Chapter 6 Diagnostic microbiology and laboratory methods
Diagnostic microbiology
Diagnostic microbiology involves the study of specimens taken from patients suspected of having infections. The end result is a report that should assist the clinician in reaching a definitive diagnosis and a decision on antimicrobial therapy. Hence, clinicians should be acquainted with the techniques of taking specimens, and understand the principles and techniques behind laboratory analysis.
The diagnosis of an infectious disease entails a number of decisions and actions by many people. The diagnostic cycle begins when the clinician takes a microbiological sample and ends when the clinician receives the laboratory report and uses the information to manage the condition (Fig. 6.1). The steps in the diagnostic cycle are:
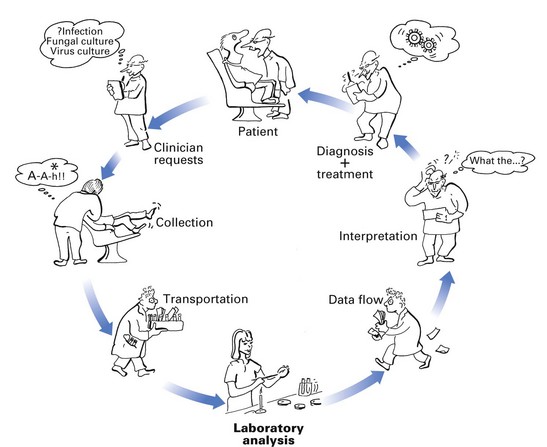
Fig. 6.1 The cycle of important events in diagnostic microbiology, depicting the interaction between the clinician and the microbiology laboratory.
Clinical request
The first stage in the diagnostic cycle comprises the specimen and the accompanying request form. The following, which influence the quality of the specimen, should be noted:
Provision of clinical information
The appropriate tests for each specimen have to be selected by the microbiologist according to clinical information given in the accompanying request form. Hence, information such as age, main clinical condition, date of onset of illness, recent/current antibiotic therapy, antibiotic allergies and history of previous specimens are all important for the rationalization of investigations and should be supplied with the specimen.
Collection and transport of specimens
Always collect appropriate specimens.
Specimens should be as fresh as possible: many organisms (e.g. anaerobes, most viruses) do not survive for long in specimens at room temperature. Others, such as coliforms and staphylococci, may multiply at room temperature, and subsequent analysis of such specimens will give misleading results.
Transport specimens in an appropriate medium (see below), otherwise dehydration and/or exposure of organisms to aerobic conditions occurs, with the resultant death and reduction in their numbers. The transport medium should be compatible with the organisms that are believed to be present in the clinical sample (e.g. virus specimens should be transported in viral transport medium, which is not suitable for bacteriological samples). Transport specimens in safe, robust containers to avoid contamination.
Laboratory analysis
A wide array of specimens are received and analyzed by a number of methods in diagnostic microbiology laboratories. The analytical process of a pus specimen from a dental abscess is given below, as an illustration (Fig. 6.2):
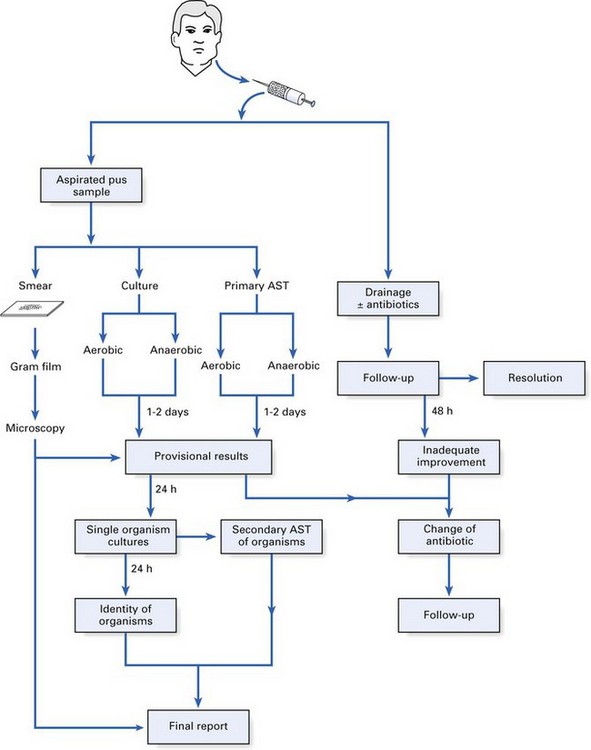
Fig. 6.2 Laboratory analysis of a pus specimen illustrating the interactions between the laboratory and the clinician. AST, antibiotic sensitivity test.
Finally, it should be noted that the microbiologist can issue a provisional report after 2 days but the final report may take longer (Fig. 6.2).
Interpretation of the microbiology report and use of information
While interpretation of most microbiology reports may be straightforward, there are situations in which the clinician should contact the microbiologist, e.g. for guidance in relation to antibiotic therapy and the necessity for further sampling. Good collaboration between the clinician and the microbiologist is essential to achieve optimal therapy.
Laboratory methods
A number of methods and techniques are used in the laboratory diagnosis of infection; they can be broadly categorized into:
Microscopic methods
Light microscopy
Bright-field or standard microscopy
Routinely used in diagnostic microbiology, stained smears from lesions are examined with the oil immersion objective (×100) using the ×10 eyepiece, yielding a magnification of ×1000. Wet films are examined with a dry objective (×40) (e.g. to demonstrate motility of bacteria).
Dark-ground microscopy
The specimen is illuminated obliquely by a special condenser so that the light rays do not enter the objective directly. Instead, the organisms appear bright, as the light rays hit them, against the dark background.
Phase-contrast microscopy
Although rarely employed in diagnostic microbiology, this technique may be used to define the detailed structure of unstained microbes.
Fluorescence microscopy
Fluorescence techniques are widely used, especially in immunology. This method employs the principle of emission of a different wavelength of light when light of one wavelength strikes a fluorescent object. Ultraviolet light is normally used, and the bacteria or cells are stained with fluorescent dyes such as auramine; for example, to detect microbial antigens in a specimen, the latter is ‘stained’ with specific antibodies tagged with fluorescent dyes (immunofluorescence; see below).
Electron microscopy
In electron microscopy, light waves are replaced by a beam of electrons, which allows resolution of extremely small organisms such as virions, e.g. 0.001 µm. Electron microscopy can be used in diagnostic virology, for instance, for direct examination of specimens (e.g. rotavirus, hepatitis A virus). Approximately 1 million virus particles are needed for such visualization. Clumps of such viral particles can be obtained by reacting the sample with antiviral antibody – immunoelectron microscopy.
Light microscopy and stains
In light microscopy, bacterial stains are used:
The most commonly used stain in diagnostic microbiology is the Gram stain.
Gram stain technique
Ziehl–Neelsen technique
Some bacteria, such as tubercle bacilli, are difficult to stain by the Gram method because they possess a thick, waxy outer cell wall. Instead, the Ziehl–Neelsen technique is used. The organisms are exposed to hot, concentrated carbolfuchsin for about 5 min, decolourized with acid and alcohol (hence, the term acid- and alcohol-fast bacilli), and finally counterstained with methylene blue or malachite green. The bacilli will stain red against a blue background.
Detection of microbes by probing for their genes
Polymerase chain reaction
Very small bacterial numbers (10–100) in patient specimens can be detected using the standard polymerase chain reaction (PCR) techniques (Chapter 3), while more sophisticated techniques can detect one human immunodeficiency virus (HIV) proviral DNA sequence in 106 cells. The main advantage of this method is its rapidity (a few hours compared with many days for conventional cultural techniques). However, PCR reactions may yield non-specific data and hence judicial selection of primers and careful conduct of the assays (to prevent contaminants giving rise to false-positive results) are important. For these reasons, PCR techniques are not common in the diagnostic laboratory, but with new developments such as microarray technology and nested PCR, it is only a matter of time before this technique becomes more popular.
Nucleic acid probes
In this technique, a labelled, single-stranded nucleic acid molecule is used to detect a complementary sequence of DNA of the pathogen in the patient sample, by hybridizing to it. The probes are obtained in the first instance from naturally occurring DNA by cloning DNA fragments into appropriate plasmid vectors and then isolating the cloned DNA. However, if the sequence of the target gene (in the pathogen) is known, oligonucleotide probes can be synthesized and labelled with a radioactive isotope or with compounds that give colour reactions under appropriate conditions.
This technique is not sensitive for detecting small numbers of organisms (i.e. few copy numbers of the gene) in clinical samples. However, a combination of the PCR technique (to produce high copy numbers) and hybridization with an oligonucleotide probe is likely to be the method of choice in identifying organisms that are slow or difficult to grow in the laboratory.
Cultural methods
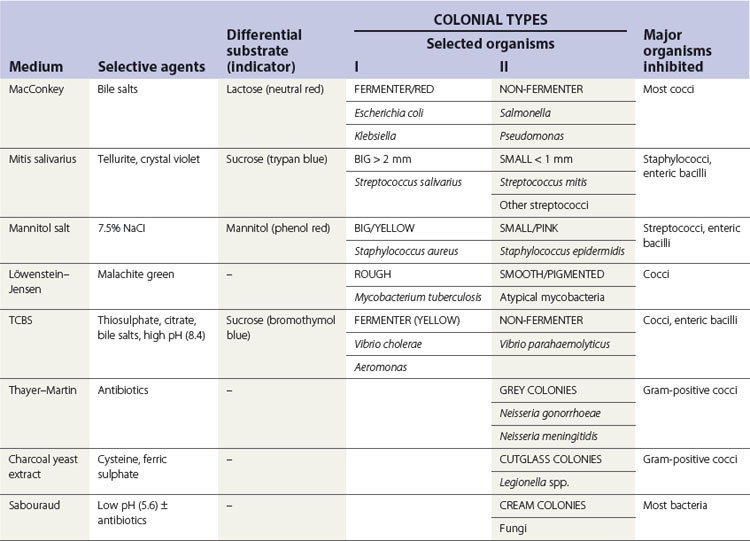
Bacteria grow well on artificial media, unlike viruses that require live cells for growth. Blood agar is the most widely used bacterial culture medium. It is an example of a non-selective medium as many organisms can grow on it. However, when chemicals are incorporated into media to prevent the growth of certain bacterial species and to promote the growth of others, selective media can be developed (e.g. the addition of bile salts helps the isolation of enterobacteria from a stool sample by suppressing the growth of most gut commensals). Some examples of selective media and their use are given in Table 6.1.
Bacteriological media
The main constituents of bacteriological media are:
Preparation of solid media and inoculation procedure
When all the necessary ingredients have been added to the molten agar, it is dispensed, while still warm, into plastic or glass Petri dishes. The agar will gradually cool and set at room temperature, yielding a plate ready for inoculation of the specimen.
The objective of inoculating the specimen or a culture of bacteria on to a solid medium is to obtain discrete colonies of organisms after appropriate incubation. Hence, a standard technique (Fig. 6.3) should be used. Solid media are more useful than liquid media as they facilitate:
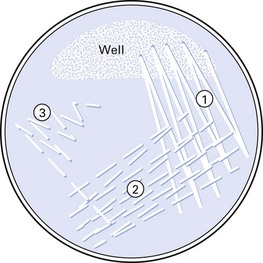
Fig. 6.3 Method of inoculating an agar plate to obtain discrete colonies of bacteria (numbers indicate inoculation steps).
Liquid media
Liquid media are used in microbiology to:
Some examples of solid and liquid media are given in Table 6.2.
Table 6.2 Constituents and uses of some commonly used solid and liquid media
| Medium | Major ingredients | Use |
|---|---|---|
| Solid media | ||
| Nutrient agar | Nutrient broth, agar | General purpose |
| Blood agar | Nutrient agar, 5–10% horse or sheep blood | Very popular, general use |
| Chocolate agar | Heated blood agar | Isolation of Haemophilus and Neisseria spp. |
| CLED agar | Peptone, l-cystine, lactose, etc. | Culture of coliforms |
| Antibiotic sensitivity | Peptone and a semisynthetic medium | Antibiotic sensitivity tests |
| Liquid media | ||
| Peptone | Peptone, sodium chloride, water | General use; base for sugar fermentation tests |
| Nutrient broth | Peptone water, meat extract | General culture |
| Robertson’s meat medium | Nutrient broth, minced meat | Mainly to culture anaerobes |
| Selenite F broth | Peptone, water, sodium selenite | Enrichment medium for Salmonella and Shigella spp. |
CLED, cystine–lactose–electrolyte-deficient.
Media for blood culture
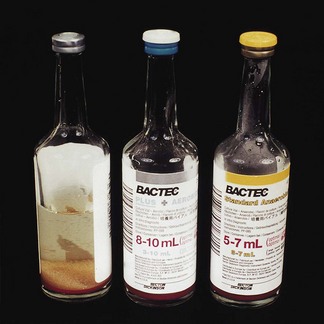
When the infectious agent is circulating in blood (e.g. in septicaemia, endocarditis, pneumonia), the latter has to be aseptically withdrawn by venepuncture and cultured. Blood culture has to be performed on special liquid media, under both aerobic and anaerobic conditions. The blood is aseptically transferred to a rich growth medium (e.g. brain–heart infusion broth) containing anticoagulants (Fig. 6.4). Cultures are checked for turbidity and gas production daily, up to a week (in many laboratories, this process is now automated and machines are used to detect bacterial growth). Positive cultures are sampled, and the organisms are isolated and identified.
Transport media
Specimens are transported from the clinic to the laboratory in a transport medium, which helps to maintain the viability of the organisms in transit.
Atmospheric requirements and incubation
Once inoculated, the agar plates may be incubated:
Bacterial identification
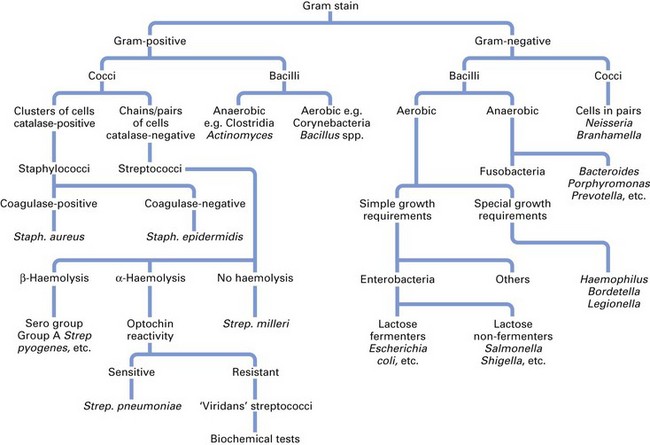
When the putative pathogen from the clinical specimen is isolated as a pure culture, it is important to identify the organism(s). Bacterial identification (Fig. 6.5) initially entails:
The foregoing will indicate the major group to which the organism belongs (e.g. streptococci, enterobacteria, clostridia). However, definitive identification to species level requires biochemical tests.
Biochemical tests
Each bacterial species has a characteristic biochemical profile valuable for its identification. These include:
Commercial identification kits
Definitive identification of an organism requires testing for a spectrum of enzymes as well as its ability to ferment (anaerobic breakdown) or assimilate (aerobic breakdown) a number of carbohydrates. This is facilitated by commercially available kits, such as the API (Analytical Profile Index) and AnIdent systems, which incorporate a wide range of the foregoing tests (usually 20) in a single kit system (Fig. 6.6).
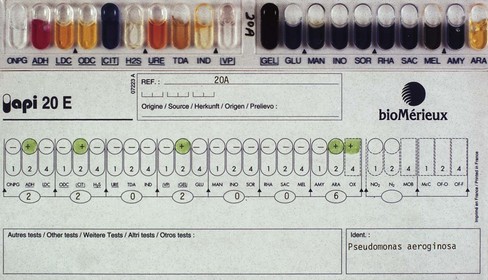
Fig. 6.6 A commercial identification kit of 20 biochemical tests used to speciate (identify to the species level) enterobacteria. Similar kits are used to speciate other genera of bacteria.
Method
A pure culture of the test organism is inoculated into each small well (cupule) containing the appropriate carbohydrate or the chemical and incubated overnight. The resultant colour or turbidity change for each test is then compared with a standard colour chart (provided by the manufacturers) and scored. The numerical profile thus obtained for the organism is compared with a profile compiled from type cultures, and the degree of concordance between the profiles of the two organisms enables identification of the test bacterium.
Sometimes the process of identifying an organism has to be extended further than speciation (i.e. identifying the bacteria beyond the species level) described above; this is called bacterial subtyping.
Subtyping organisms
It is important to realize that organisms belonging to the same species may have different characteristics (just as individual members of the species Homo sapiens vary in characteristics such as skin colour, stature, etc.). This is especially important when tracing the epidemic spread of an organism either in the community or in a hospital ward (like tracing a criminal in a vast population). Tracing such an organism can be performed by strain differentiation using the following typing procedures:
Genetic typing
A number of novel genetic typing methods such as those described in Chapter 3 are now available, and these produce very accurate ‘fingerprints’ of bacteria. These methods are gradually supplanting the foregoing traditional subtyping methods and are likely to replace them in a few years’ time. As genetic typing methods are highly discriminatory compared with the foregoing, they are used both in the diagnostic and research laboratories to detect clonality of organisms with respect to microbes from a common-source outbreak. If an infectious organism arises from a single parent cell, then in order to detect the lineage of the progeny daughter cells that are, for all intents and purposes, genetically identical, a number of detection methods can be used. These include:
Immunological methods
Immunological methods are useful in diagnostic microbiology to identify organisms and to detect antibodies in a patient’s body fluids (e.g. serum, saliva), especially when the organism cannot be cultured in laboratory media.
Identification of organisms using immunological techniques
Agglutination
Slide agglutination
Antibodies against the specific serotypes of the organism (e.g. Salmonella and Shigella species) can be used in identification. When a suspension of the organisms and a few drops of the specific antibody are mixed on a glass slide, visible agglutination (clumping) of the organism indicates a positive reaction.
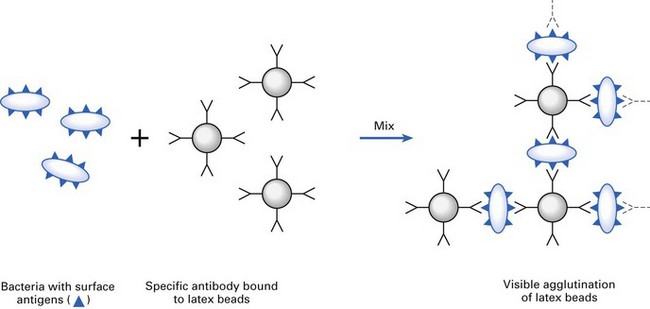
Latex agglutination
Here the agglutination of latex beads coated with the specific antibody directed against the unknown organism is used, as above (e.g. Neisseria meningitidis, Haemophilus influenzae, the yeast Cryptococcus neoformans) (Fig. 6.7).
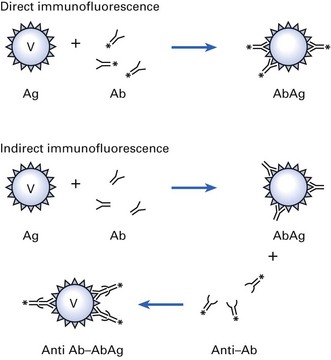
Immunofluorescence
If an organism is exposed to the specific antibody tagged with a fluorescent dye, then the organism binds to the antibody and can be visualized through an ultraviolet microscope. Principles of direct (one-step) and indirect (two-step) immunofluorescence techniques are shown in Figure 6.8.
Enzyme-linked immunosorbent assay
The enzyme-linked immunosorbent assay (ELISA) is a modification of the above test in which the fluorescent dye tagged to the antibody is replaced by an enzyme. The organism binds to the antibody and the tagged enzyme, and the amount of bound enzyme can then be demonstrated by reaction with the enzyme substrate. This is a highly popular test.
Detection of antibodies in a patient’s serum
An example of this technique is the serological tests for syphilis. The agent of syphilis, Treponema pallidum, does not grow in laboratory media. Hence, serological tests are useful. These are:
Laboratory investigations related to antimicrobial therapy
Once the putative pathogen has been identified from a specimen, its antimicrobial sensitivity can be predicted with some degree of accuracy, based on previous experience and available data. Prescribing in this manner is called empirical therapy (e.g. based on the sensitivity of staphylococci to flucloxacillin). However, it is essential to base rational therapy on the results of laboratory antibiotic tests performed on the isolated pathogen.
Susceptibility of organisms to antimicrobial agents
In clinical microbiology, a microbe is considered sensitive (or susceptible) to an antimicrobial agent if it is inhibited by a concentration of the drug normally obtained in human tissues after a standard therapeutic dose. The reverse is true for a resistant organism. Organisms are considered intermediate in susceptibility if the inhibiting concentration of the antimicrobial agent is slightly higher than that obtained with a therapeutic dose.
Laboratory testing for antimicrobial sensitivity
The action of an antimicrobial drug against an organism can be measured:
A semiquantitative technique, called the break-point test, is not described here. These in vitro tests indicate whether the expected therapeutic concentration of the drug given in standard dosage inhibits the growth of a given organism in vivo. Laboratory results can only give an indication of the activity of the drug in vitro, and its effect in vivo depends on factors such as the ability of the drug to reach the site of infection and the immune status of the host. A strong host defence response may give the impression of ‘successful’ drug therapy, even though the infecting organism was ‘resistant’ to a specific drug when laboratory tests were used.
Disc diffusion test
The disc diffusion test is the most commonly used method of testing the sensitivity of a microorganism to an antimicrobial agent. Here, the isolate to be tested is seeded over the entire surface of an agar plate, and drug-impregnated filter paper discs are applied. After overnight incubation at 37°C, zones of growth inhibition are observed around each disc, depending on the sensitivity of a particular organism to a given agent (Figs 6.9 and 6.10).
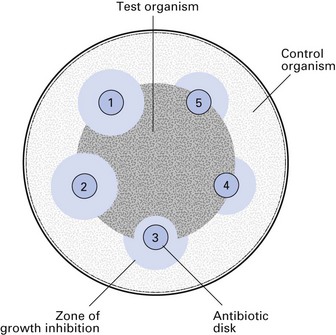
Fig. 6.9 The antibiotic susceptibility of an organism can be tested by an application of filter-paper discs impregnated with different antibiotics onto a lawn of the organism seeded on an agar plate. After overnight incubation, zones of growth inhibition around discs indicate sensitivity to the antibiotic, whereas growth of the organism up to the disc indicates resistance. In this example, the test organism is sensitive to antibiotics 1 and 2, moderately sensitive to antibiotic 3, and resistant to antibiotics 4 and 5.
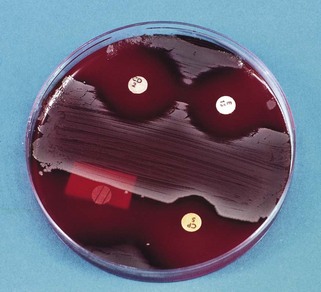
Fig. 6.10 Another example of an antibiotic sensitivity test; here the control organism is inoculated on the polar aspects of the plate and the test organism is inoculated in the middle. In this example, the organism is resistant to ampicillin (AM disc, bottom left) and sensitive to the other three antibiotics (AM, ampicillin; CD, clindamycin; CP, cephalosporin; E, erythromycin).
Antimicrobial sensitivity tests of this type can be divided into primary sensitivity (direct) and secondary sensitivity (indirect). A primary test is carried out by inoculating the clinical sample, say pus, directly on to the test zone of the plate. The advantage of this is that the overall sensitivity results for the organisms present in pus will be available after 24- to 48-h incubation (see Fig. 6.2). This is particularly useful when treating debilitated patients with acute infections such as dentoalveolar abscesses. However, because this is a rough estimate, secondary sensitivity tests are therefore performed on a pure culture of the isolated organism, but the results are not available for at least 2–4 days after sampling.
Assessment of MIC and MBC
Determining the MIC and MBC gives a quantitative assessment of the potency of an antibiotic (Fig. 6.11).
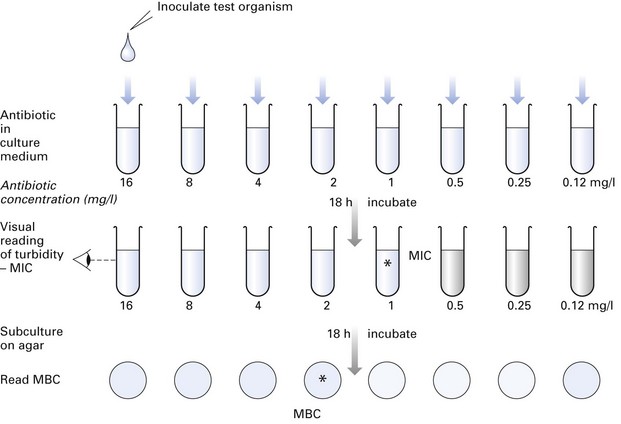
Fig. 6.11 Determination of the minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) of an antibiotic required to inhibit or kill a specific organism. This gives a quantitative estimate of the antibiotic sensitivity of the organism, compared with the disc diffusion method described in Figure 6.9. (In this example, MIC = 1 mg/l, MBC = 2 mg/l.)
Method
A range of twofold dilutions of an antimicrobial agent can be incorporated into a suitable broth in a series of tubes (tube dilution technique). The broth is inoculated with a standardized suspension of the test organism and incubated for 18 h. The minimum concentration of the drug that inhibits the growth of the test organism in the tube is recorded as the MIC, i.e. the lowest concentration that will inhibit the visible growth in vitro. Subsequently, a standard inoculum from each of the tubes in which no growth occurred may be subcultured on blood agar to determine the minimum concentration of the drug required to kill the organism (MBC). The MBC is defined as the minimum concentration of drug that kills 99.9% of the test microorganisms in the original inoculum.
These tests are not routinely performed but are useful in patients with serious infections where optimal antimicrobial therapy is essential, for example, to establish sensitivity of streptococci isolated from blood cultures from patients with infective endocarditis, and of bacteria causing septicaemia in immunosuppressed patients.
Appropriate specimens in medical microbiology
See Table 6.3.
Table 6.3 Some appropriate specimens for microbiological investigations
| Tissue or system | Specimen | Comments |
|---|---|---|
| Skin | Swab | Examine for bacteria and yeasts |
| Scrapings | Examine for fungi | |
| Vesicle fluid | Examine for viruses (electron microscopy and culture) | |
| Serum | Viral serology | |
| Blood (bacteraemia and septicaemia) | Blood culture | Sterile precautions necessary; multiple specimens required |
| Gastrointestinal tract | Faeces | Culture for bacteria and viruses; toxin detection for Clostridium difficile; light microscopy for parasites and protozoa; electron microscopy for viruses |
| Serum | Serological tests for enteric fevers | |
| Urinary tract | Midstream specimen of urine/suprapubic aspirate/catheter specimen of urine (not from a collecting bag) | For quantitative and qualitative bacteriology |
| Upper respiratory tract | Pernasal, throat and nose swabs; saliva | Culture for Bordetella pertussis; culture for β-haemolytic streptococci, other bacteria and viruses |
| Throat washings or nose and throat aspirates | Culture and immunofluorescence for viruses | |
| Lower respiratory tract | Sputum | Culture for bacteria, viruses and fungi; fluorescent microscopy for many viruses, Mycobacterium tuberculosis and Legionella spp. |
| Serum | Viral and fungal serology | |
| Meninges | Cerebrospinal fluid | Cell count, microscopy and culture |
| Serum | Viral serology | |
| Genital tract | Swab in Amies’ or Stuart’s transport medium | For bacterial and yeast culture and microscopy for gonococci and Trichomonas spp. (wet film) |
| Swabs in Chlamydia and viral transport medium | Culture of Chlamydia and viruses | |
| Smear of discharge | For detection of gonococci | |
| Serum | Serological test for syphilis | |
| Abscess | Pus | Aspirates for culture and identification |
| Wounds | Pus or swab | Avoid contamination from skin; pus preferred |
| Tissue | Send small samples in dry sterile containers for homogenization, culture and microscopy | |
| Mucosal lesions | Swab | Avoid contamination with normal flora. Use transport medium if necessary; culture for bacteria, fungi and viruses |
| Smear | Fluorescent microscopy; useful for gonococci and yeasts | |
| Serum | Serological tests for staphylococcal and streptococcal infection and viruses |
Modified from Ross, PW, and Hollbrook, WP (1984). Clinical and oral microbiology. Blackwell, Oxford.
Appropriate specimens for oral infections
Sampling for pathogens within the oral environment poses many problems due to the multitude of indigenous commensal flora that thrive in the oral cavity. Further, many of the pathogens are endogenous in origin and cause disease when an opportunity arises (opportunistic pathogens). In addition, obtaining an uncontaminated sample from sites such as the depths of periodontal pockets where disease activity, and hence the numbers of periodontopathogens, are likely to be high is extremely difficult. For these reasons, judicial and appropriate sampling techniques should be used when diagnosing oral infections (Table 6.4).
Table 6.4 Appropriate specimens for microbiological examination of oral infections
| Lesion or site of lesion | Specimen | Comments |
|---|---|---|
| Lips and perioral skin | Moistened swab | Culture for yeasts and bacteria |
| Vesicle fluid, swab | Virus culture and electron microscopy | |
| Aspirate of abscess | Microscopy and culture (see Fig. 6.2) | |
| Serum | Serological tests for viruses and syphilis | |
| Tongue and oral mucosa | Swab | Culture for bacteria, yeasts and viruses |
| Smear of scraping (heat-fixed) | Microscopy for yeasts and bacteria | |
| Vesicle fluid | Microscopy for yeasts and bacteria | |
| Biopsy tissue | Culture for bacteria and viruses: microscopy for yeasts and suspected tuberculosis | |
| Serum | Culture for bacteria and viruses: microscopy for yeasts and suspected tuberculosis | |
| Dental abscess or suspected infected cyst | Aspirate | Smear and culture (see Fig. 6.2) |
| Infected root canal | Paper point or barbed broach | Aseptic collection; use semisolid transport medium; semiquantitative culture |
| Dental plaque | Scraping | A variety of sampling tools and procedures available |
| Gingivae and gingival crevice | Scraping on a sterile sealer | Smear can be diagnostic for fusospirochaetal infection; viral culture possible, DNA tests, BANA tests for periodontopathogens |
| Severe caries | Saliva | Lactobacillus/Streptococcus mutans counts |
| Prosthesis (dentures) | Swab and smear | In suspected denture stomatitis, examine for yeasts |
BANA, N-benzoyl-dl-arginine-2-naphthylamide.
The specimens submitted to an oral microbiology laboratory can be categorized as those useful for the management of purulent infections, mucosal infections, and periodontal infections and caries.
Purulent infections
The appropriate specimen is an aspirated sample of pus, if possible. Take care to avoid needlestick injuries when resheathing the needle cap; drainage of residual pus by incision, after aspiration sampling, is obligatory. The laboratory steps in the diagnosis of a purulent infection are shown in Figure 6.2.
Mucosal infections
A common oral mucosal infection is oral candidiasis. Here, the lesion is sampled with a dry swab, and a smear taken immediately thereafter (see the section on Candidal infections).
When evaluating the oral carriage of yeasts (or other organisms such as Enterobacteriaceae), then an oral rinse should be collected. This entails requesting the patient to rinse the mouth for 60 s with 10 ml of phosphate-buffered saline and then expectorating the rinse into a container, which is transported to the laboratory for quantification of yeast growth (in terms of CFUs).
Diagnosis of viral infections of the oral mucosa is described below.
Periodontal infections and caries
The value of microbiological sampling for the diagnosis of caries and periodontal diseases is limited. In the case of dental caries, salivary counts of lactobacilli and Streptococcus mutans could be used, and for this purpose, saliva samples should be collected (Chapter 32).
The diagnosis of periodontal disease by microbiological means is problematic. A deep gingival smear is useful for the diagnosis of acute necrotizing ulcerative gingivitis, while paper point samples appear useful for DNA analysis of periodontopathic bacteria. However, the latter is not a conclusive test.
Laboratory isolation and identification of viruses
The techniques for isolation and identification of viruses are significantly different from bacteriological techniques. Laboratory procedures for the diagnosis of viral infections are of four main types:
Direct microscopy of clinical material
Direct microscopy is the quickest method of diagnosis. Virus or virus antigen may be detected in tissues from lesions, aspirated fluid samples or excretions from the patient. The common techniques used are:
Isolation and identification from tissues
Viruses do not grow on inanimate media, and they must be cultivated in living cells. Since no single type of host cell will support the growth of all viruses, a number of different methods of culturing viruses have been developed:
Tissue culture
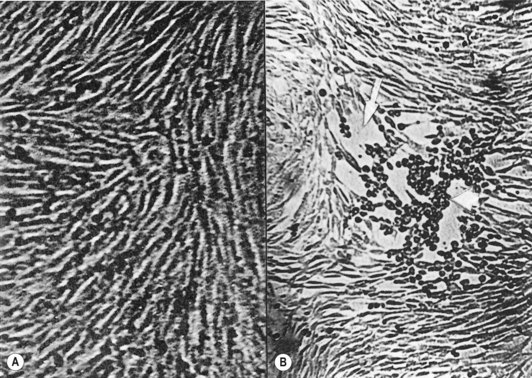
After the inoculation of a monolayer of tissue culture with a clinical sample, it is examined daily for microscopic evidence of viral growth, for about 10 days. Viruses produce different kinds of degenerative changes or cytopathic effects, such as rounding of cells and net or syncytial formation, in susceptible cells (Fig. 6.12). The cell type supporting virus growth and the nature of the cytopathic effect help identification of individual viruses (e.g. herpesviruses growing in monkey kidney cells produce fused cells in which nuclei aggregate to form multinucleate giant cells). The time required for the cytopathic effect to be seen can vary from 24 h up to several days, depending on the virus strain and the concentration of the inoculum. Once the virus is cultured, it can be identified by:
Serodiagnosis of viral infections
Many virus infections produce a short period of acute illness in which viral shedding occurs, and thereafter, it is difficult to culture viral samples from clinical specimens. Hence, diagnosis of viral infections by serology is widely used. A diagnosis of a recent viral infection depends on:
Serological test results are interpreted by comparing the antibody titres of the acute and convalescent sera. Antibody titre is defined as the reciprocal of the highest serum dilution that shows antibody activity, in a given test (e.g. if the patient’s serum shows antibody activity when diluted by 1 in 64, then the antibody titre is 64). A greater than fourfold rise in titre between the acute and convalescent samples is considered to be a positive result, indicating the patient has had an acute illness due to the specific virus.
Serological tests
A wide array of serological tests are used in virology. The classic and oldest technique of complement fixation is now being supplanted by a number of other tests. These include immunofluorescence (see Fig. 6.8), ELISA and radioimmunoassay. The advantages of the latter methods over the comparison of viral titre in paired sera, described above, are that only a single serum sample is needed and results are available quickly. Immunological and tissue culture detection methods have been successfully combined to shorten the time required to identify viral infections. Other methods frequently used in serodiagnosis of viral infections include haemagglutination.
Serodiagnosis using multiple antigen systems
Some viruses, such as mumps virus and hepatitis B virus, present with more than a single antigen (and hence antibody), which appear at different periods of the illness. This feature can be exploited to detect the state of illness by using a single sample of serum without waiting for convalescence. A variety of antigens and antibodies used in the detection of various phases of hepatitis B virus infection is described in some detail in Chapter 29.
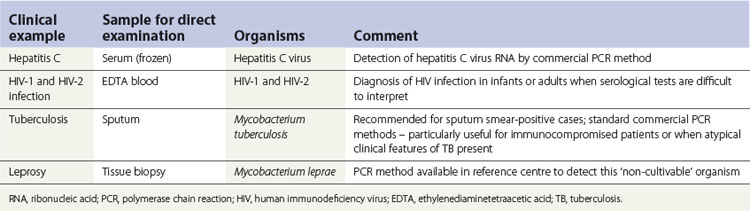
Molecular amplification methods for rapid viral diagnosis
Molecular methods are increasingly useful and should gradually supplant conventional methods of viral detection, as has already been discussed in Chapter 4. For example, the PCR technique can detect even a few DNA molecules of a specific virus in a sample. Also, radioactive virus DNA can detect virus genome or mRNA in tissues by molecular hybridization (Table 6.5).
Diagnosis of fungal infections
These principles of diagnosis of fungal diseases are essentially the same as for bacterial and viral infections. Fungal diseases can be diagnosed by:
Candidal infections
Smears, swabs and oral rinse samples are the common specimens received in the laboratory for the diagnosis of oral candidal infections. For this, the lesion is sampled with a dry swab and a smear is taken immediately thereafter (a smear is taken by scraping the lesion with the edge of a flat plastic instrument and transferring the sample to a glass microscope slide). In patients with possible Candida-associated denture stomatitis, a smear of the fitting surface of the denture as well as a swab should be taken.
In the laboratory, the smear is stained with the Gram stain or periodic acid-Schiff (PAS) reagent and examined microscopically to visualize the hyphae and/or blastospores (synonym: blastoconidia, yeast phase) of Candida. Their presence in large numbers suggests infection. The swabs are cultured on Sabouraud medium and incubated for 48–72 h, when Candida albicans appears as cream-coloured large convex colonies. Other species of Candida co-infecting with C. albicans (e.g. Candida glabrata, Candida krusei) can be identified if the specimen is cultured in commercially available media, such as CHROMagar or Pagano–Levin agar, in which different species produce colonies with varying colours and hues (Fig. 6.13).
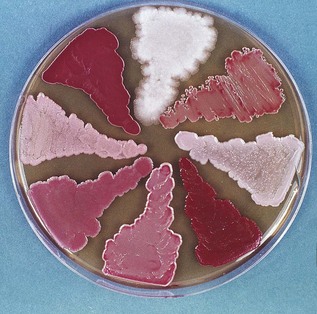
Fig. 6.13 Growth of different Candida species on Pagano–Levin agar exhibiting varying colony colours and hues.
Yeasts so derived are speciated by sugar fermentation and assimilation tests and the germ tube test. The latter is a useful quick test to differentiate C. albicans and C. dubliniensis from the other Candida species such as C. glabrata and C. krusei.
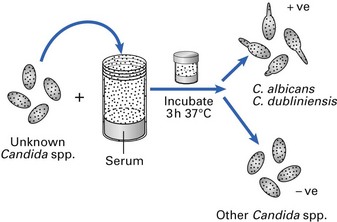
Germ tube test
A small inoculum of the isolated yeast is incubated in serum at 37°C for about 3 h and a few drops of the suspension are then examined microscopically. Virtually all strains of C. albicans and C. dubliniensis produce short, cylindrical extensions termed ‘germ tubes’, as opposed to the other Candida species, which do not exhibit this characteristic (Figs 6.14 and 6.15).
Histopathology
Incisional and excisional biopsies are useful in the diagnosis of persistent oral white lesions thought to be related to candidal infection. As a significant proportion of chronic candidal leukoplakic lesions are premalignant, a biopsy in addition to a swab is essential if the lesion does not resolve after antifungal therapy (see Chapter 35).
Other laboratory investigations
On occasions, chronic candidal infections are associated with nutritional and haematological abnormalities and appropriate laboratory investigations (e.g. iron, vitamin levels) should also be carried out.
Key facts
Collee J.G., Fraser A.G., Marmion B.P., Simmons A. Practical medical microbiology, 14th ed. Edinburgh: Churchill Livingstone; 1996.
De la Maza L.M., Pezzlo M.T., Baron E.J. Color atlas of diagnostic microbiology. St Louis: Mosby Year Book; 1997.
Mims C., Playfair J., Roitt I., Wakelin D., Williams R. Diagnostic principles of clinical manifestations, Sect. 13. Medical microbiology, 2nd ed. London: Mosby. 1998.
Samaranayake L.P. The wastage of microbial samples in clinical practice. Dental Update. 1987;14:53-61.
Scully C., Samaranayake L.P. Clinical virology in oral medicine and dentistry, Ch. 4. Cambridge: Cambridge University Press. 1992.
Review questions (answers on p. 351)
Please indicate which answers are true, and which are false.