Chapter 2 Bacterial structure and taxonomy
Classification of all living beings, including microbes has been attempted by many over centuries (Table 2.1). Traditionally, though they were all classified into two kingdoms, plants and animals, classification was arbitrary and based on morphological and growth characteristics. With the development of novel techniques, the latter classification was expanded to include five kingdoms: monera, protista, plantae, fungi and animalia. However, the current understanding based on their genetic relatedness is that all forms of life fall into three domains: Archaea, Bacteria and Eucarya. The main differences among Archaea, Bacteria and Eucarya are listed in Table 2.2. Note that taken together, Archaea and Bacteria are also known as prokaryotes (see below).
Table 2.2 Major differences among the three domains of life
| Bacteria | Archaea | Eucarya |
|---|---|---|
| Organization of the genetic material and replication | ||
| DNA free in the cytoplasm | DNA free in the cytoplasm | DNA is contained with a membrane-bound nucleus. A nucleolus is also present |
| Only one chromosome | Only one chromosome | More than one chromosome. Two copies of each chromosome may be present (diploid) |
| DNA associated with histone-like proteins | DNA associated with histone-like proteins | DNA complexed with histone proteins |
| May contain extrachromosomal elements called plasmids | Plasmids may be found | Plasmids only found in yeast |
| Introns not found in mRNA | Introns not found in most genes | Introns found in all genes |
| Cell division by binary fission – asexual replication only | Reproduce asexually and spores are not found | Cells divide by mitosis |
| Transfer of genetic information occurs by conjugation, transduction and transformation (see Chapter 3) | Processes similar to bacterial conjugation enables exchange of genetic material | Exchange of genetic information occurs during sexual reproduction. Meiosis leads to the production of haploid cells (gametes), which can fuse |
| Cellular organization | ||
| Cytoplasmic membrane contains hopanoids | Membranes contain isoprenes | Cytoplasmic membrane contains sterols |
| Lipopolysaccharides and teichoic acids found | No lipopolysaccharides or teichoic acids found | |
| Energy metabolism associated with the cytoplasmic membrane | Mitochondria present in most cases | |
| Photosynthesis associated with membrane systems and vesicles in cytoplasm | Chloroplasts present in algal and plant cells | |
| Internal membranes, endoplasmic reticulum and Golgi apparatus present associated with protein synthesis and targeting | ||
| Membrane vesicles such as lysosomes and peroxisomes present | ||
| Cytoskeleton of microtubules present | ||
| Flagella consist of one protein, flagellin | Contains flagella that derive energy from proton pumps | Flagella have a complex structure with 9 + 2 microtubular arrangement |
| Ribosomes – 70S | Ribosomes behave more like eucarya when exposed to inhibitors | Ribosomes – 80S (mitochondrial and chloroplast ribosomes are 70S) |
| Peptidoglycan cell walls | Cell walls lack peptidoglycan | Polysaccharide cell walls, where present, are generally either cellulose or chitin |
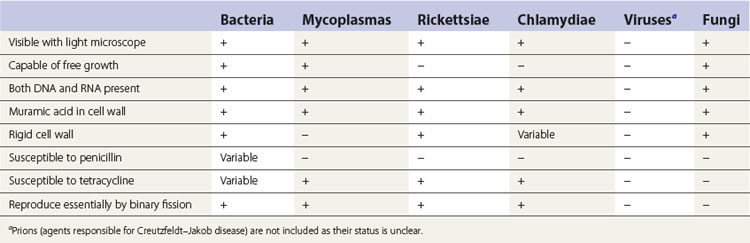
Viruses are not included in this classification as they are unique, acellular, metabolically inert organisms and therefore replicate only within living cells. Other differences between viruses and cellular organisms include:
Eukaryotes and prokaryotes
As mentioned above, another modification of classifying cellular organisms is to divide them into prokaryotes (i.e. Archaea and Bacteria) and eukaryotes (Greek karyon: nucleus). Fungi, protozoa and humans, for instance, are eukaryotic, whereas bacteria are prokaryotic. In prokaryotes, the bacterial genome, or chromosome, is a single, circular molecule of double-stranded DNA, lacking a nuclear membrane (smaller, single or multiple circular DNA molecules called plasmids may also be present in bacteria), whereas the eukaryotic cell has a true nucleus with multiple chromosomes surrounded by a nuclear membrane.
Bacteria comprise the vast majority of human pathogens, while archaea appear rarely to cause human disease and live in extreme environments (e.g. high temperature or salt concentrations). Archaea received little attention traditionally as they cannot be easily cultured in the laboratory. Interestingly, recent studies using novel techniques such as pyrosequencing have uncovered their presence in the oral cavity. Some studies have even shown that certain species of archaea are more frequently found in subgingival plaque in periodontal disease.
Morphology
Shape and size
The shape of a bacterium is determined by its rigid cell wall. Bacteria are classified by shape into three basic groups (Fig. 2.1A and B):
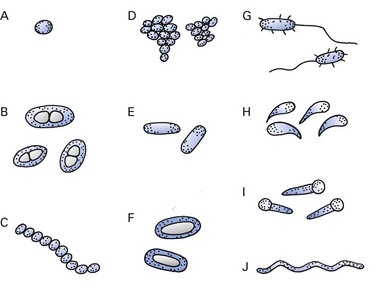
Fig. 2.1 Common bacterial forms. (A) Coccus; (B) capsulated diplococci; (C, D) cocci in chains (e.g. streptococcus) and clusters (e.g. staphylococcus); (E) bacillus; (F, G) capsulated and flagellated bacillus (e.g. Escherichia coli); (H) curved bacilli (e.g. Vibrio spp.); (I) spore-bearing bacilli (e.g. Clostridium tetani); (J) spirochaete.
Some bacteria with variable shapes, appearing both as coccal and bacillary forms, are called pleomorphic (pleo: many; morphic: shaped) in appearance.
The size of bacteria ranges from about 0.2 to 5 µm. The smallest bacteria approximate the size of the largest viruses (poxviruses), whereas the longest bacilli attain the same length as some yeasts and human red blood cells (7 µm).
Arrangement
Bacteria, whichever shape they may be, arrange themselves (usually according to the plane of successive cell division) as pairs (diplococci), chains (streptococci), grape-like clusters (staphylococci) or as angled pairs or palisades (corynebacteria).
Gram-staining characteristics
In clinical microbiology, bacteria can be classified into two major subgroups according to the staining characteristics of their cell walls. The stain used, called the Gram stain (first developed by a Danish physician, Christian Gram), divides the bacteria into Gram-positive (purple) and Gram-negative (pink) groups. The Gram-staining property of bacteria is useful both for their identification and in the therapy of bacterial infections because, in general, Gram-positive bacteria are more susceptible to penicillins than Gram-negative bacteria.
Structure
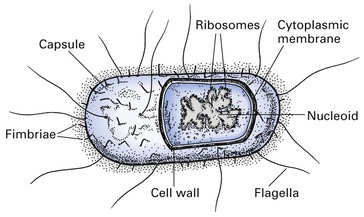
The structure of a typical bacterium is shown in Figure 2.2. Bacteria have a rigid cell wall protecting a fluid protoplast comprising a cytoplasmic membrane and a variety of other components (described below).
Structures external to the cell wall
Flagella
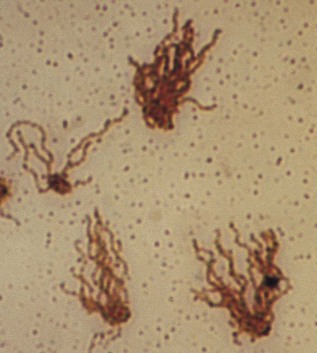
Flagella are whip-like filaments that act as propellers and guide the bacteria towards nutritional and other sources (Fig. 2.3). The filaments are composed of many subunits of a single protein, flagellin. Flagella may be located at one end (monotrichous, a single flagellum; lophotrichous, many flagella) or all over the outer surface (peritrichous). Many bacilli (rods) have flagella, but most cocci do not and are therefore non-motile. Spirochaetes move by using a flagellum-like structure called the axial filament, which wraps around the cell to produce an undulating motion.
Fimbriae and pili
Fimbriae and pili are fine, hair-like filaments, shorter than flagella, that extend from the cell surface. Pili, found mainly on Gram-negative organisms, are composed of subunits of a protein, pilin, and mediate the adhesion of bacteria to receptors on the human cell surface – a necessary first step in the initiation of infection. A specialized type of pilus, the sex pilus, forms the attachment between the male (donor) and the female (recipient) bacteria during conjugation, when genes are transferred from one bacterium to another.
Glycocalyx (slime layer)
The glycocalyx is a polysaccharide coating that covers the outer surfaces of many bacteria and allows the bacteria to adhere firmly to various structures, e.g. oral mucosa, teeth, heart valves and catheters, and contribute to the formation of biofilms. This is especially true in the case of Streptococcus mutans, a major cariogenic organism, which has the ability to produce vast quantities of extracellular polysaccharide in the presence of dietary sugars such as sucrose.
Capsule
An amorphous, gelatinous layer (usually more substantial than the glycocalyx) surrounds the entire bacterium; it is composed of polysaccharide, and sometimes protein (e.g. anthrax bacillus). The sugar components of the polysaccharide vary in different bacterial species and frequently determine the serological type within a species (e.g. 84 different serological types of Streptococcus pneumoniae can be distinguished by the antigenic differences of the sugars in the polysaccharide capsule). The capsule is important because:
Cell wall
The cell wall confers rigidity upon the bacterial cell. It is a multilayered structure outside the cytoplasmic membrane. It is porous and permeable to substances of low molecular weight.
The inner layer of the cell wall is made of peptidoglycan and is covered by an outer membrane that varies in thickness and chemical composition, depending upon the Gram-staining property of the bacteria (Fig. 2.4). The term ‘peptidoglycan’ is derived from the peptides and the sugars (glycan) that make up the molecule. (Synonyms for peptidoglycan are murein and mucopeptide.)
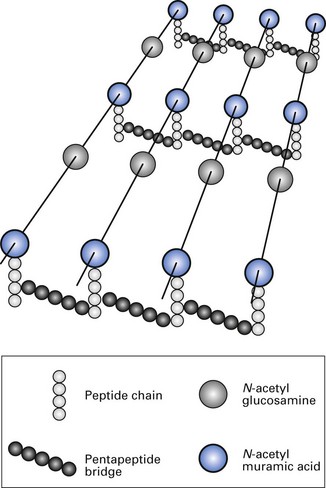
Fig. 2.4 Chemical structure of cross-linking peptidoglycan component of cell wall, common to both Gram-positive and Gram-negative bacteria.
(After Sharon, N (1969). The bacterial cell wall. Scientific American 220, 92.)
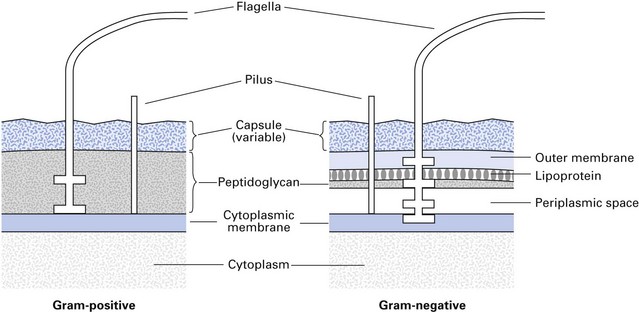
The cell walls of Gram-positive and Gram-negative bacteria have important structural and chemical differences (Fig. 2.5):
Bacteria with defective cell walls
Some bacteria can survive with defective cell walls. These include mycoplasmas, L-forms, spheroplasts and protoplasts.
Mycoplasmas do not possess a cell wall and do not need hypertonic media for their survival. They occur in nature and may cause human disease (e.g. pneumonia).
L-forms are usually produced in the laboratory and may totally or partially lack cell walls. They may be produced in patients treated with penicillin and, like mycoplasmas, can replicate on ordinary media.
Both spheroplasts (derived from Gram-negative bacteria) and protoplasts (derived from Gram-positive bacteria) lack cell walls, cannot replicate on laboratory media and are unstable and osmotically fragile. They require hypertonic conditions for maintenance and are produced in the laboratory by the action of enzymes or antibiotics.
Cytoplasmic membrane
The cytoplasmic membrane lies just inside the peptidoglycan layer of the cell wall and is a ‘unit membrane’ composed of a phospholipid bilayer similar in appearance to that of eukaryotic cells. However, eukaryotic membranes contain sterols, whereas prokaryotes generally do not (the only exception being mycoplasmas). The membrane has the following major functions:
Cytoplasm
The cytoplasm comprises an inner, nucleoid region (composed of DNA), which is surrounded by an amorphous matrix that contains ribosomes, nutrient granules, metabolites and various ions.
Nuclear material or nucleoid
Bacterial DNA comprises a single, supercoiled, circular chromosome that contains about 2000 genes, approximately 1 mm long in the unfolded state. (It is analogous to a single, haploid chromosome.) During cell division, it undergoes semiconservative replication bidirectionally from a fixed point.
Ribosomes
Ribosomes are the sites of protein synthesis. Bacterial ribosomes differ from those of eukaryotic cells in both size and chemical composition. They are organized in units of 70S, compared with eukaryotic ribosomes of 80S. These differences are the basis of the selective action of some antibiotics that inhibit bacterial, but not human, protein β-synthesis.
Bacterial spores
Spores are formed in response to adverse conditions by the medically important bacteria that belong to the genus Bacillus (which includes the agent of anthrax) and the genus Clostridium (which includes the agents of tetanus and botulism). These bacteria sporulate (form spores) when nutrients, such as sources of carbon and nitrogen, are scarce (Fig. 2.6). The spore develops at the expense of the vegetative cell and contains bacterial DNA, a small amount of cytoplasm, cell membrane, peptidoglycan, very little water and, most importantly, a thick, keratin-like coat. This coat, which contains a high concentration of calcium dipicolinate, is remarkably resistant to heat, dehydration, radiation and chemicals. Once formed, the spore is metabolically inert and can remain dormant for many years. Spores are called either terminal or subterminal, depending on their position in relation to the cell wall of the bacillus from which they developed.
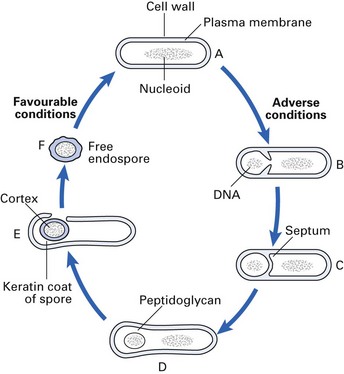
Fig. 2.6 The cycle of sporulation. (A) Vegetative cell; (B) ingrowth of cytoplasmic membrane; (C) developing forespore; (D) forespore completely cut off from the cell cytoplasm; (E) development of cortex and keratin spore coat; (F) liberation of spore and conversion to vegetative state under favourable conditions.
When appropriate conditions supervene (i.e. water, nutrients), there is enzymatic degradation of the coat, and the spore transforms itself into a metabolizing, reproducing bacterial cell once again (Fig. 2.6).
Clinical relevance of bacterial spores
The clinical importance of spores lies in their extraordinary resistance to heat and chemicals. Because of this, sterilization cannot be easily achieved by boiling; other, more efficacious methods of sterilization, such as steam under pressure (autoclaving), are required to ensure the sterility of products used for surgical purposes (Chapter 37). This property of bacterial spores is exploited when they are used for evaluating the sterilization efficacy of autoclaves; spores of Bacillus stearothermophilus and other species are used for this purpose.
Taxonomy
The systematic classification and categorization of organisms into ordered groups are called taxonomy. A working knowledge of taxonomy is useful for diagnostic microbiology and for studies in epidemiology and pathogenicity.
As mentioned at the beginning of this chapter, organisms encountered in medical microbiology fall into the domains of Bacteria, Archaea and Eucarya. Although this system of classification is based on the evolutionary relatedness or the genetic homogeneity of the species represented in each domain, a more pragmatic means of classification is employed in the clinical microbiology laboratory. Such bacterial classification is somewhat artificial in that they are categorized according to phenotypic (as opposed to genotypic) features, which facilitate their laboratory identification. These comprise:
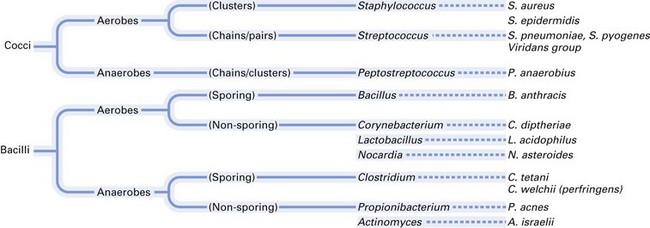
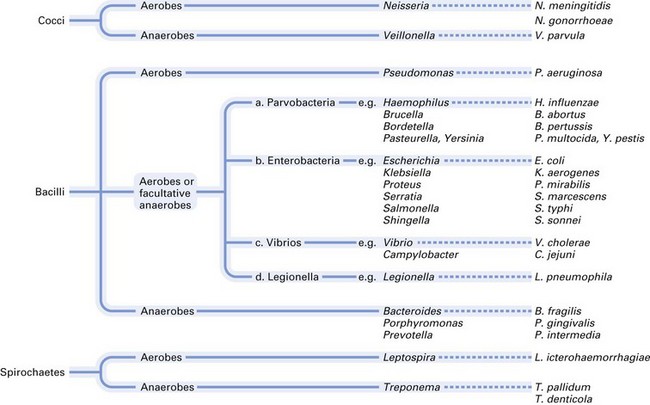
Most of the medically and dentally important bacteria are classified according to their morphology, Gram-staining characteristics and atmospheric requirements. A simple classification of medically important bacteria is given in Figures 2.7 and 2.8.
Genotypic taxonomy
In contrast to the classical phenotypic classification methods outlined above, genotypic classification and speciation of organisms are becoming increasingly important and useful. Genotypic taxonomy exploits the genetic characteristics, which are more stable than the sometimes transient phenotypic features of organisms. These methods essentially evaluate the degree of DNA homology of organisms in order to speciate them, for example by assessing molecular guanine and cytosine (GC) content, ribotyping, random amplification of polymorphic DNA (RAPD) analysis and pulsed-field gel electrophoresis (PFGE). Novel bacterial typing methods based on the nucleotide sequences of ribosomal RNA (rRNA) genes have become a robust way of assessing bacterial identity. Further details of these methods are given in Chapter 3.
Additionally, recent research indicates that endogenous bacterial habitats in humans, including the oral cavity, harbour a flora that cannot be cultured using routine laboratory techniques. These so-called unculturable species comprise both bacteria and archaea, mentioned above and can only be detected by molecular techniques or metagenomics (e.g. by direct amplification of 16S RNA). The role of these totally new phylotypes of bacteria in either disease or health awaits clarification.
Both the culturable and unculturable organisms in the healthy oral cavity are now given the term ‘core microbiome’. The analysis of this core microbiome has been greatly facilitated by a recently developed technique called pyrosequencing (a method of DNA sequencing). The data from pyrosequencing studies have revealed that the oral cavity in health may contain more than 1000 different bacterial species (see Chapter 31)!
How do organisms get their names?
Organisms are named according to a hierarchical system, beginning with the taxonomic rank domain, followed by kingdom, phylum, class, order, family, genus and species (Table 2.3). The scientific name of an organism is classically a binomial of the last two ranks, i.e. a combination of the generic name followed by the species name, e.g. Streptococcus salivarius (note that the species name does not begin with a capital letter). The name is usually written in italics with the generic name abbreviated (e.g. S. salivarius). When bacterial names are used adjectivally or collectively, the names are not italicized and do not begin with a capital letter (e.g. staphylococcal enzymes, lactobacilli).
Table 2.3 Hierarchical ranks in classification of organisms
| Taxonomic rank | Example |
|---|---|
| Domain | Bacteria |
| Kingdom | Bacteria |
| Phylum | Firmicutes |
| Class | Bacilli |
| Order | Lactobacillales |
| Family | Lactobacillaceae |
| Genus | Lactobacillus |
| Species | Lactobacillus acidophilus |
Key facts
Note: clinically relevant facts and practice points are italicized; key words are in bold.
Crielaard W. Pyrosequencing analysis of the oral microflora of healthy adults. Journal of Dental Research. 2008;87:1016-1020.
Mims C., Playfair J., Roitt I., Wakelin D., Williams R. Microbes and parasites; and the host-parasite response, Chs 1 and 2. Medical microbiology, 2nd ed. London: Mosby. 1998.
Murray P.R., Rosenthal K.S., Kobayashi G.S., Pfaller M.A. Bacterial morphology and cell wall structure and synthesis; and bacterial metabolism and growth, Chs 3 and 4. Medical microbiology, 3rd ed. St Louis: Mosby Year Book. 1998.
Parahitiyawa N., Scully C., Leung W., Yam W., Jin L., Samaranayake L.P. Exploring the oral bacterial flora: current status and future directions. Oral Diseases. 2010;16:136-145.
Raoult D. The journey from Rickettsia to mimivirus. ASM News. 2005;71:278-284.
Villareal L.P. Viruses and the evolution of life. Washington, DC: ASM Press; 2005.
Wade W.G. Non-culturable bacteria in complex commensal populations. Advances in Applied Microbiology. 2004;54:93-106.
Review questions (answers on p. 351)
Please indicate which answers are true, and which are false.