44 Viral Classification, Structure, and Replication
Viruses were first described as “filterable agents.” Their small size allows them to pass through filters designed to retain bacteria. Unlike most bacteria, fungi, and parasites, viruses are obligate intracellular parasites that depend on the biochemical machinery of the host cell for replication. In addition, reproduction of viruses occurs by assembly of the individual components rather than by binary fission (Boxes 44-1 and 44-2).
Box 44-1
Definition and Properties of a Virus
Viruses are filterable agents.
Viruses are obligate intracellular parasites.
Viruses cannot make energy or proteins independently of a host cell.
Viral genomes may be RNA or DNA but not both.
Viruses have a naked capsid or an envelope morphology.
Viral components are assembled and do not replicate by “division.”
Box 44-2
Consequences of Viral Properties
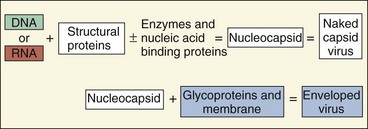
The simplest viruses consist of a genome of deoxyribonucleic acid (DNA) or ribonucleic acid (RNA) packaged in a protective shell of protein and, for some viruses, a membrane (Figure 44-1). Viruses lack the capacity to make energy or substrates, cannot make their own proteins, and cannot replicate their genome independently of the host cell. To use the cell’s biosynthetic machinery, the virus must be adapted to the biochemical rules of the cell.
The physical structure and genetics of viruses have been optimized by mutation and selection to infect humans and other hosts. To do this, the virus must be capable of transmission through potentially harsh environmental conditions, must traverse the skin or other protective barriers of the host, must be adapted to the biochemical machinery of the host cell for replication, and must escape elimination by the host immune response.
Knowledge of the structural (size and morphology) and genetic (type and structure of nucleic acid) features of a virus provides insight into how the virus replicates, spreads, and causes disease. The concepts presented in this chapter are repeated in greater detail in the discussions of specific viruses in later chapters.
Classification
Viruses range from the structurally simple and small parvoviruses and picornaviruses to the large and complex poxviruses and herpesviruses. Their names may describe viral characteristics, the diseases they are associated with, or even the tissue or geographic locale where they were first identified. Names such as picornavirus (pico, “small”; rna, “ribonucleic acid”) or togavirus (toga, Greek for “mantle,” referring to a membrane envelope surrounding the virus) describe the structure of the virus. The name retrovirus (retro, “reverse”) refers to the virus-directed synthesis of DNA from an RNA template, whereas the poxviruses are named for the disease smallpox, caused by one of its members. The adenoviruses (adenoids) and the reoviruses (respiratory, enteric, orphan) are named for the body site from which they were first isolated. Reovirus was discovered before it was associated with a specific disease, and thus it was designated an “orphan” virus. Norwalk virus is named for Norwalk, Ohio; coxsackievirus is named for Coxsackie, New York; and many of the togaviruses, arenaviruses, and bunyaviruses are named after African places where they were first isolated.
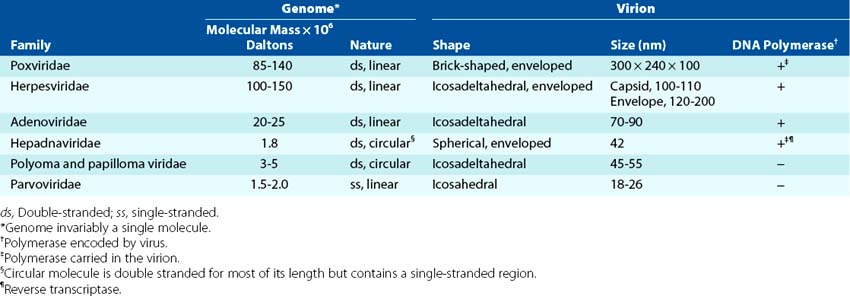
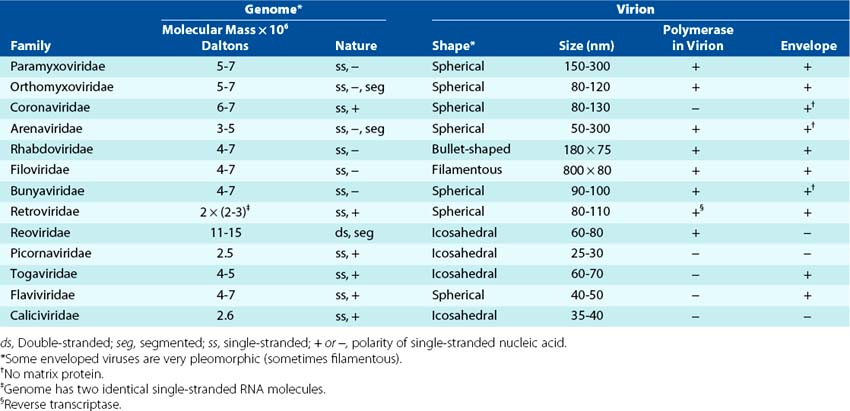
Viruses can be grouped by characteristics such as disease (e.g., hepatitis), target tissue, means of transmission (e.g., enteric, respiratory), or vector (e.g., arboviruses; arthropod-borne virus) (Box 44-3). The most consistent and current means of classification is by physical and biochemical characteristics, such as size, morphology (e.g., presence or absence of a membrane envelope), type of genome, and means of replication (Figures 44-2 and 44-3). DNA viruses associated with human disease are divided into seven families (Tables 44-1 and 44-2). The RNA viruses may be divided into at least 13 families (Tables 44-3 and 44-4).
Box 44-3
Means of Classification and Naming of Viruses
Structure: size, morphology, and nucleic acid (e.g., picornavirus [small RNA], togavirus)
Biochemical characteristics: structure and mode of replication*
Disease: encephalitis and hepatitis viruses, for example
Means of transmission: arbovirus spread by insects, for example
Host cell (host range): animal (human, mouse, bird), plant, bacteria
Tissue or organ (tropism): adenovirus and enterovirus, for example
* This is the current means of taxonomic classification of viruses.
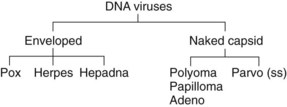
Figure 44-2 The DNA viruses and their morphology. The viral families are determined by the structure of the genome and the morphology of the virion.
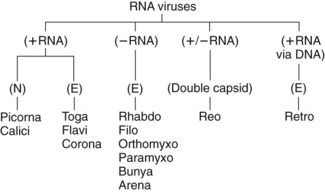
Figure 44-3 The RNA viruses, their genome structure, and their morphology. The viral families are determined by the structure of the genome and the morphology of the virion. E, Enveloped; N, naked capsid.
Table 44-1 Families of DNA Viruses and Some Important Members
| Family | Members* |
|---|---|
| POXVIRIDAE† | Smallpox virus, vaccinia virus, monkeypox, canarypox, molluscum contagiosum |
| Herpesviridae | Herpes simplex virus types 1 and 2, varicella-zoster virus, Epstein-Barr virus, cytomegalovirus, human herpesviruses 6, 7, and 8 |
| Adenoviridae | Adenovirus |
| Hepadnaviridae | Hepatitis B virus |
| Papillomaviridae | Papilloma virus |
| Polyomaviridae | JC virus, BK virus, SV40 |
| Parvoviridae | Parvovirus B19, adeno-associated virus |
* The italicized virus is the prototype virus for the family.
† The size of type is indicative of the relative size of the virus.
Table 44-3 Families of RNA Viruses and Some Important Members
| Family* | Members† |
|---|---|
| PARAMYXOVIRIDAE | Parainfluenza virus, Sendai virus, measles virus, mumps virus, respiratory syncytial virus, metapneumovirus |
| ORTHOMYXOVIRIDAE | Influenza virus types A, B, and C |
| CORONAVIRIDAE | Coronavirus, severe acute respiratory syndrome |
| Arenaviridae | Lassa fever virus, Tacaribe virus complex (Junin and Machupo viruses), lymphocytic choriomeningitis virus |
| Rhabdoviridae | Rabies virus, vesicular stomatitis virus |
| Filoviridae | Ebola virus, Marburg virus |
| Bunyaviridae | California encephalitis virus, La Crosse virus, sandfly fever virus, hemorrhagic fever virus, Hanta virus |
| Retroviridae | Human T-cell leukemia virus types I and II, human immunodeficiency virus, animal oncoviruses |
| Reoviridae | Rotavirus, Colorado tick fever virus |
| Togaviridae | Rubella virus; western, eastern, and Venezuelan equine encephalitis virus; Ross River virus; Sindbis virus; Semliki Forest virus |
| Flaviviridae | Yellow fever virus, dengue virus, St. Louis encephalitis virus, West Nile virus, hepatitis C virus |
| Caliciviridae | Norwalk virus, calicivirus |
| Picornaviridae | Rhinoviruses, poliovirus, echoviruses, coxsackievirus, hepatitis A virus |
| Delta | Delta agent |
* The size of the type is indicative of the relative size of the virus.
† The italicized virus is the prototype virus for the family.
Virion Structure
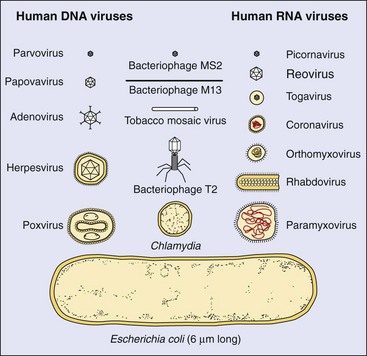
The units for measurement of virion size are nanometers (nm). The clinically important viruses range from 18 nm (parvoviruses) to 300 nm (poxviruses) (Figure 44-4). The latter are almost visible with a light microscope and are approximately one fourth the size of staphylococcal bacteria. Larger virions can hold a larger genome that can encode more proteins, and they are generally more complex.
The virion (the virus particle) consists of a nucleic acid genome packaged into a protein coat (capsid) or a membrane (envelope) (Figure 44-5). The virion may also contain certain essential or accessory enzymes or other proteins to facilitate initial replication in the cell. Capsid or nucleic acid–binding proteins may associate with the genome to form a nucleocapsid, which may be the same as the virion or surrounded by an envelope.
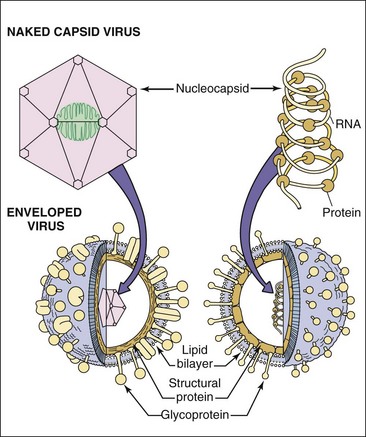
Figure 44-5 The structures of a naked capsid virus (top left) and enveloped viruses (bottom) with an icosahedral (left) nucleocapsid or a helical (right) ribonucleocapsid. The helical ribonucleocapsid is formed by viral proteins associated with an RNA genome.
The genome of the virus consists either of DNA or RNA. The DNA can be single or double stranded, linear or circular. The RNA can be either positive sense (+) (like messenger RNA [mRNA]) or negative sense (−) (analogous to a photographic negative), double stranded (+/−) or ambisense (containing + and − regions of RNA attached end to end). The RNA genome may also be segmented into pieces, with each piece encoding one or more genes. Just as there are many different types of computer memory devices, all of these forms of nucleic acid can maintain and transmit the genetic information of the virus. Similarly, the larger the genome, the more information (genes) it can carry and the larger the capsid or envelope structure required to contain the genome.
The outer layer of the virion is the capsid or envelope. These structures are the package, protection, and delivery vehicle during transmission of the virus from one host to another and for spread within the host to the target cell. The surface structures of the capsid and envelope mediate the interaction of the virus with the target cell through a viral attachment protein (VAP) or structure. Removal or disruption of the outer package inactivates the virus. Antibodies generated against the components of these structures prevent virus infection.
The capsid is a rigid structure able to withstand harsh environmental conditions. Viruses with naked capsids are generally resistant to drying, acid, and detergents, including the acid and bile of the enteric tract. Many of these viruses are transmitted by the fecal-oral route and can endure transmission even in sewage.
The envelope is a membrane composed of lipids, proteins, and glycoproteins. The membranous structure of the envelope can be maintained only in aqueous solutions. It is readily disrupted by drying, acidic conditions, detergents, and solvents such as ether, which results in inactivation of the virus. As a result, enveloped viruses must remain wet and are generally transmitted in fluids, respiratory droplets, blood, and tissue. Most cannot survive the harsh conditions of the gastrointestinal tract. The influence of virion structure on viral properties is summarized in Boxes 44-4 and 44-5.
Box 44-5 Virion Structure
Envelope
Properties*
Consequences*
Cannot survive the gastrointestinal tract
Spreads in large droplets, secretions, organ transplants, and blood transfusions
Does not need to kill the cell to spread
May need antibody and cell-mediated immune response for protection and control
Elicits hypersensitivity and inflammation to cause immunopathogenesis
* Exceptions exist.
Capsid Viruses
The viral capsid is assembled from individual proteins associated into progressively larger units. All of the components of the capsid have chemical features that allow them to fit together and to assemble into a larger unit. Individual structural proteins associate into subunits, which associate into protomers, capsomeres (distinguishable in electron micrographs), and finally, a recognizable procapsid or capsid (Figure 44-6). A procapsid requires further processing to the final, transmissible capsid. For some viruses, the capsid forms around the genome; for others the capsid forms as an empty shell (procapsid) to be filled by the genome.
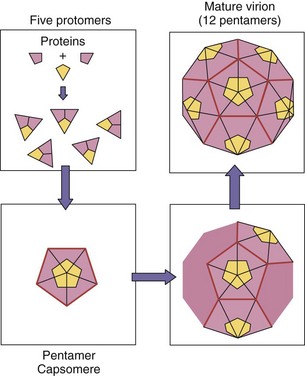
Figure 44-6 Capsid assembly of the icosahedral capsid of a picornavirus. Individual proteins associate into subunits, which associate into protomers, capsomeres, and an empty procapsid. Inclusion of the (+) RNA genome triggers its conversion to the final capsid form.
The simplest viral structures that can be built stepwise are symmetric and include helical and icosahedral structures. Helical structures appear as rods, whereas the icosahedron is an approximation of a sphere assembled from symmetric subunits (Figure 44-7). Nonsymmetric capsids are complex forms and are associated with certain bacterial viruses (phages).
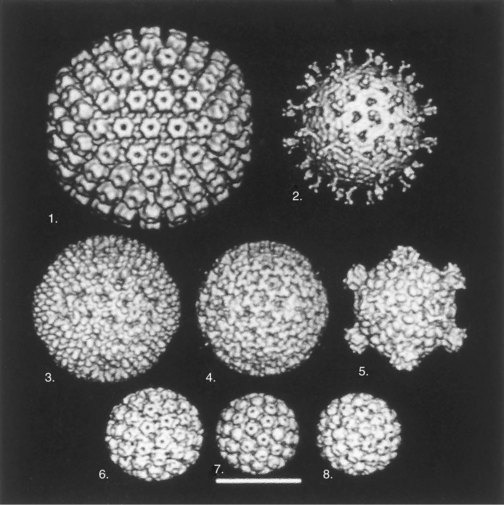
Figure 44-7 Cryoelectron microscopy and computer-generated three-dimensional image reconstructions of several icosahedral capsids. These images show the symmetry of capsids and the individual capsomeres. During assembly, the genome may fill the capsid through the holes in the herpesvirus and papovavirus capsomeres. 1, Equine herpesvirus nucleocapsid; 2, simian rotavirus; 3, reovirus type 1 (Lang) virion; 4, intermediate subviral particle (reovirus); 5, core (inner capsid) particle (reovirus); 6, human papillomavirus type 19; 7, mouse polyomavirus; 8, cauliflower mosaic virus. Bar = 50 nm.
(Courtesy Dr. Tim Baker, Purdue University, West Lafayette, Ind.)
The classic example of a virus with helical symmetry is the tobacco mosaic plant virus. Its capsomeres self-assemble on the RNA genome into rods that extend the length of the genome. The capsomeres cover and protect the RNA. Helical nucleocapsids are observed within the envelope of most negative-strand RNA viruses (see Figure 56-1).
Simple icosahedrons are used by small viruses such as the picornaviruses and parvoviruses. The icosahedron is made of 12 capsomeres, each with fivefold symmetry (pentamer or penton). For the picornaviruses, every pentamer is made up of five protomers, each of which is composed of three subunits of four separate proteins (see Figure 44-6). X-ray crystallography and image analysis of cryoelectron microscopy have defined the structure of the picornavirus capsid to the molecular level. These studies have depicted a canyon-like cleft, which is a “docking site” to bind to the receptor on the surface of the target cell (see Figure 54-2).
Larger capsid virions are constructed by inserting structurally distinct capsomeres between the pentons at the vertices. These capsomeres have six nearest neighbors (hexons). This extends the icosahedron and is called an icosadeltahedron, and its size is determined by the number of hexons inserted along the edges and within the surfaces between the pentons. A soccer ball is an icosadeltahedron. For example, the herpesvirus nucleocapsid has 12 pentons and 150 hexons. The herpesvirus nucleocapsid is also surrounded by an envelope. The adenovirus capsid is composed of 252 capsomeres, with 12 pentons and 240 hexons. A long fiber is attached to each penton of adenovirus to serve as the VAP to bind to target cells, and it also contains the type-specific antigen (see Figure 50-1). The reoviruses have an icosahedral double capsid with fiber-like proteins partially extended from each vertex. The outer capsid protects the virus and promotes its uptake across the gastrointestinal tract and into target cells, whereas the inner capsid contains enzymes for the synthesis of RNA (see Figures 44-7 and 59-2).
Enveloped Viruses
The virion envelope is composed of lipids, proteins, and glycoproteins (see Figure 44-5 and Box 44-5). It has a membrane structure similar to cellular membranes. Cellular proteins are rarely found in the viral envelope, even though the envelope is obtained from cellular membranes. Most enveloped viruses are round or pleomorphic (see Figures 44-2 and 44-3 for the complete listing of enveloped viruses). Two exceptions are the poxvirus, which has a complex internal and a bricklike external structure, and the rhabdovirus, which is bullet shaped.
Most viral glycoproteins have asparagine-linked (N-linked) carbohydrate and extend through the envelope and away from the surface of the virion. For many viruses, these can be observed as spikes (Figure 44-8). Some glycoproteins act as VAPs, capable of binding to structures on target cells. VAPs that also bind to erythrocytes are termed hemagglutinins (HAs). Some glycoproteins have other functions, such as the neuraminidase (NA) of orthomyxoviruses (influenza) and the Fc receptor and the C3b receptor associated with herpes simplex virus (HSV) glycoproteins, or the fusion glycoproteins of paramyxoviruses. Glycoproteins, especially the VAP, are also major antigens that elicit protective immunity.
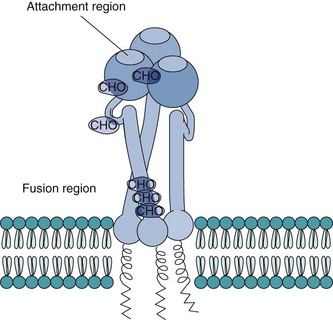
Figure 44-8 Diagram of the hemagglutinin glycoprotein trimer of influenza A virus, a representative spike protein. The region for attachment to the cellular receptor is exposed on the spike protein’s surface. Under mild acidic conditions, the hemagglutinin folds over to bring the virion envelope and cellular membrane together and exposes a hydrophobic sequence to promote fusion. CHO, N-linked carbohydrate attachment sites.
(Modified from Schlesinger MJ, Schlesinger S: Domains of virus glycoproteins, Adv Virus Res 33:1–44, 1987.)
The envelope of the togaviruses surrounds an icosahedral nucleocapsid containing a positive-strand RNA genome. The envelope contains spikes consisting of two or three glycoprotein subunits anchored to the virion’s icosahedral capsid. This causes the envelope to adhere tightly and conform (shrink-wrap) to an icosahedral structure discernible by cryoelectron microscopy.
All of the negative-strand RNA viruses are enveloped. Components of the viral RNA-dependent RNA polymerase associate with the (−) RNA genome of the orthomyxoviruses, paramyxoviruses, and rhabdoviruses to form helical nucleocapsids (see Figure 44-5). These enzymes are required to initiate virus replication, and their association with the genome ensures their delivery into the cell. Matrix proteins lining the inside of the envelope facilitate the assembly of the ribonucleocapsid into the virion. Influenza A (orthomyxovirus) is an example of a (−) RNA virus with a segmented genome. Its envelope is lined with matrix proteins and has two glycoproteins: the HA, which is the VAP, and an NA (see Figure 57-1). Bunyaviruses do not have matrix proteins.
The herpesvirus envelope is a baglike structure that encloses the icosadeltahedral nucleocapsid (see Figure 51-1). Depending on the specific herpesvirus, the envelope may contain as many as 11 glycoproteins. The interstitial space between the nucleocapsid and the envelope is called the tegument, and it contains enzymes, other proteins, and even RNA that facilitate the viral infection.
The poxviruses are enveloped viruses with large, complex, bricklike shapes (see Figure 52-1). The envelope encloses a dumbbell-shaped, DNA-containing nucleoid structure; lateral bodies; fibrils; and many enzymes and proteins, including the enzymes and transcriptional factors required for mRNA synthesis.
Viral Replication
The major steps in viral replication are the same for all viruses (Figure 44-9 and Box 44-6). The cell acts as a factory, providing the substrates, energy, and machinery necessary for the synthesis of viral proteins and replication of the genome. Processes not provided by the cell must be encoded in the genome of the virus. The manner in which each virus accomplishes these steps and overcomes the cell’s biochemical limitations is different for different structures of the genome and of the virion (whether it is enveloped or has a naked capsid). This is illustrated in the examples in Figures 44-12 to 44-14 (see later).
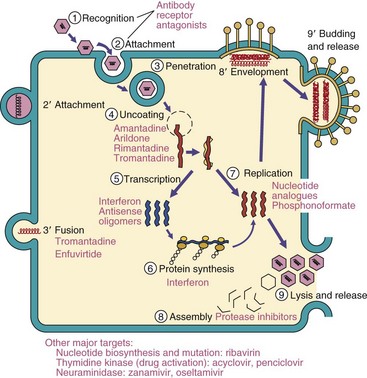
Figure 44-9 A general scheme of viral replication. Enveloped viruses have alternative means of entry (steps 2′ and 3′), assembly, and exit from the cell (8′ and 9′). The antiviral drugs for susceptible steps in viral replication are listed in magenta.
Box 44-6
Steps in Viral Replication
A single round of the viral replication cycle can be separated into several phases. During the early phase of infection, the virus must recognize an appropriate target cell, attach to the cell, penetrate the plasma membrane and be taken up by the cell, release (uncoat) its genome into the cytoplasm, and if necessary, deliver the genome to the nucleus. The late phase begins with the start of genome replication and viral macromolecular synthesis and proceeds through viral assembly and release. Uncoating of the genome from the capsid or envelope during the early phase abolishes its infectivity and identifiable structure, thus initiating the eclipse period. The eclipse period, like a solar eclipse, ends with the appearance of new virions after virus assembly. The latent period, during which extracellular infectious virus is not detected, includes the eclipse period and ends with the release of new viruses (Figure 44-10). Each infected cell may produce as many as 100,000 particles; however, only 1% to 10% of these particles may be infectious. The noninfectious particles (defective particles) result from mutations and errors in the manufacture and assembly of the virion. The yield of infectious virus per cell, or burst size, and the time required for a single cycle of virus reproduction are determined by the properties of the virus and the target cell.
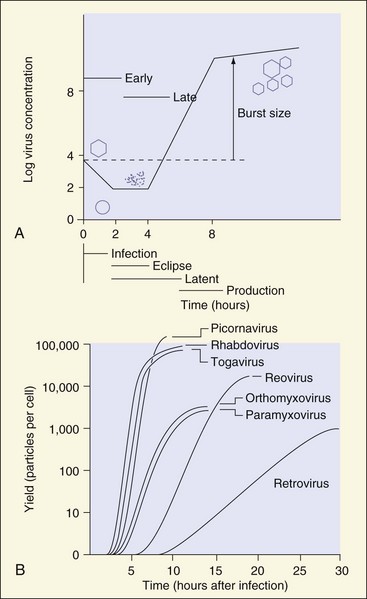
Figure 44-10 A, Single-cycle growth curve of a virus that is released on cell lysis. The different stages are defined by the absence of visible viral components (eclipse period), infectious virus in the media (latent period), or presence of macromolecular synthesis (early/late phases). B, Growth curve and burst size of representative viruses.
(A, Modified from Davis BD, et al: Microbiology, ed 4, Philadelphia, 1990, Lippincott; B, modified from White DO, Fenner F: Medical virology, ed 3, New York, 1986, Academic.)
Recognition of and Attachment to the Target Cell
The binding of the VAPs or structures on the surface of the virion capsid (Table 44-5) to receptors on the cell (Table 44-6) initially determines which cells can be infected by a virus. The receptors for the virus on the cell may be proteins or carbohydrates on glycoproteins or glycolipids. Viruses that bind to receptors expressed on specific cell types may be restricted to certain species (host range) (e.g., human, mouse) or specific cell types. The susceptible target cell defines the tissue tropism (e.g., neurotropic, lymphotropic). Epstein-Barr virus (EBV), a herpesvirus, has a very limited host range and tropism because it binds to the C3d receptor (CR2) expressed on human B cells. The B19 parvovirus binds to globoside (blood group P antigen) expressed on erythroid precursor cells.
Table 44-5 Examples of Viral Attachment Proteins
| Virus Family | Virus | Viral Attachment Protein |
|---|---|---|
| Picornaviridae | Rhinovirus | VP1-VP2-VP3 complex |
| Adenoviridae | Adenovirus | Fiber protein |
| Reoviridae | Reovirus | σ-1 |
| Rotavirus | VP7 | |
| Togaviridae | Semliki Forest virus | E1-E2-E3 complex gp |
| Rhabdoviridae | Rabies virus | G protein gp |
| Orthomyxoviridae | Influenza A virus | HA gp |
| Paramyxoviridae | Measles virus | HA gp |
| Herpesviridae | Epstein-Barr virus | gp350 and gp220 |
| Retroviridae | Murine leukemia virus | gp70 |
| Human immunodeficiency virus | gp120 |
gp, Glycoprotein; HA, hemagglutinin.
Table 44-6 Examples of Viral Receptors
| Virus | Target Cell | Receptor* |
|---|---|---|
| Epstein-Barr virus | B cell | C3d complement receptor CR2 (CD21) |
| Human immunodeficiency virus | Helper T cell | CD4 molecule and chemokine coreceptor |
| Rhinovirus | Epithelial cells | ICAM-1 (immunoglobulin superfamily protein) |
| Poliovirus | Epithelial cells | Immunoglobulin superfamily protein |
| Herpes simplex virus | Many cells | Herpesvirus entry mediator (HVEM), nectin-1 |
| Rabies virus | Neuron | Acetylcholine receptor, NCAM |
| Influenza A virus | Epithelial cells | Sialic acid |
| B19 parvovirus | Erythroid precursors | Erythrocyte P antigen (globoside) |
CD, Cluster of differentiation; ICAM-1, intercellular adhesion molecule; NCAM, neural cell adhesion molecule.
* Other receptors for these viruses may also exist.
The viral attachment structure for a capsid virus may be part of the capsid or a protein that extends from the capsid. A canyon on the surface of picornaviruses, such as the rhinovirus 14, serves as a “keyhole” for the insertion of a portion of the intercellular adhesion molecule (ICAM-1) from the cell surface. The fibers of the adenoviruses and the σ-1 proteins of the reoviruses at the vertices of the capsid interact with receptors expressed on specific target cells.
VAPs are specific glycoproteins of enveloped viruses. The HA of influenza A virus binds to sialic acid expressed on many different cells and has a broad host range and tissue tropism. Similarly, the α-togaviruses and the flaviviruses are able to bind to receptors expressed on cells of many animal species, including arthropods, reptiles, amphibians, birds, and mammals. This allows them to infect animals, mosquitoes, and other insects and to be spread by them.
Penetration
Interactions between multiple VAPs and cellular receptors initiate the internalization of the virus into the cell. The mechanism of internalization depends on the virion structure and cell type. Most nonenveloped viruses enter the cell by receptor-mediated endocytosis or by viropexis. Endocytosis is a normal process used by the cell for the uptake of receptor-bound molecules such as hormones, low-density lipoproteins, and transferrin. Picornaviruses and papovaviruses may enter by viropexis. Hydrophobic structures of capsid proteins may be exposed after viral binding to the cells, and these structures help the virus or the viral genome slip through (direct penetration) the membrane.
Enveloped viruses fuse their membranes with cellular membranes to deliver the nucleocapsid or genome directly into the cytoplasm. The optimum pH for fusion determines whether penetration occurs at the cell surface at neutral pH or whether the virus must be internalized by endocytosis, and fusion occurs in an endosome at acidic pH. The fusion activity may be provided by the VAP or another protein. The HA of influenza A (see Figure 44-8) binds to sialic acid receptors on the target cell. Under the mild acidic conditions of the endosome, the HA undergoes a dramatic conformational change to expose hydrophobic portions capable of promoting membrane fusion. Paramyxoviruses have a fusion protein that is active at neutral pH to promote virus-to-cell fusion. Paramyxoviruses can also promote cell-to-cell fusion to form multinucleated giant cells (syncytia). Some herpesviruses and retroviruses fuse with cells at a neutral pH and induce syncytia after replication.
Uncoating
Once internalized, the nucleocapsid must be delivered to the site of replication within the cell and the capsid or envelope removed. The genome of DNA viruses, except for poxviruses, must be delivered to the nucleus, whereas most RNA viruses remain in the cytoplasm. The uncoating process may be initiated by attachment to the receptor or promoted by the acidic environment or proteases found in an endosome or lysosome. Picornavirus capsids are weakened by the release of the VP4 capsid protein to allow uncoating. VP4 is released by insertion of the receptor into the keyhole-like canyon attachment site of the capsid. Enveloped viruses are uncoated on fusion with cell membranes. Fusion of the herpesvirus envelope with the plasma membrane releases its nucleocapsid, which then “docks” with the nuclear membrane to deliver its DNA genome directly to the site of replication. The release of the influenza nucleocapsid from its matrix and envelope is facilitated by the passage of protons from inside the endosome through the ion pore formed by the influenza M2 membrane protein to acidify the virion.
The reovirus and poxvirus are only partially uncoated on entry. The outer capsid of reovirus is removed, but the genome remains in an inner capsid, which contains the polymerases necessary for RNA synthesis. The initial uncoating of the poxviruses exposes a subviral particle to the cytoplasm, allowing synthesis of mRNA by virion-contained enzymes. An uncoating enzyme can then be synthesized to release the DNA-containing core into the cytoplasm.
Macromolecular Synthesis
Once inside the cell, the genome must direct the synthesis of viral mRNA and protein and generate identical copies of itself. The genome is useless unless it can be transcribed into functional mRNAs capable of binding to ribosomes and being translated into proteins. The means by which each virus accomplishes these steps depends on the structure of the genome (Figure 44-11) and the site of replication.
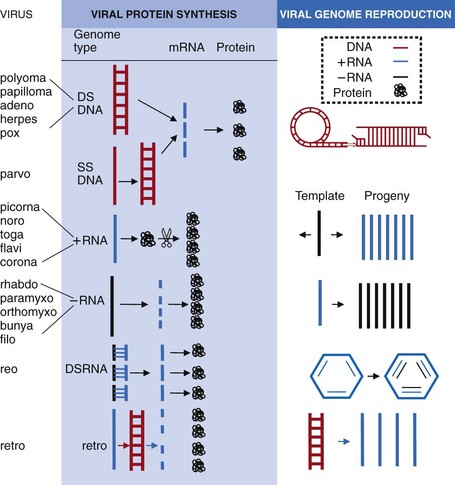
Figure 44-11 Viral macromolecular synthesis steps: the mechanism of viral mRNA and protein synthesis and genome replication are determined by the structure of the genome. 1, Double-stranded DNA (DS DNA) uses host machinery in the nucleus (except poxviruses) to make mRNA, which is translated by host cell ribosomes into proteins. Replication of viral DNA occurs by semiconservative means, by rolling circle, linear, and in other ways. 2, Single-stranded DNA (SS DNA) is converted into DS DNA and replicates like DS DNA. 3, (+) RNA resembles an mRNA that binds to ribosomes to make a polyprotein that is cleaved into individual proteins. One of the viral proteins is an RNA polymerase that makes a (−) RNA template and then more (+) RNA genome progeny and mRNAs. 4, (−) RNA is transcribed into mRNAs and a full-length (+) RNA template by the RNA polymerase carried in the virion. The (+) RNA template is used to make (−) RNA genome progeny. 5, DS RNA acts like (−) RNA. The (−) strands are transcribed into mRNAs by an RNA polymerase in the capsid. New (+) RNAs get encapsidated and (−) RNAs are made in the capsid. 6, Retroviruses have (+) RNA that is converted to complementary DNA (cDNA) by reverse transcriptase carried in the virion. cDNA integrates into the host chromosome, and the host makes mRNAs, proteins, and full-length RNA genome copies.
The cell’s machinery for transcription and mRNA processing is found in the nucleus. Most DNA viruses use the cell’s DNA-dependent RNA polymerase II and other enzymes to make mRNA. For example, eukaryotic mRNAs acquire a 3′ polyadenylated (polyA) tail and a 5′ methylated cap (for binding to the ribosome) and are processed to remove introns before being exported to the cytoplasm. Viruses that replicate in the cytoplasm must provide these functions or an alternative. Although poxviruses are DNA viruses, they replicate in the cytoplasm and therefore must encode enzymes for all these functions. Most RNA viruses replicate and produce mRNA in the cytoplasm, except for orthomyxoviruses and retroviruses. RNA viruses must encode the necessary enzymes for transcription and replication, because the cell has no means of replicating RNA. The mRNAs for RNA viruses may or may not acquire a 5′ cap or polyA tail.
The naked genome of DNA viruses (except poxviruses) and the positive-sense RNA viruses (except retroviruses) are sometimes referred to as infectious nucleic acids because they are sufficient for initiating replication on injection into a cell. These genomes can interact directly with host machinery to promote mRNA or protein synthesis.
In general, mRNA for nonstructural proteins is transcribed first (Figure 44-12). Early gene products (nonstructural proteins) are often DNA-binding proteins and enzymes, including virus-encoded polymerases. These proteins are catalytic, and only a few are required. Replication of the genome usually initiates the transition to transcription of late gene products. Late viral genes encode structural and other proteins. Many copies of these proteins are required to package the virus, but are generally not required before the genome is replicated. Newly replicated genomes also provide new templates for more late gene mRNA synthesis. Different DNA and RNA viruses control the time and amount of viral gene and protein synthesis in different ways.
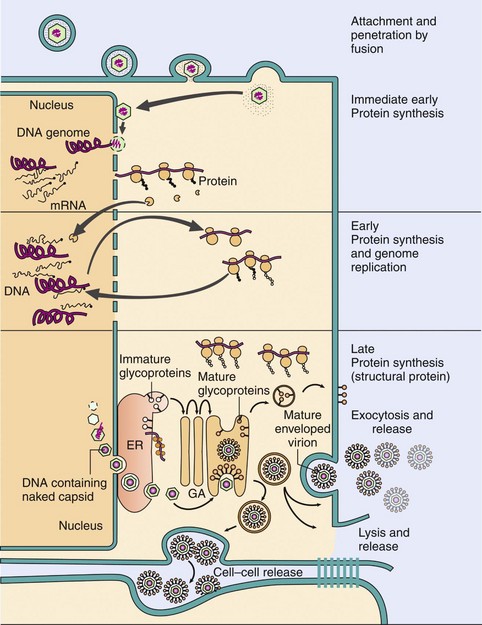
Figure 44-12 Replication of herpes simplex virus, a complex enveloped DNA virus. The virus binds to specific receptors and fuses with the plasma membrane. The nucleocapsid then delivers the DNA genome to the nucleus. Transcription and translation occur in three phases: immediate early, early, and late. Immediate early proteins promote the takeover of the cell; early proteins consist of enzymes, including the DNA-dependent DNA polymerase; and the late proteins are structural and other proteins, including the viral capsid and glycoproteins. The genome is replicated before transcription of the late genes. Capsid proteins migrate into the nucleus, assemble into icosadeltahedral capsids, and are filled with the DNA genome. The capsids filled with genomes bud through the nuclear and endoplasmic reticulum (ER) membranes into the cytoplasm, acquire tegument proteins, and then acquire their envelope as they bud through the viral glycoprotein-modified membranes of the trans-Golgi network. The virus is released by exocytosis or cell lysis. GA, Golgi apparatus.
DNA Viruses
Replication of the DNA genome requires a DNA-dependent DNA polymerase, other enzymes, and deoxyribonucleotide triphosphates, especially thymidine (Box 44-7). Transcription of the DNA virus genome (except for poxviruses) occurs in the nucleus, using host cell polymerases and other enzymes for viral mRNA synthesis. Transcription of the viral genes is regulated by the interaction of specific DNA-binding proteins with promoter and enhancer elements in the viral genome. The viral promoter and enhancer elements are similar in sequence to those of the host cell to allow binding of the cell’s transcriptional activation factors and DNA-dependent RNA polymerase. Cells from some tissues do not express the DNA-binding proteins necessary for activating the transcription of viral genes, and replication of the virus in that cell is thus prevented or limited.
Box 44-7
Properties of DNA Viruses
DNA is not transient or labile.
Many DNA viruses establish persistent infections (e.g., latent, immortalizing).
DNA genomes reside in the nucleus (except for poxviruses).
Viral DNA resembles host DNA for transcription and replication.
Viral genes must interact with host transcriptional machinery (except for poxviruses).
Viral gene transcription is temporally regulated.
Early genes encode DNA-binding proteins and enzymes.
Late genes encode structural and other proteins.
DNA polymerases require a primer to replicate the viral genome.
The larger DNA viruses encode means to promote efficient replication of their genome.
Parvovirus: requires cells undergoing DNA synthesis to replicate.
Papovavirus: stimulates cell growth and DNA synthesis.
Hepadnavirus: stimulates cell growth, cell makes RNA intermediate, encodes a reverse transcriptase.
Adenovirus: stimulates cellular DNA synthesis and encodes its own polymerase.
Herpesvirus: stimulates cell growth, encodes its own polymerase and enzymes to provide deoxyribonucleotides for DNA synthesis, establishes latent infection in host.
Poxvirus: encodes its own polymerases and enzymes to provide deoxyribonucleotides for DNA synthesis, replication machinery, and transcription machinery in the cytoplasm.
Different DNA viruses control the duration, timing, and quantity of viral gene and protein synthesis in different ways. The more complex viruses encode their own transcriptional activators, which enhance or regulate the expression of viral genes. For example, HSV encodes many proteins that regulate the kinetics of viral gene expression, including the VMW 65 (α-TIF protein, VP16). VMW 65 is carried in the virion, binds to the host cell transcription-activating complex (Oct-1), and enhances its ability to stimulate transcription of the immediate early genes of the virus.
Genes may be transcribed from either DNA strand of the genome and in opposite directions. For example, the early and late genes of the SV40 papovavirus are on opposite, nonoverlapping DNA strands. Viral genes may have introns requiring posttranscriptional processing of the mRNA by the cell’s nuclear machinery (splicing). The late genes of papovaviruses and adenoviruses are initially transcribed as a large RNA from a single promoter and then processed to produce several different mRNAs after removal of different intervening sequences (introns).
Replication of viral DNA follows the same biochemical rules as for cellular DNA. Replication is initiated at a unique DNA sequence of the genome called the origin (ori). This is a site recognized by cellular or viral nuclear factors and the DNA-dependent DNA polymerase. Viral DNA synthesis is semiconservative, and viral and cellular DNA polymerases require a primer to initiate synthesis of the DNA chain. The parvoviruses have DNA sequences that are inverted and repeated to allow the DNA to fold back and hybridize with itself to provide a primer. Replication of the adenovirus genome is primed by deoxycytidine monophosphate attached to a terminal protein. A cellular enzyme (primase) synthesizes an RNA primer to start the replication of the papovavirus genome, whereas the herpesviruses encode a primase.
Replication of the genome of the simple DNA viruses (e.g., parvoviruses, papovaviruses) uses the host DNA-dependent DNA polymerases, whereas the larger, more complex viruses (e.g., adenoviruses, herpesviruses, poxviruses) encode their own polymerases. Viral polymerases are usually faster but less precise than host cell polymerases, causing a higher mutation rate in viruses and providing a target for nucleotide analogues as antiviral drugs.
Hepadnavirus replication is unique in that a larger than genome positive-strand RNA copy is first synthesized by the cell’s DNA-dependent RNA polymerase and circularizes. Viral proteins surround the RNA, a viral encoded RNA-dependent DNA polymerase (reverse transcriptase) in this virion core makes a negative-strand DNA, and then the RNA is degraded. Positive-strand DNA synthesis is initiated but stops when the genome and core are enveloped, yielding a partially double-stranded, circular DNA genome.
Major limitations for replication of a DNA virus include availability of the DNA polymerase and deoxyribonucleotide substrates. Most cells in the resting phase of growth are not undergoing DNA synthesis because the necessary enzymes are not present, and deoxythymidine pools are limited. The smaller the DNA virus, the more dependent the virus is on the host cell to provide these functions (see Box 44-7). The parvoviruses are the smallest DNA viruses and replicate only in growing cells, such as erythroid precursor cells or fetal tissue. Speeding up the growth of the cell can enhance viral DNA and mRNA synthesis. The T antigen of SV40, the E6 and E7 of papillomavirus, and the E1a and E1b proteins of adenovirus bind to and prevent the function of growth-inhibitory proteins (p53 and the retinoblastoma gene product), resulting in cell growth, which also promotes virus replication. The larger DNA viruses may encode a DNA polymerase and other proteins to facilitate DNA synthesis and are more independent. HSV encodes a DNA polymerase and scavenging enzymes, such as deoxyribonuclease, ribonucleotide reductase, and thymidine kinase, to generate the necessary deoxyribonucleotide substrates for replication of its genome.
RNA Viruses
Replication and transcription of RNA viruses are similar processes, because the viral genomes are usually either an mRNA (positive-strand RNA) (Figure 44-13) or a template for mRNA (negative-strand RNA) (Box 44-8 and Figure 44-14). During replication and transcription, a double-stranded RNA replicative intermediate is formed. Double-stranded RNA is not normally found in uninfected cells and is a strong inducer of innate host protections.
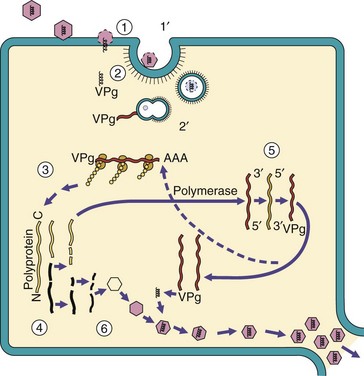
Figure 44-13 Replication of picornaviruses: a simple (+) RNA virus. 1, Interaction of the picornaviruses with receptors on the cell surface defines the target cell and weakens the capsid. 2, The genome is injected through the virion and across the cell membrane. 2′, Alternatively, the virion is endocytosed, and then the genome is released. 3, The genome is used as mRNA for protein synthesis. One large polyprotein is translated from the virion genome. 4, Then the polyprotein is proteolytically cleaved into individual proteins, including an RNA-dependent RNA polymerase. 5, The polymerase makes a (−) strand template from the genome and replicates the genome. A protein (VPg) is covalently attached to the 5′ end of the viral genome. 6, The structural proteins associate into the capsid structure, the genome is inserted, and the virions are released on cell lysis.
Box 44-8
Properties of RNA Viruses
Most RNA viruses replicate in the cytoplasm.
Cells cannot replicate RNA. RNA viruses must encode an RNA-dependent RNA polymerase.
The genome structure determines the mechanism of transcription and replication.
RNA viruses are prone to mutation.
The genome structure and polarity determine how viral messenger RNA (mRNA) is generated and proteins are processed.
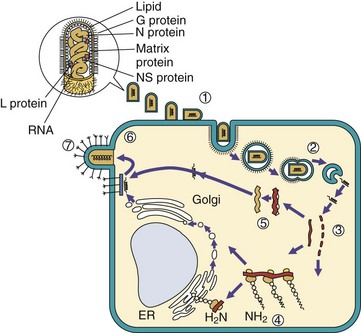
Figure 44-14 Replication of rhabdoviruses: a simple enveloped (−) RNA virus. 1, Rhabdoviruses bind to the cell surface and are (2) endocytosed. The envelope fuses with the endosome vesicle membrane to deliver the nucleocapsid to the cytoplasm. The virion must carry a polymerase, which (3) produces five individual messenger RNAs (mRNAs) and a full-length (+) RNA template. 4, Proteins are translated from the mRNAs, including one glycoprotein (G) which is co-translationally glycosylated in the endoplasmic reticulum (ER), processed in the Golgi apparatus, and delivered to the cell membrane. 5, The genome is replicated from the (+) RNA template, and N, L, and NS proteins associate with the genome to form the nucleocapsid. 6, The matrix protein associates with the G protein–modified membrane, which is followed by assembly of the nucleocapsid. 7, The virus buds from the cell in a bullet-shaped virion.
The RNA virus genome must code for RNA-dependent RNA polymerases (replicases and transcriptases) because the cell has no means of replicating RNA. The replicases and transcriptases are generated by addition of subunits or cleavage of a core polymerase. Because RNA is degraded relatively quickly, the RNA-dependent RNA polymerase must be provided or synthesized soon after uncoating to generate more viral RNA, or the infection will be aborted. Most viral RNA polymerases work at a fast pace but are also error prone, causing mutations. Replication of the genome provides new templates for production of more mRNA and genomes, which amplifies and accelerates virus replication.
The positive-strand RNA viral genomes of the picornaviruses, caliciviruses, coronaviruses, flaviviruses, and togaviruses act as mRNA, bind to ribosomes, and direct protein synthesis. The naked positive-strand RNA viral genome is sufficient to initiate infection by itself. After the virus-encoded, RNA-dependent RNA polymerase is produced, a negative-strand RNA template (antigenome) is synthesized. The template can then be used to generate more mRNA and to replicate the genome. For the togaviruses, coronaviruses, and caliciviruses, the negative-sense RNA is also used as a template to produce mRNAs for structural and other proteins (late genes). The mRNAs for picornaviruses are not capped at the 5′ end, but RNA for other viruses have 5′ caps and polyA tails. Transcription and replication of coronaviruses share many of these aspects but are more complex.
The negative-strand RNA virus genomes of the rhabdoviruses, orthomyxoviruses, paramyxoviruses, filoviruses, and bunyaviruses are the templates for production of mRNA. The negative-strand RNA genome is not infectious by itself, and a polymerase must be carried into the cell with the genome (associated with the genome as part of the nucleocapsid) to make individual mRNA for the different viral proteins. As a result, a full-length positive-strand RNA must also be produced by the viral polymerase to act as a template to generate more copies of the genome. The (−) RNA genome is like the negatives from a roll of movie film: each frame encodes a photo/mRNA, but a full-length positive is required for replicating the roll. Except for influenza viruses, transcription and replication of negative-strand RNA viruses occur in the cytoplasm. The influenza transcriptase requires a primer to produce mRNA. It uses the 5′ ends of cellular mRNA in the nucleus as primers for its polymerase and, in the process, steals the 5′ cap from the cellular mRNA. The influenza genome is also replicated in the nucleus.
The reoviruses have a segmented, double-stranded RNA genome and undergo a more complex means of replication and transcription. The reovirus RNA polymerase is part of the inner capsid core; mRNA units are transcribed from each of the 10 or more segments of the genome while they are still in the core. The negative strands of the genome segments are used as templates for mRNA in a manner similar to that of the negative-strand RNA viruses. Reovirus-encoded enzymes contained in the inner capsid core add the 5′ cap to viral mRNA. The mRNA does not have polyA. The mRNAs are released into the cytoplasm, where they direct protein synthesis or are sequestered into new cores. The positive-strand RNA in the new cores acts as a template for negative-strand RNA, and the core polymerase produces the progeny double-stranded RNA.
The arenaviruses have an ambisense genome with (−) sequences adjacent to (+) sequences. The early mRNAs of the virus are transcribed from the negative-sense portion of the genome, a full-length replicative intermediate is produced to generate a new genome, and the late mRNAs of the virus are transcribed from the region complementary to the (+) sequences in the replicative intermediate.
Although the retroviruses have a positive-strand RNA genome, the virus provides no means for replication of the RNA in the cytoplasm. Instead, the retroviruses carry two copies of the genome, two transfer RNA (tRNA) molecules, and an RNA-dependent DNA polymerase (reverse transcriptase) in the virion. The tRNA is used as a primer for synthesis of a circular complementary DNA (cDNA) copy of the genome. The cDNA is synthesized in the cytoplasm, travels to the nucleus, and is then integrated into the host chromatin. The viral genome becomes a cellular gene. Promoters at the end of the integrated viral genome enhance the transcription of the viral DNA sequences by the cell. Full-length RNA transcripts are used as new genomes, and individual mRNAs are generated by differential splicing of this RNA.
The most unusual mode of replication is reserved for the deltavirus. The deltavirus resembles a viroid. The genome is a circular, rod-shaped, single-stranded RNA, which is extensively hybridized to itself. As the exception, the deltavirus RNA genome is replicated by the host cell DNA-dependent RNA polymerase II in the nucleus. A portion of the genome forms an RNA structure called a ribozyme, which cleaves the RNA circle to produce an mRNA.
Viral Protein Synthesis
All viruses depend on the host cell ribosomes, tRNA, and mechanisms for posttranslational modification to produce their proteins. The binding of mRNA to the ribosome is mediated by a 5′ cap structure of methylated guanosine or a special RNA loop structure (internal ribosome entry sequence [IRES]), which binds within the ribosome to initiate protein synthesis. The cap structure, if used, is acquired in different ways by different viruses. The IRES structure was discovered first in the picornavirus genome and then in selected cellular mRNAs. Most but not all viral mRNA have a polyadenosine (polyA) tail, like eukaryotic mRNAs.
Unlike bacterial ribosomes, which can bind to a polycistronic mRNA and translate several gene sequences into separate proteins, the eukaryotic ribosome binds to mRNA and can make only one continuous protein, and then it falls off the mRNA. Each virus deals with this limitation differently, depending on the structure of the genome. For example, the entire genome of a positive-strand RNA virus is read by the ribosome and translated into one giant polyprotein. The polyprotein is subsequently cleaved by cellular and viral proteases into functional proteins. DNA viruses, retroviruses, and most negative-strand RNA viruses transcribe separate mRNA for smaller polyproteins or individual proteins. The orthomyxovirus and reovirus genomes are segmented, and most of the segments code for single proteins for this reason.
Viruses use different tactics to promote preferential translation of their viral mRNA instead of cellular mRNA. In many cases, the concentration of viral mRNA in the cell is so large that it occupies most of the ribosomes, preventing translation of cellular mRNA. Adenovirus infection blocks the egress of cellular mRNA from the nucleus. HSV and other viruses inhibit cellular macromolecular synthesis and induce degradation of the cell’s DNA and mRNA. To promote selective translation of its mRNA, poliovirus uses a virus-encoded protease to inactivate the 200,000-Da cap-binding protein of the ribosome to prevent binding and translation of the cell’s 5′capped cellular mRNA. Togaviruses and many other viruses increase the permeability of the cell’s membrane; thus the ribosomal affinity for most cellular mRNA is decreased. All these actions also contribute to the cytopathology of the virus infection. The pathogenic consequences of these actions are discussed further in Chapter 45.
Some viral proteins require posttranslational modifications, such as phosphorylation, glycosylation, acylation, or sulfation. Protein phosphorylation is accomplished by cellular or viral protein kinases and is a means of modulating, activating, or inactivating proteins. Several herpesviruses and other viruses encode their own protein kinase. Viral glycoproteins are synthesized on membrane-bound ribosomes and have the amino acid sequences to allow insertion into the rough endoplasmic reticulum and N-linked glycosylation. The high-mannose precursor form of the glycoproteins progresses from the endoplasmic reticulum through the vesicular transport system of the cell and is processed through the Golgi apparatus. The sialic acid–containing mature glycoprotein is expressed on the plasma membrane of the cell unless the glycoprotein expresses protein sequences for retention in an intracellular organelle. The presence of the glycoproteins determines where the virion will assemble within the cell. Other modifications, such as O-glycosylation, acylation, and sulfation of the proteins, can also occur during progression through the Golgi apparatus.
Assembly
Virion assembly is analogous to a three-dimensional interlocking puzzle that puts itself together in the box. The virion is built from small, easily manufactured parts that enclose the genome in a functional package. Each part of the virion has recognition structures that allow the virus to form the appropriate protein-protein, protein-nucleic acid, and (for enveloped viruses) protein-membrane interactions needed to assemble into the final structure. The assembly process begins when the necessary pieces are synthesized, and the concentration of structural proteins in the cell is sufficient to drive the process thermodynamically, much like a crystallization reaction. The assembly process may be facilitated by scaffolding proteins or other proteins, some of which are activated or release energy on proteolysis. For example, cleavage of the VP0 protein of poliovirus releases the VP4 peptide, which solidifies the capsid.
The site and mechanism of virion assembly in the cell depend on where genome replication occurs and whether the final structure is a naked capsid or an enveloped virus. Assembly of the DNA viruses, other than poxviruses, occurs in the nucleus and requires transport of the virion proteins into the nucleus. RNA virus and poxvirus assembly occurs in the cytoplasm.
Capsid viruses may be assembled as empty structures (procapsids) to be filled with the genome (e.g., picornaviruses), or they may be assembled around the genome. Nucleocapsids of the retroviruses, togaviruses, and the negative-strand RNA viruses assemble around the genome and are subsequently enclosed in an envelope. The helical nucleocapsid of negative-strand RNA viruses includes the RNA-dependent RNA polymerase necessary for mRNA synthesis in the target cell.
For enveloped viruses, newly synthesized and processed viral glycoproteins are delivered to cellular membranes by vesicular transport. Acquisition of an envelope occurs after association of the nucleocapsid with the viral glycoprotein-containing regions of host cell membranes in a process called budding. Matrix proteins for negative-strand RNA viruses line and promote the adhesion of nucleocapsids with the glycoprotein-modified membrane. As more interactions occur, the membrane surrounds the nucleocapsid, and the virus buds from the membrane.
The type of genome and the protein sequence of the glycoproteins determine the site of budding. Most RNA viruses bud from the plasma membrane, and the virus is released from the cell at the same time without killing the cell. The flaviviruses, coronaviruses, and bunyaviruses acquire their envelope by budding into the endoplasmic reticulum and Golgi membranes and may remain cell-associated in these organelles. The HSV nucleocapsid assembles in the nucleus and buds into and then out of the endoplasmic reticulum. The nucleocapsid is dumped into the cytoplasm, viral proteins associate with the capsid, and then the envelope is acquired by budding into a trans-Golgi network membrane decorated with the 10 viral glycoproteins. The virion is transported to the cell surface and released by exocytosis, on cell lysis, or transmitted through cell-to-cell bridges.
Viruses use different tricks to ensure that all the parts of the virus are assembled into complete virions. The RNA polymerase required for infection by negative-strand RNA viruses is carried on the genome as part of a helical nucleocapsid. The human immunodeficiency virus (HIV) and other retrovirus genomes are packaged in a procapsid consisting of a polyprotein containing the protease, polymerase, integrase, and structural proteins. This procapsid binds to viral glycoprotein-modified membranes, and the virion buds from the membrane. The virus-encoded protease is activated within the virion and cleaves the polyprotein to produce the final infectious nucleocapsid and the required proteins within the envelope.
Assembly of viruses with segmented genomes, such as influenza or reovirus, requires accumulation of at least one copy of each gene segment. This can be accomplished if the segments assemble together like capsid subunits or randomly package more segments per virion than necessary. This will statistically generate a small but acceptable percentage of functional viruses.
Errors are made by the viral polymerase and during viral assembly. Empty virions and virions containing defective genomes are produced. As a result, the particle to infectious virus ratio, also called particle to plaque-forming unit ratio, is high, usually greater than 10, and during rapid viral replication can even be 104. Defective viruses can occupy the machinery (e.g., bind to the receptor) required for normal virus replication to prevent (interfere with) virus production (defective interfering particles).
Release
Viruses can be released from cells after lysis of the cell, by exocytosis, or by budding from the plasma membrane. Naked capsid viruses are generally released after lysis of the cell. Release of most enveloped viruses occurs after budding from the plasma membrane without killing the cell. Survival of the cell allows continual release of virus from the factory. Lysis and plasma membrane budding are efficient means of release. Viruses that bud or acquire their membrane in the cytoplasm (e.g., flaviviruses, poxviruses) remain cell associated and are released by exocytosis or cell lysis. Viruses that bind to sialic acid receptors (e.g., orthomyxoviruses, certain paramyxoviruses) may also have an NA. The NA removes potential sialic acid receptors on the glycoproteins of the virion and the host cell to prevent clumping and facilitate release.
Reinitiation of the Replication
Spread of the infection occurs from virus released to the extracellular medium, but alternatively, the virus, nucleocapsid, or genome can be transmitted through cell-to-cell bridges, upon cell-to-cell fusion, or vertically to daughter cells. These alternate routes allow the virus to escape antibody detection. Some herpesviruses, retroviruses, and paramyxoviruses can induce cell-to-cell fusion to merge the cells into multinucleated giant cells (syncytia), which become huge virus factories. The retroviruses and some DNA viruses can transmit their integrated copy of the genome vertically to daughter cells on cell division.
Viral Genetics
Mutations spontaneously and readily occur in viral genomes, creating new virus strains with properties differing from the parental, or wild-type, virus. These variants can be identified by their nucleotide sequences, antigenic differences (serotypes), or differences in functional or structural properties. Most mutations have no effect or are detrimental to the virus. Mutations in essential genes inactivate the virus, but mutations in other genes can produce antiviral drug resistance or alter the antigenicity or pathogenicity of the virus.
Errors in copying the viral genome during virus replication produce many mutations. This is because of the poor fidelity of the viral polymerase and the rapid rate of genome replication. In addition, RNA viruses do not have a genetic error-checking mechanism. As a result, the rates of mutation for RNA viruses are usually greater than for DNA viruses.
Mutations that inactivate essential genes are termed lethal mutations. These mutants are difficult to isolate, because the virus cannot replicate. A deletion mutant results from the loss or selective removal of a portion of the genome and the function that it encodes. Other mutations may produce plaque mutants, which differ from the wild type in the size or appearance of the infected cells; host range mutants, which differ in the tissue type or species of target cell that can be infected; or attenuated mutants, which are variants that cause less serious disease in animals or humans. Conditional mutants, such as temperature-sensitive (ts) or cold-sensitive mutants, have a mutation in a gene for an essential protein that allows virus production only at certain temperatures. Whereas ts mutants generally grow well or relatively better at 30° C to 35° C, the encoded protein is inactive at elevated temperatures of 38° C to 40° C, preventing virus production. Live virus vaccines are often conditional or host range mutants and attenuated for human disease.
New virus strains can also arise by genetic interactions between viruses or between the virus and the cell (Figure 44-15). Intramolecular genetic exchange between viruses or the virus and the host is termed recombination. Recombination can occur readily between two related DNA viruses. For example, co-infection of a cell with the two closely related herpesviruses (HSV types 1 and 2) yields intertypic recombinant strains. These new hybrid strains have genes from types 1 and 2. Integration of retroviruses into host cell chromatin is a form of recombination. Recombination of two related RNA viruses, Sindbis and eastern equine encephalitis virus, resulted in creation of another togavirus, western equine encephalitis (WEE) virus.
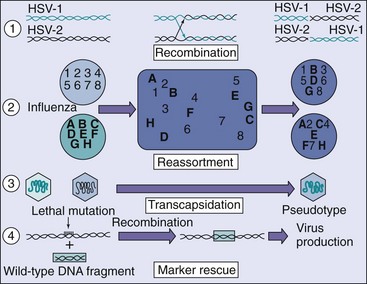
Figure 44-15 Genetic exchange between viral particles can give rise to new viral types, as illustrated. Representative viruses include the following: 1, intertypic recombination of herpes simplex virus type 1 (HSV-1) and type 2 (HSV-2); 2, reassortment of two strains of influenza virus; 3, rescue of a papovavirus defective in assembly by a complementary defective virus (transcapsidation); and 4, marker rescue of a lethal or conditional mutation.
Viruses with segmented genomes (e.g., influenza viruses and reoviruses) form hybrid strains on infection of one cell with more than one virus strain. This process, termed reassortment, is analogous to picking 10 marbles out of a box containing 10 black and 10 white marbles. Very different strains of influenza A virus are created on co-infection with a virus from different species (see Figure 57-5).
In some cases, a defective viral strain can be rescued by the replication of another mutant, by the wild-type virus, or by a cell line bearing a replacement viral gene. Replication of the other virus or expression of the gene in the cell provides the missing function required by the mutant (complementation), allowing replication to occur. An experimental disabled infectious single-cycle HSV (DISC-HSV) vaccine lacks an essential gene and is grown in a cell line that expresses that gene product to “complement” the virus. The vaccine virus can infect the normal cells of the individual, but the virions that are produced lack the function necessary for replication in other cells and cannot spread. Rescue of a lethal or conditional-lethal mutant with a defined genetic sequence, such as a restriction endonuclease DNA fragment, is called marker rescue. Marker rescue is used to map the genomes of viruses such as HSV. Virus produced from cells infected with different virus strains may be phenotypically mixed and have the proteins of one strain but the genome of the other (transcapsidation). Pseudotypes are generated when transcapsidation occurs between different types of virus, but this is rare.
Individual virus strains or mutants are selected by their ability to use the host cell machinery and to withstand the conditions of the body and the environment. Cellular properties that can act as selection pressures include the growth rate of the cell and tissue-specific expression of certain proteins required by the virus (e.g., enzymes, glycoproteins, transcription factors) and proteins that prevent essential virus functions. The conditions of the body, its elevated temperature, innate and immune defenses, and tissue structure are also selection pressures for viruses. The viruses that cannot endure these conditions or evade the host defenses are eliminated. A small selective advantage in a mutant virus can shortly lead to its becoming the predominant viral strain. The high mutation rate of HIV promotes a switch in target cell tropism to include different types of T cells, the development of antiviral drug-resistant strains, and the generation of antigenic variants during a patient’s course of infection.
The growth of virus under benign laboratory conditions allows weaker strains to survive because of the absence of the selective pressures of the human body. This process is used to select attenuated virus strains for use in vaccines.
Viral Vectors for Therapy
Genetically manipulated viruses can be excellent delivery systems for foreign genes. Viruses can provide gene replacement therapy, can be used as vaccines to promote immunity to other agents or tumors, and can act as targeted killers of tumors. The advantages of using viruses are that they can be readily amplified by replication in appropriate cells, and they target specific tissues and deliver the DNA or RNA into the cell. Viruses that are being developed as vectors include retroviruses, adenoviruses, HSV, adeno-associated virus (parvovirus), poxviruses (e.g., vaccinia and canarypox) (see Figure 52-3), and even some togaviruses. The viral vectors are usually defective or attenuated viruses, in which the foreign DNA replaces a virulence or unessential gene. The foreign gene may be under the control of a viral promoter or even a tissue-specific promoter. Defective virus vectors are grown in cell lines that express the missing viral functions “complementing” the virus. The progeny can deliver their nucleic acid but not produce infectious virus. Retroviruses and adeno-associated viruses can integrate into cells and permanently deliver a gene into the cell’s chromosome. Adenovirus and HSV promote targeted delivery of the foreign gene to receptor-bearing cells. Genetically attenuated HSVs are being developed to specifically kill the growing cells of glioblastomas while sparing the surrounding neurons. Adenovirus and canarypox virus are being used to carry and express HIV genes as vaccines. Vaccinia virus carrying a gene for the rabies glycoprotein is already being used successfully to immunize raccoons, foxes, and skunks in the wild. Some day, virus vectors may be routinely used to treat cystic fibrosis, Duchenne muscular dystrophy, lysosomal storage diseases, and immunologic disorders.
1. Describe the features of these viruses that are similar, and those that are different.
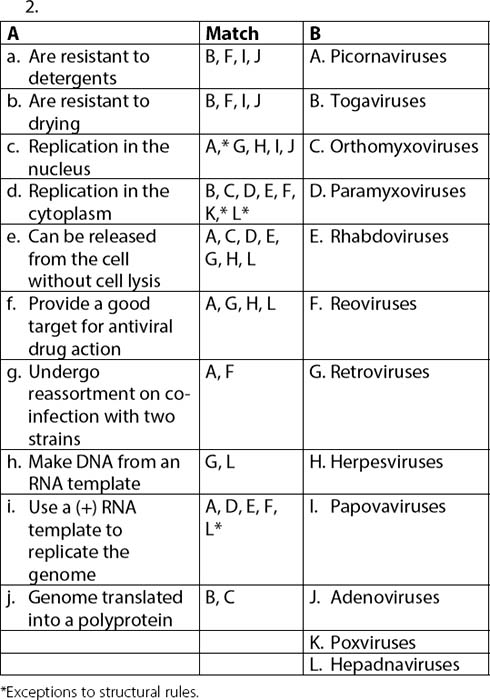
2. Match the characteristics from column A with the appropriate viral families in column B, based on your knowledge of their physical and genome structure and their implications.
| A | B | |
| a. | Are resistant to detergents | Picornaviruses |
| b. | Are resistant to drying | Togaviruses |
| c. | Replication in the nucleus | Orthomyxoviruses |
| d. | Replication in the cytoplasm | Paramyxoviruses |
| e. | Can be released from the cell without cell lysis | Rhabdoviruses |
| f. | Provide a good target for antiviral drug action | Reoviruses |
| g. | Undergo reassortment on co-infection with two strains | Retroviruses |
| h. | Make DNA from an RNA template | Herpesviruses |
| i. | Use a (+) RNA template to replicate the genome | Papovaviruses |
| j. | Genome translated into a polyprotein | Adenoviruses, poxviruses hepadnaviruses |
3. Based on structural considerations, which of the virus families listed in question 2 should be able to endure fecal-oral transmission?
4. List the essential enzymes encoded by the virus families listed in question 2.
5. A mutant defective in the HSV type 1 DNA polymerase gene replicates in the presence of HSV type 2. The progeny virus contains the HSV type 1 genome but is recognized by antibodies to HSV type 2. Which genetic mechanisms may be occurring?
6. How are the early and late genes of the togaviruses, papovaviruses, and herpesviruses distinguished, and how is the time of their expression regulated?
7. What are the consequences (no effect, decreased efficiency, or inhibition of replication) of a deletion mutation in the following viral enzymes?
1. a. Both are picornaviruses and have similar structures and modes of replication, but unlike poliovirus, rhinoviruses are acid and temperature labile.
b. Polio and rotaviruses are capsid viruses that are spread by the fecal-oral route. Polio has a +RNA genome; rotaviruses have a double-stranded RNA genome.
c. Polio and WEE virus have positive-stranded RNA, and their genomes are infectious. WEE is a togavirus, which can generate early and late proteins from a full-length or partial-length RNA. WEE is enveloped and spread in mosquito saliva and blood.
d. Yellow fever virus and dengue virus are flaviviruses that are enveloped, +RNA viruses, both of which are spread by mosquitoes in blood and mosquito saliva.
e. EBV and CMV are herpesviruses, which have large DNA genomes enclosed in an icosahedral capsid surrounded by an envelope. These viruses have complex replication schemes that are controlled at the transcription level by some cells. Both viruses are strictly human viruses, but EBV infects B lymphocytes while CMV has a broader tissue tropism.
3. Adenovirus, picornavirus, reovirus, and papovavirus.
4. DNA-dependent DNA polymerase: adenovirus, herpes, poxviruses; RNA-dependent DNA polymerase: hepadnavirus, retroviruses; RNA-dependent RNA polymerase: all the RNA viruses, except retrovirus and hepadnavirus; also, poxvirus. Integrase, protease: retrovirus.
5. Complementation: An HSV-2 gene may provide the missing activity for the mutant. Transcapsidation: The HSV genome may be encapsidated and enveloped in an HSV-2 virion particle. Recombination: HSV-1 and HSV-2 share enough similarity to allow recombination of the two genomes and the generation of a hybrid virus.
6. The early genes of togaviruses are expressed from the infecting +RNA genome (42S). Later, a subgenomic mRNA (26S) is transcribed from the replicative intermediate that encodes the late, structural proteins. The polyomavirus genome is circular, and the early genes are transcribed in one direction, and the late genes are transcribed in the opposite direction. The herpesvirus immediate early genes are activated by host DNA-binding proteins. The early genes are recognized by viral proteins, and different combinations of viral proteins activate the late proteins after genome replication is initiated.
7. a. EBV polymerase: no virus production
b. HSV thymidine kinase: inefficient virus production, especially in neurons
c. HIV reverse transcriptase: no virus production
d. Influenza B virus NA: very inefficient virus production
e. Rabies virus (rhabdovirus) G protein: no virus production