis by the 70S ribosome represents an important target of antimicrobial action. The aminoglycosides (e.g., streptomycin and gentamicin) and the tetracyclines act by binding to the small ribosomal subunit and inhibiting normal ribosomal function. Similarly, the macrolide (e.g., erythromycin) and lincosamide (e.g., clindamycin) groups of antibiotics act by binding to the large ribosomal subunit. Also, formyl methionine peptides (e.g., fmet-leu-phe) attract neutrophils to the site of an infection.
Control of Gene Expression
Bacteria have developed mechanisms to adapt quickly and efficiently to changes and triggers from the environment. This allows them to coordinate and regulate the expression of genes for multicomponent structures or the enzymes of one or more metabolic pathways. For example, temperature change could signify entry into the human host and indicate the need for a global change in metabolism and up-regulation of genes important for parasitism or virulence. Many bacterial genes are controlled at multiple levels and by multiple methods.
A coordinated change in the expression of many genes, as would be required for sporulation, occurs through use of a different sigma factor for the RNA polymerase. This would change the specificity of the RNA polymerase and allow mRNA synthesis for the necessary genes while ignoring unnecessary genes. Bacteria might produce more than six different sigma factors to provide global regulation in response to stress, shock, starvation, or to coordinate production of complicated structures such as flagella.
Coordination of a large number of processes on a global level can also be mediated by small molecular activators, such as cyclic adenosine monophosphate (cAMP). Increased cAMP levels indicate low glucose levels and the need to utilize alternative metabolic pathways. Similarly, in a process called quorum sensing, when a sufficient number of bacteria are present and producing a specific small molecule, virulence and other genes are turned on. The trigger for biofilm production by Pseudomonas spp. is triggered by a critical concentration of N-acyl homoserine lactone (AHL) produced when sufficient numbers of bacteria (a quorum) are present. Activation of toxin production and more virulent behavior by Staphylococcus aureus accompanies the increase in concentration of a cyclic peptide.
To coordinate the expression of a more limited group of genes, such as for a specific metabolic process, the genes for the necessary enzymes would be organized into an operon. The operon would be under the control of a promoter or repressor DNA sequence that can activate or turn off the expression of a gene or a group of genes to coordinate production of the necessary enzymes and allow the bacteria to react to changes in concentrations of nutrients. The genes for some virulence mechanisms are organized into a pathogenicity island under the control of a single promoter to allow their expression under appropriate (to the bacteria) conditions. The many components of the type III secretion devices of E. coli, Salmonella, or Yersinia are grouped together within a pathogenicity island.
Transcription can also be regulated by the translation process. Unlike eukaryotes, the absence of a nuclear membrane in prokaryotes allows the ribosome to bind to the mRNA as it is being transcribed from the DNA. The position and speed of ribosomal movement along the mRNA can affect the presence of loops in the mRNA and the ability of the polymerase to transcribe new mRNA. This allows control of gene expression at both the transcriptional and translational levels.
Initiation of transcription may be under positive or negative control. Genes under negative control are expressed unless they are switched off by a repressor protein. This repressor protein prevents gene expression by binding to a specific DNA sequence called the operator, blocking the RNA polymerase from initiating transcription at the promoter sequence. Inversely, genes whose expression is under positive control are not transcribed unless an active regulator protein, called an apoinducer, is present. The apoinducer binds to a specific DNA sequence and assists the RNA polymerase in the initiation steps by an unknown mechanism.
Operons can be inducible or repressible. Introduction of a substrate (inducer) into the growth medium may induce an operon to increase the expression of the enzymes necessary for its metabolism. An abundance of the end products (co-repressors) of a pathway may signal that a pathway should be shut down or repressed by reducing the synthesis of its enzymes.
The lactose (lac) operon responsible for the degradation of the sugar lactose is an inducible operon under positive and negative regulation (see Figure 3-6). Normally the bacteria use glucose and not lactose. In the absence of lactose the operon is repressed by the binding of the repressor protein to the operator sequence, thus impeding the RNA polymerase function. In the absence of glucose, however, the addition of lactose reverses this repression. Full expression of the lac operon also requires a protein-mediated, positive-control mechanism. In E. coli, when glucose decreases in the cell, cAMP increases to promote usage of other sugars for metabolism. Binding of cAMP to a protein called the catabolite gene-activator protein (CAP) allows it to bind to a specific DNA sequence present in the promoter. The CAP-cAMP complex enhances binding of the RNA polymerase to the promoter, thus allowing an increase in the frequency of transcription initiation.
The tryptophan operon (trp operon) contains the structural genes necessary for tryptophan biosynthesis and is under dual transcriptional control mechanisms (Figure 13-8). Although tryptophan is essential for protein synthesis, too much tryptophan in the cell can be toxic; therefore its synthesis must be regulated. At the DNA level the repressor protein is activated by an increased intracellular concentration of tryptophan to prevent transcription. At the protein synthesis level, rapid translation of a “test peptide” at the beginning of the mRNA in the presence of tryptophan allows the formation of a double-stranded loop in the RNA, which terminates transcription. The same loop is formed if no protein synthesis is occurring, a situation in which tryptophan synthesis would similarly not be required. This regulates tryptophan synthesis at the mRNA level in a process termed attenuation, in which mRNA synthesis is prematurely terminated.
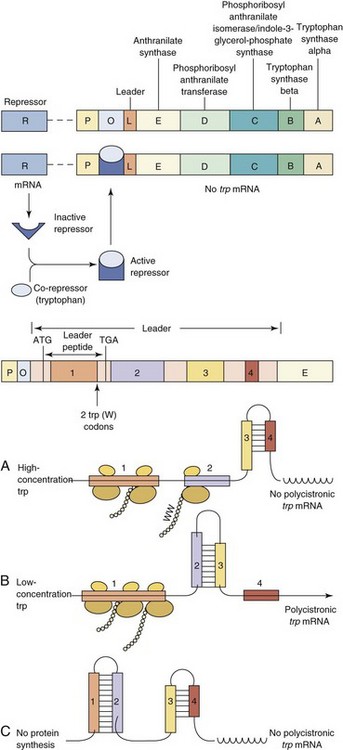
Figure 13-8 Regulation of the tryptophan (trp) operon. A, The trp operon encodes the five enzymes necessary for tryptophan biosynthesis. This trp operon is under dual control. B, The conformation of the inactive repressor protein is changed after its binding by the co-repressor tryptophan. The resulting active repressor (R) binds to the operator (O), blocking any transcription of the trp mRNA by the RNA polymerase. C, The trp operon is also under the control of an attenuation-antitermination mechanism. Upstream of the structural genes are the promoter (P), the operator, and a leader (L), which can be transcribed into a short peptide containing two tryptophans (W), near its distal end. The leader mRNA possesses four repeats (1, 2, 3, and 4), which can pair differently according to the tryptophan availability, leading to an early termination of transcription of the trp operon or its full transcription. In the presence of a high concentration of tryptophan, regions 3 and 4 of the leader mRNA can pair, forming a terminator hairpin, and no transcription of the trp operon occurs. However, in the presence of little or no tryptophan the ribosomes stall in region 1 when translating the leader peptide because of the tandem of tryptophan codons. Then regions 2 and 3 can pair, forming the antiterminator hairpin and leading to transcription of the trp genes. Finally, the regions 1 : 2 and 3 : 4 of the free leader mRNA can pair, also leading to cessation of transcription before the first structural gene trpE. A, Adenine; G, guanine; T, thymidine.
The expression of the components of virulence mechanisms are also coordinately regulated from an operon. Simple triggers, such as temperature, osmolarity, pH, nutrient availability, or the concentration of specific small molecules, such as oxygen or iron, can turn on or turn off the transcription of a single gene or a group of genes. Salmonella invasion genes within a pathogenicity island are turned on by high osmolarity and low oxygen, conditions present in the gastrointestinal tract or an endosomal vesicle within a macrophage. E. coli senses its exit from the gut of a host by a drop in temperature and inactivates its adherence genes. Low iron levels can activate expression of hemolysin in E. coli or diphtheria toxin from Corynebacterium diphtheriae, potentially to kill cells and provide iron. Quorum sensing for virulence factors of S. aureus and biofilm production by Pseudomonas spp. were discussed above. An example of coordinated control of virulence genes for S. aureus based on the growth rate, availability of metabolites, and the presence of a quorum is presented in Figure 13-9.
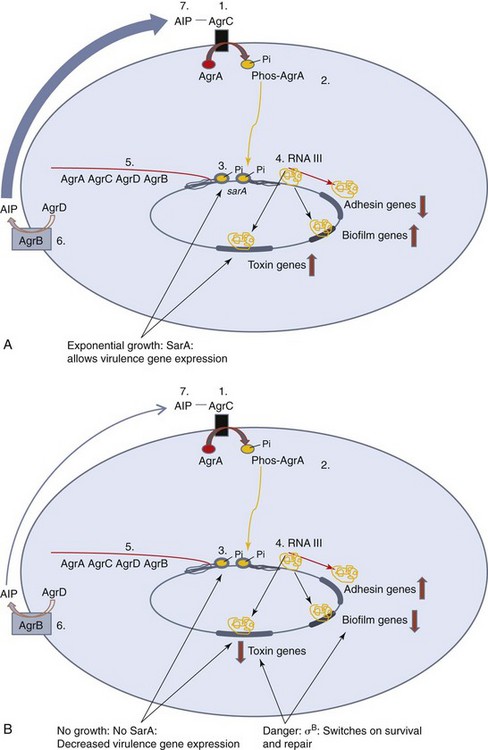
Figure 13-9 Control of virulence genes in Staphylococcus aureus. S. aureus switches on virulence factors when in exponential growth and when their numbers increase to a quorum. Toxin and protease are produced to kill host cells and supply the colony with food, and the colony produces a biofilm for protection. Cell wall thickness and adhesion factors are less important within the colony and are repressed. Quorum sensing is mediated and autoinduced by the Agr (A-D) proteins. A, 1. The autoinducing peptide (AIP) binds to AgrC. 2, AgrC is a receptor that phosphorylates AgrA. 3, Phosphorylated AgrA activates the promoter for the agr operon and the promoter for a regulatory RNA called RNA III. 4, RNA III contains the 26–amino acid δ-hemolysin RNA sequence. In addition, RNA III activates toxin and other virulence genes while decreasing expression of adhesion and cell wall synthesis genes. 5, AgrD interacts with AgrB, in the membrane, to be converted into the AIP. As long as the bacteria are in exponential phase growth, they produce SarA, which also binds and activates the promoters for the agr and RNAIII genes. B, Upon metabolic problems and danger, SarA production is decreased and a new sigma factor, σB is produced to decrease production of these virulence factors and σB turns on DNA and cellular repair mechanisms. Large red arrows indicate increases or decreases in expression.
Replication of DNA
Replication of the bacterial genome is triggered by a cascade of events linked to the growth rate of the cell. Replication of bacterial DNA is initiated at a specific sequence in the chromosome called oriC. The replication process requires many enzymes, including an enzyme (helicase) to unwind the DNA at the origin to expose the DNA, an enzyme (primase) to synthesize primers to start the process, and the enzyme or enzymes (DNA-dependent DNA polymerases) that synthesize a copy of the DNA, but only if there is a primer sequence to add onto and only in the 5′ to 3′ direction.
New DNA is synthesized semiconservatively, using both strands of the parental DNA as templates. New DNA synthesis occurs at growing forks and proceeds bidirectionally. One strand (the leading strand) is copied continuously in the 5′ to 3′ direction, whereas the other strand (the lagging strand) must be synthesized as many pieces of DNA using RNA primers (Okazaki fragments). The lagging-strand DNA must be extended in the 5′ to 3′ direction as its template becomes available. Then the pieces are ligated together by the enzyme DNA ligase (Figure 13-10). To maintain the high degree of accuracy required for replication, the DNA polymerases possess “proofreading” functions, which allow the enzyme to confirm that the appropriate nucleotide was inserted and to correct any errors that were made. During log-phase growth in rich medium, many initiations of chromosomal replication may occur before cell division. This process produces a series of nested bubbles of new daughter chromosomes, each with its pair of growth forks of new DNA synthesis. The polymerase moves down the DNA strand, incorporating the appropriate (complementary) nucleotide at each position. Replication is complete when the two replication forks meet 180 degrees from the origin. The process of DNA replication puts great torsional strain on the chromosomal circle of DNA; this strain is relieved by topoisomerases (e.g., gyrase), which supercoil the DNA. Topoisomerases are essential to the bacteria and are targets for the quinolone antibiotics.
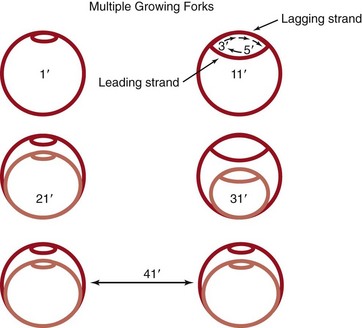
Figure 13-10 Bacterial DNA replication. New DNA synthesis occurs at growing forks and proceeds bidirectionally. DNA synthesis progresses in the 5′ to 3′ direction continuously (leading strand) or in pieces (lagging strand). Assuming it takes 40 minutes to complete one round of replication, and assuming new initiation every 20 minutes, initiation of DNA synthesis precedes cell division. Multiple growing forks may be initiated in a cell before complete septum formation and cell division. The daughter cells are “born pregnant.”
Bacterial Growth
Bacterial replication is a coordinated process in which two equivalent daughter cells are produced. For growth to occur, there must be sufficient metabolites to support the synthesis of the bacterial components and especially the nucleotides for DNA synthesis. A cascade of regulatory events (synthesis of key proteins and RNA), much like a countdown at the Kennedy Space Center, must occur on schedule to initiate a replication cycle. However, once it is initiated, DNA synthesis must run to completion, even if all nutrients have been removed from the medium.
Chromosome replication is initiated at the membrane, and each daughter chromosome is anchored to a different portion of membrane. Bacterial membrane, peptidoglycan synthesis, and cell division are linked together such that inhibition of peptidoglycan synthesis will also inhibit cell division. As the bacterial membrane grows, the daughter chromosomes are pulled apart. Commencement of chromosome replication also initiates the process of cell division, which can be visualized by the start of septum formation between the two daughter cells (Figure 13-11; see also Chapter 12). New initiation events may occur even before completion of chromosome replication and cell division.
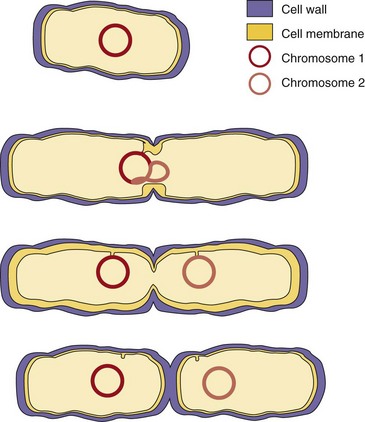
Figure 13-11 Bacterial cell division. Replication requires extension of the cell wall and replication of the chromosome and septum formation. Membrane attachment of the DNA pulls each daughter strand into a new cell.
Depletion of metabolites (starvation) or a buildup of toxic byproducts (e.g., ethanol) triggers the production of chemical alarmones, which causes synthesis to stop, but degradative processes continue. DNA synthesis continues until all initiated chromosomes are completed, despite the detrimental effect on the cell. Ribosomes are cannibalized for deoxyribonucleotide precursors, peptidoglycan and proteins are degraded for metabolites, and the cell shrinks. Septum formation may be initiated, but cell division may not occur. Many cells die. Similar signals may initiate sporulation in species capable of this process (see Chapter 12).
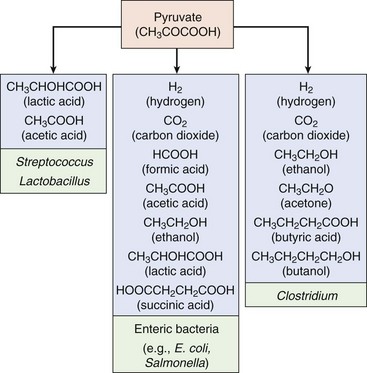
Population Dynamics
When bacteria are added to a new medium, they require time to adapt to the new environment before they begin dividing (Figure 13-12). This hiatus is known as the lag phase of growth. During the logarithmic (log) or exponential phase, the bacteria will grow and divide with a doubling time characteristic of the strain and determined by the conditions. The number of bacteria will increase to 2n, in which n is the number of generations (doublings). The culture eventually runs out of metabolites, or a toxic substance builds up in the medium; the bacteria then stop growing and enter the stationary phase.
Bacterial Genetics
Mutation, Repair, and Recombination
Accurate replication of DNA is important to the survival of the bacteria, but mistakes and accidental damage to the DNA occurs. The bacteria have efficient DNA repair systems, but mutations and alterations to the DNA still occur. Most of these mutations have little effect on the bacteria or are detrimental, but some mutations may provide a selective advantage for survival of the bacteria when challenged by the environment, the host, or therapy.
Mutations and Their Consequences
A mutation is any change in the base sequence of the DNA. A single base change can result in a transition in which one purine is replaced by another purine, or in which a pyrimidine is replaced by another pyrimidine. A transversion, in which, for example, a purine is replaced by a pyrimidine and vice versa, may also result. A silent mutation is a change at the DNA level that does not result in any change of amino acid in the encoded protein. This type of mutation occurs because more than one codon may encode an amino acid. A missense mutation results in a different amino acid being inserted in the protein, but this may be a conservative mutation if the new amino acid has similar properties (e.g., valine replacing alanine). A nonsense mutation changes a codon encoding an amino acid to a stop codon (e.g., TAG [thymidine-adenine-guanine]), which will cause the ribosome to fall off the mRNA and end the protein prematurely. Conditional mutations, such as temperature-sensitive mutations, may result from a conservative mutation which changes the structure or function of an important protein at elevated temperatures.
More drastic changes can occur when numerous bases are involved. A small deletion or insertion that is not in multiples of three produces a frameshift mutation. This results in a change in the reading frame, usually leading to a useless peptide and premature truncation of the protein. Null mutations, which completely destroy gene function, arise when there is an extensive insertion, deletion, or gross rearrangement of the chromosome structure. Insertion of long sequences of DNA (many thousands of base pairs) by recombination, by transposition, or during genetic engineering can produce null mutations by separating the parts of a gene and inactivating the gene.
Many mutations occur spontaneously in nature (e.g., by polymerase mistakes); however, physical or chemical agents can also induce mutations. Among the physical agents used to induce mutations in bacteria are heat, which results in deamination of nucleotides; ultraviolet light, which causes pyrimidine dimer formation; and ionizing radiation, such as x-rays, which produce very reactive hydroxyl radicals that may be responsible for opening a ring of a base or causing single- or double-stranded breaks in the DNA. Chemical mutagens can be grouped into three classes. Nucleotide-base analogues lead to mispairing and frequent DNA replication mistakes. For example, incorporation of 5-bromouracil into DNA instead of thymidine allows base pairing with guanine instead of adenine, changing a T-A base pair to a G-C base pair. Frameshift mutagens, such as polycyclic flat molecules like ethidium bromide or acridine derivatives, insert (or intercalate) between the bases as they stack with each other in the double helix. The increase in spacing of successive base pairs cause the addition or deletion of a single base and lead to frequent mistakes during DNA replication. DNA-reactive chemicals act directly on the DNA to change the chemical structure of the base. These include nitrous acid (HNO2) and alkylating agents, including nitrosoguanidine and ethyl methane sulfonate, which are known to add methyl or ethyl groups to the rings of the DNA bases. The modified bases may pair abnormally or not at all. The damage may also cause the removal of the base from the DNA backbone.
Repair Mechanisms of DNA
A number of repair mechanisms have evolved in bacterial cells to minimize damage to DNA. These repair mechanisms can be divided into the following five groups:
1. Direct DNA repair is the enzymatic removal of damage, such as pyrimidine dimers and alkylated bases.
2. Excision repair is the removal of a DNA segment containing the damage, followed by synthesis of a new DNA strand. Two types of excision-repair mechanisms, generalized and specialized, exist.
3. Recombinational or postreplication repair is the retrieval of missing information by genetic recombination when both DNA strands are damaged.
4. The SOS response is the induction of many genes (approximately 15) after DNA damage or interruption of DNA replication.
5. Error-prone repair is the last resort of a bacterial cell before it dies. It is used to fill in gaps with a random sequence when a DNA template is not available for directing an accurate repair.
Gene Exchange in Prokaryotic Cells
Many bacteria, especially many pathogenic bacterial species, are promiscuous with their DNA. The exchange of DNA between cells allows the exchange of genes and characteristics between cells, thus producing new strains of bacteria. This exchange may be advantageous for the recipient, especially if the exchanged DNA encodes antibiotic resistance. The transferred DNA can be integrated into the recipient chromosome or stably maintained as an extrachromosomal element (plasmid) or a bacterial virus (bacteriophage) and passed on to daughter bacteria as an autonomously replicating unit.
Plasmids are small genetic elements that replicate independently of the bacterial chromosome. Most plasmids are circular, double-stranded DNA molecules varying from 1500 to 400,000 base pairs. However, Borrelia burgdorferi, the causative agent of Lyme disease, and the related Borrelia hermsii are unique among all eubacteria because they possess linear plasmids. Like the bacterial chromosomal DNA, plasmids can autonomously replicate and, as such, are referred to as replicons. Some plasmids, such as the E. coli F plasmid, are episomes, which means that they can integrate into the host chromosome.
Plasmids carry genetic information, which may not be essential but can provide a selective advantage to the bacteria. For example, plasmids may encode the production of antibiotic resistance mechanisms, bacteriocins, toxins, virulence determinants, and other genes that may provide the bacteria with a unique growth advantage over other microbes or within the host (Figure 13-13). The number of copies of plasmid produced by a cell is determined by the particular plasmid. The copy number is the ratio of copies of the plasmid to the number of copies of the chromosome. This may be as few as one in the case of large plasmids or as many as 50 in smaller plasmids.
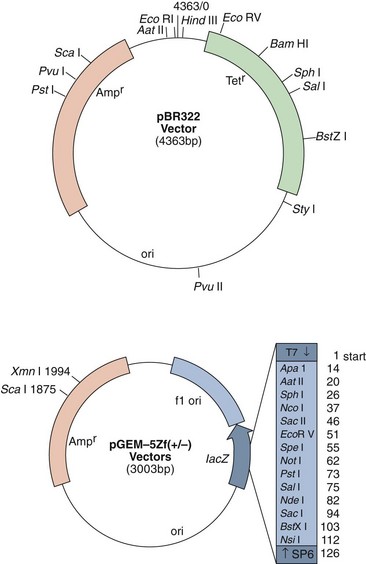
Figure 13-13 Plasmids. The pBR322 plasmid is one of the plasmids used for cloning DNA. This plasmid encodes resistance to ampicillin (Amp) and tetracycline (Tet) and an origin of replication (ori). The multiple cloning site in the pGEM-5Zf(+/−) plasmid provides different restriction enzyme cleavage sites for insertion of DNA within the β-galactosidase gene (lacZ). The insert is flanked by bacteriophage promoters to allow directional messenger RNA expression of the cloned sequence.
Large plasmids (20 to 120 kb), such as the fertility factor F found in E. coli or the resistance transfer factor (80 kb), can often mediate their own transfer from one cell to another by a process called conjugation (see the section on conjugation later in this chapter). These conjugative plasmids encode all the necessary factors for their transfer. Other plasmids can be transferred into a bacterial cell by means other than conjugation, such as transformation or transduction. These terms are also discussed later in the chapter.
Bacteriophages are bacterial viruses with a DNA or RNA genome usually protected by a protein shell. These extrachromosomal genetic elements can survive outside of a host cell and be transmitted from one cell to another. Bacteriophages infect bacterial cells and either replicate to large numbers and cause the cell to lyse (lytic infection) or, in some cases, integrate into the host genome without killing the host (the lysogenic state), such as the E. coli bacteriophage lambda. Some lysogenic bacteriophages carry toxin genes (e.g., corynephage beta carries the gene for the diphtheria toxin). Bacteriophage lambda remains lysogenic as long as a repressor protein is synthesized and prevents the phage genome from becoming unintegrated in order to replicate and exit the cell. Damage to the host cell DNA by radiation or by another means or inability to produce the repressor protein is a signal that the host cell is unhealthy and is no longer a good place for “freeloading.”
Transposons (jumping genes) are mobile genetic elements (Figure 13-14) that can transfer DNA within a cell, from one position to another in the genome, or between different molecules of DNA (e.g., plasmid to plasmid or plasmid to chromosome). Transposons are present in prokaryotes and eukaryotes. The simplest transposons are called insertion sequences and range in length from 150 to 1500 base pairs, with inverted repeats of 15 to 40 base pairs at their ends and the minimal genetic information necessary for their own transfer (i.e., the gene coding for the transposase). Complex transposons carry other genes, such as genes that provide resistance against antibiotics. Transposons sometimes insert into genes and inactivate those genes. If insertion and inactivation occur in a gene that encodes an essential protein, the cell dies.
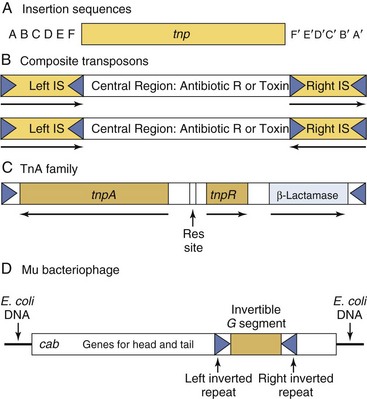
Figure 13-14 Transposons. A, The insertion sequences code only for a transposase (tnp) and possess inverted repeats (15 to 40 base pairs) at each end. B, The composite transposons contain a central region coding for antibiotic resistances or toxins flanked by two insertion sequences (IS), which can be either directly repeated or reversed. C, Tn3, a member of the TnA transposon family. The central region encodes three genes—a transposase (tnpA), a resolvase (tnpR), and a β-lactamase—conferring resistance to ampicillin. A resolution site (Res site) is used during the replicative transposition process. This central region is flanked on both ends by direct repeats of 38 base pairs. D, Phage-associated transposon is exemplified by the bacteriophage mu.
Some pathogenic bacteria use a transposon-like mechanism to coordinate the expression of a system of virulence factors. The genes for the activity may be grouped together in a pathogenicity or virulence island, which is surrounded by transposon-like mobile elements, allowing them to move within the chromosome and to other bacteria. The entire genetic unit can be triggered by an environmental stimulus (e.g., pH, heat, contact with the host cell surface) as a way to coordinate the expression of a complex process. For example, the SPI-1 island of Salmonella encodes 25 genes for a type III secretion device that allow the bacteria to enter nonphagocytic cells.
Mechanisms of Genetic Transfer between Cells
The exchange of genetic material between bacterial cells may occur by one of three mechanisms (Figure 13-15): (1) conjugation, which is the mating or quasisexual exchange of genetic information from one bacterium (the donor) to another bacterium (the recipient); (2) transformation, which is an active uptake and incorporation of exogenous or foreign DNA; or (3) transduction, which is the transfer of genetic information from one bacterium to another by a bacteriophage. Once inside a cell, a transposon can jump between different DNA molecules (e.g., plasmid to plasmid or plasmid to chromosome).
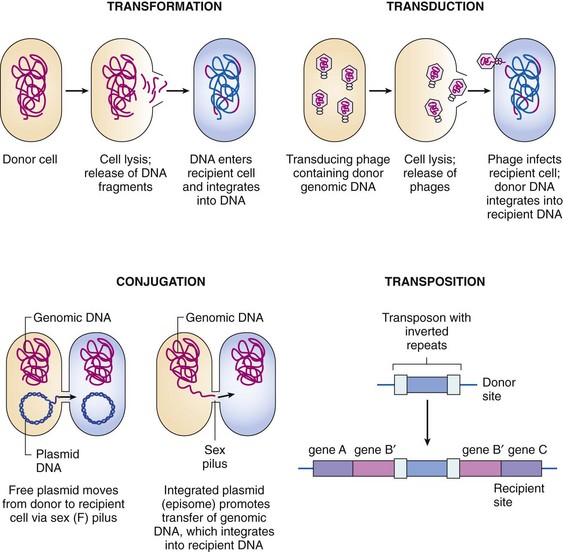
Figure 13-15 Mechanisms of bacterial gene transfer.
(From Rosenthal KS, Tan J: Rapid reviews microbiology and immunology, St Louis, 2002, Mosby.)
Transformation
Transformation is the process by which bacteria take up fragments of naked DNA and incorporate them into their genomes. Transformation was the first mechanism of genetic transfer to be discovered in bacteria. In 1928, Griffith observed that pneumococcus virulence was related to the presence of a polysaccharide capsule and that extracts of encapsulated bacteria producing smooth colonies could transmit this trait to nonencapsulated bacteria, normally appearing with rough edges. Griffith’s studies led to Avery, MacLeod, and McCarty’s identification of DNA as the transforming principle some 15 years later.
Gram-positive and gram-negative bacteria can take up and stably maintain exogenous DNA. Certain species are naturally capable of taking up exogenous DNA (such species are then said to be competent), including Haemophilus influenzae, Streptococcus pneumoniae, Bacillus species, and Neisseria species. Competence develops toward the end of logarithmic growth. E. coli and most other bacteria lack the natural ability for DNA uptake, and competence must be induced or chemical methods or electroporation (the use of high-voltage pulses), used to facilitate uptake of plasmid and other DNA.
Conjugation
Conjugation results in one-way transfer of DNA from a donor (or male) cell to a recipient (or female) cell through the sex pilus. Conjugation occurs with most, if not all, eubacteria, usually between members of the same or related species, but it has also been demonstrated to occur between prokaryotes and cells from plants, animals, and fungi. Many of the large conjugative plasmids specify colicins or antibiotic resistance.
The mating type (sex) of the cell depends on the presence (male) or absence (female) of a conjugative plasmid, such as the F plasmid of E. coli. The F plasmid is defined as conjugative because it carries all the genes necessary for its own transfer, including the ability to make sex pili and to initiate DNA synthesis at the transfer origin (oriT) of the plasmid. The sex pili is a specialized type IV secretion device. Upon transfer of the F plasmid, the recipients become F+ male cells. If a fragment of chromosomal DNA has been incorporated into the plasmid, it is designated an F prime (F′) plasmid. When it transfers into the recipient cell, it carries that fragment with it and converts it into an F′ male. If the F plasmid sequence is integrated into the bacterial chromosome, the cell is designated an Hfr (high-frequency recombination) cell.
The DNA that is transferred by conjugation is not a double helix but a single-stranded molecule. Mobilization begins when a plasmid-encoded protein makes a single-stranded, site-specific cleavage at the oriT. The nick initiates rolling circle replication, and the displaced linear strand is directed to the recipient cell. The transferred single-stranded DNA is recircularized and its complementary strand synthesized. Conjugation results in transfer of a part of the plasmid sequence and some portion of the bacterial chromosomal DNA. Because of the fragile connection between the mating pairs, the transfer is usually aborted before being completed such that only the chromosomal sequences adjacent to the integrated F are transferred. Artificial interruption of a mating between an Hfr and an F− pair has been helpful in constructing a consistent map of the E. coli chromosomal DNA. In such maps, the position of each gene is given in minutes (based on 100 minutes for complete transfer at 37° C), according to its time of entry into a recipient cell in relation to a fixed origin.
Transduction
Genetic transfer by transduction is mediated by bacterial viruses (bacteriophages), which pick up fragments of DNA and package them into bacteriophage particles. The DNA is delivered to infected cells and becomes incorporated into the bacterial genomes. Transduction can be classified as specialized if the phages in question transfer particular genes (usually those adjacent to their integration sites in the genome) or generalized if incorporation of DNA sequences is random because of accidental packaging of host DNA into the phage capsid. For example, a nuclease from the P1 phage degrades the host E. coli chromosomal DNA, and some of the DNA fragments are packaged into phage particles. The encapsulated DNA, instead of phage DNA, is injected into a new host cell, where it can recombine with the homologous host DNA. Generalized transducing particles are valuable in the genetic mapping of bacterial chromosomes. The closer two genes are within the bacterial chromosome, the more likely it is that they will be co-transduced in the same fragment of DNA.
Recombination
Incorporation of extrachromosomal (foreign) DNA into the chromosome occurs by recombination. There are two types of recombination: homologous and nonhomologous. Homologous (legitimate) recombination occurs between closely related DNA sequences and generally substitutes one sequence for another. The process requires a set of enzymes produced (in E. coli) by the rec genes. Nonhomologous (illegitimate) recombination occurs between dissimilar DNA sequences and generally produces insertions or deletions or both. This process usually requires specialized (sometimes site-specific) recombination enzymes, such as those produced by many transposons and lysogenic bacteriophages.
Generation of Vancomycin-Resistant Staphylococcus aureus by Multiple Genetic Manipulations
Until recently, vancomycin was the last-resort drug for S. aureus strains resistant to β-lactam (penicillin-related) antibiotics (e.g., methicillin-resistant S. aureus [MRSA]). S. aureus acquired the vancomycin resistance gene during a mixed infection with Enterococcus faecalis (Figure 13-16). The gene for vancomycin resistance was contained within a transposon (Tn1546) on a multiresistance conjugative plasmid. The plasmid was probably transferred by conjugation between E. faecalis and S. aureus. Alternatively, after lysis of the E. faecalis, S. aureus acquired the DNA by transduction and became transformed by the new DNA. The transposon then jumped from the E. faecalis plasmid, recombined, and integrated into the S. aureus multiresistance plasmid, and the E. faecalis DNA was degraded. The resulting S. aureus plasmid encodes resistance to β-lactams, vancomycin, trimethoprim, and gentamycin/kanamycin/tobramycin antibiotics and to quaternary ammonium disinfectants and can transfer to other S. aureus strains by conjugation. (For more information, refer to Weigel in the Bibliography at the end of the chapter.)
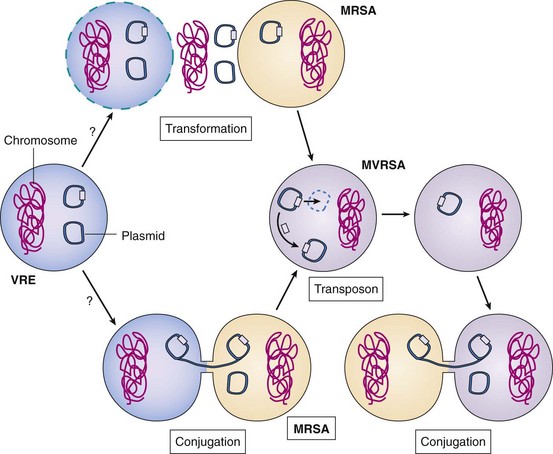
Figure 13-16 Genetic mechanisms of evolution of methicillin- and vancomycin-resistant Staphylococcus aureus (MRSA and MVRSA). Vancomycin-resistant enterococcus (VRE) (in red) contains plasmids with multiple antibiotic resistance and virulence factors. During co-infection, a MRSA may have acquired the enterococcal resistance plasmid (e-plasmid) by transformation (after lysis of the enterococcal cell and release of its DNA) or more likely, by conjugation. A transposon in the e-plasmid containing the vancomycin resistance gene jumped out and inserted into the multiple antibiotic resistance plasmid of the MRSA. The new plasmid is readily spread to other S. aureus bacteria by conjugation.
Genetic Engineering
Genetic engineering, also known as recombinant DNA technology, uses the techniques and tools developed by the bacterial geneticists to purify, amplify, modify, and express specific gene sequences. The use of genetic engineering and “cloning” has revolutionized biology and medicine. The basic components of genetic engineering are (1) cloning and expression vectors, which can be used to deliver the DNA sequences into receptive bacteria and amplify the desired sequence, (2) the DNA sequence to be amplified and expressed, (3) enzymes, such as restriction enzymes, which are used to cleave DNA reproducibly at defined sequences (Table 13-1), and (4) DNA ligase, the enzyme that links the fragment to the cloning vector.
Table 13-1 Common Restriction Enzymes Used in Molecular Biology
| Microorganism | Enzyme | Recognition Site |
|---|---|---|
| Acinetobacter calcoaceticus | Acc I |
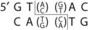 |
| Bacillus amyloliquefaciens H | Bam HI |
 |
| Escherichia coli RY13 | Eco RI |
 |
| Haemophilus influenzae Rd | Hind III |
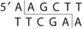 |
| H. influenzae serotype c, 1160 | Hinc II |
 |
| Providencia stuartii 164 | Pst I |
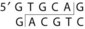 |
| Serratia marcescens | Sma I |
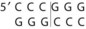 |
| Staphylococcus aureus 3A | Sau 3AI |
 |
| Xanthomonas malvacearum | Xma I |
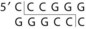 |
Cloning and expression vectors must allow foreign DNA to be inserted into them, but still must be able to replicate normally in a bacterial or eukaryotic host. Many types of vectors are currently used. Plasmid vectors, such as pUC, pBR322, and pGEM (Figure 13-17), are used for DNA fragments up to 20 kb. Bacteriophages, such as lambda, are used for larger fragments up to 25 kb, and cosmid vectors have combined some of the advantages of plasmids and phages for fragments up to 45 kb.
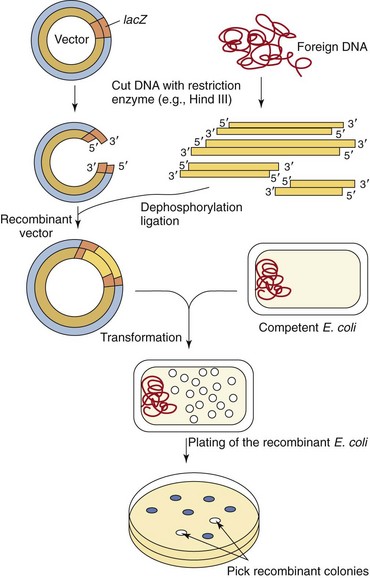
Figure 13-17 Cloning of foreign DNA in vectors. The vector and the foreign DNA are first digested by a restriction enzyme. Insertion of foreign DNA into the lacZ gene inactivates the β-galactosidase gene, allowing subsequent selection. The vector is then ligated to the foreign DNA, using bacteriophage T4 DNA ligase. The recombinant vectors are transformed into competent Escherichia coli cells. The recombinant E. coli cells are plated onto agar containing antibiotic, an inducer of the lac operon, and a chromophoric substrate that turns blue in cells having a plasmid but not an insert; those cells with a plasmid containing the insert remain white.
Most cloning vectors have been “engineered” to have a site for insertion of foreign DNA, a means of selection of the bacteria that have incorporated any plasmid (e.g., antibiotic resistance), and a means of distinguishing the bacteria that have incorporated those plasmids that contain inserted DNA. Expression vectors have DNA sequences to facilitate their replication in bacteria and eukaryotic cells and also the transcription of the gene into mRNA.
The DNA to be cloned can be obtained by purification of chromosomal DNA from cells, viruses, or other plasmids or by selective amplification of DNA sequences by a technique known as polymerase chain reaction (PCR) (PCR is explained further in Chapter 5). Both the vector and the foreign DNA are cleaved with restriction enzymes (see Figure 13-17). Restriction enzymes recognize a specific palindromic sequence and make a staggered cut, which generates sticky ends, or a blunt cut, which generates blunt ends (see Table 13-1). Most cloning vectors have a sequence called the multiple cloning site that can be cleaved by many restriction enzymes. Ligation of the vector with the DNA fragments generates a molecule called recombinant DNA, which is capable of replicating the inserted sequence. The total number of recombinant vectors obtained when cloning all the fragments that result from cleavage of chromosomal DNA is known as a genomic library because there should be at least one representative of each gene in the library. An alternative approach to cloning the gene for a protein is to use a retrovirus enzyme called reverse transcriptase (RNA-dependent DNA polymerase) to convert the mRNA in the cell into DNA to produce a complementary DNA (cDNA). A cDNA library represents the genes that are expressed as mRNA in a particular cell.
The recombinant DNA is then transformed into a bacterial host, usually E. coli, and the plasmid-containing bacteria are selected for antibiotic resistance (e.g., ampicillin resistance). The library can then be screened to find an E. coli clone possessing the desired DNA fragment. Various screening techniques can be used to identify the bacteria containing the appropriate recombinant DNA. The multiple cloning site used for inserting the foreign DNA is often part of the lacZ gene of the lac operon. Insertion of the foreign DNA into the lacZ gene inactivates the gene (acting almost like a transposon) and prevents the plasmid-directed synthesis of β-galactosidase in the recipient cell, which results in white bacterial colonies instead of blue colonies if β-galactosidase were produced and able to cleave an appropriate chromophore.
Genetic engineering has been used to isolate and express the genes for useful proteins, such as insulin, interferon, growth hormones, and interleukin in bacteria, yeast, or even insect cells. Similarly, large amounts of pure immunogen for a vaccine can be prepared without the need to work with the intact disease organisms.
The vaccine against hepatitis B virus represents the first successful use of recombinant DNA technology to make a vaccine approved for human use by the U.S. Food and Drug Administration. The hepatitis B surface antigen is produced by the yeast Saccharomyces cerevisiae. In the future, it may be sufficient to inject plasmid DNA capable of expressing the desired immunogen (DNA vaccine) into an individual to let the host cells express the immunogen and generate the immune response. Recombinant DNA technology has also become essential to laboratory diagnosis, forensic science, agriculture, and many other disciplines.
1. How many moles of ATP are generated per mole of glucose in glycolysis, the TCA cycle, and electron transport? Which of these occur in anaerobic conditions and in aerobic conditions? Which is most efficient?
2. What products of anaerobic fermentation would be detrimental to host (human) tissue (e.g., C. perfringens)?
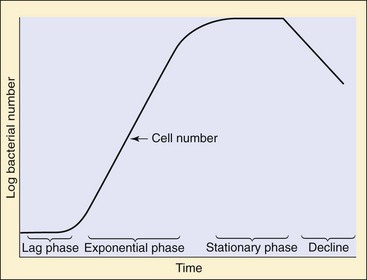
3. If the number of bacteria during log phase growth can be calculated by the following equation:
in which Nt is the number of bacteria after time (t), t/d is the amount of time divided by the doubling time, and N0 is the initial number of bacteria, how many bacteria will be in the culture after 4 hours if the doubling time is 20 minutes and the initial bacterial inoculum contained 1000 bacteria?
4. What are the principal properties of a plasmid?
5. Give two mechanisms of regulation of bacterial gene expression. Use specific examples.
6. What types of mutations affect DNA, and what agents are responsible for such mutations?
7. Which mechanisms may be used by a bacterial cell for the exchange of genetic material? Briefly explain each mechanism.
8. Discuss the applications of molecular biotechnology to medicine, including contributions and uses in diagnoses.
1. Glycolysis: During fermentation, each mole of glucose yields two moles of ATP and two moles of NADH. Conversion of pyruvate to acetyl-CoA produces two more NADH.
TCA cycle, two moles GTP (equivalent to ATP) are produced plus two moles FADH2 and six moles NADH, which are fed into the electron transport system.
Electron transport: The 2 FADH2 (4 ATP) and 6 NADH (18 ATP) plus the 2 GTP (equivalent to 2 ATP) from the TCA cycle plus the 2 NADH (6 ATP) from gycolysis and the 2 NADH (6 ATP) from conversion of pyruvate to Acetyl-CoA and the 2 ATP from glycolysis add up to 38 ATP.
Anaerobic conditions: Glycolysis occurs in a process called fermentation. This is not an efficient process.
Aerobic conditions: Glycolysis, TCA cycle, and electron transport occur under aerobic conditions. This is the most efficient process for conversion of glucose to energy.
2. Anaerobic fermentation produces acids, CO2, and, sometimes, methane. The detrimental effect of these actions is seen in gas gangrene.
3. Nt = 1000 × 2480 min/20 min
4. A plasmid is extrachromosomal, circular DNA with an origin of replication (allows replication) and often contains genes for antibiotic resistance, metabolism of unusual molecules (e.g., Pseudomonas) or virulence.
5. Repression: A repressor binds to a site on the Lac operon to prevent expression of the gene unless lactose is present. Binding of lactose to the repressor causes it to dissociate from the DNA and allows expression.
Induction: The CAP binds cAMP to form a complex that enhances gene expression. cAMP is produced when levels of glucose are depleted to indicate a metabolic problem. This would enhance the expression of the lac operon in the presence of galactose.
Attenuation: Translation of a protein can regulate the transcription of the gene because there is no nuclear membrane to separate these processes. The amount of tryptophan in a cell will determine the rate of synthesis of a test mRNA and peptide, which will determine whether the mRNA forms a hairpin loop. The loop will stop transcription.
• Transversion: pyrimidine 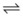 purine
purine
• Missense: change in amino acid in protein
• Nonsense: change codon to insert a stop codon into the protein
• Frameshift: inserts or deletes of one or two nucleotides to disrupt the reading of three nucleotide codons
• Null: destroys protein function (e.g., nonsense, frameshift)
7. Transformation: acquisition of DNA from the extracellular space, which becomes part of the chromatin
Transduction: infection by a bacteriophage that has acquired DNA sequences from another bacteria
Conjugation: transfer of DNA via a sex-pilus
Transposition: acquisition of a transposon that inserts into the chromosome
8. Genetic engineering has been used to isolate genes for hormones (e.g., growth hormone, insulin), viral genes for vaccines (e.g., hepatitis B virus), and cytokine genes (e.g., interferon-α, interferon-γ). These genes have been cloned into plasmids and expressed in large quantities to produce these proteins as drugs. In addition, DNA vaccines have been prepared in which viral or other genes are inserted into plasmids that can be expressed in mammalian cells. Expression of the gene and its protein in the vaccinated person will lead to the development of an immune response.
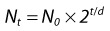
 purine
purine