2 Morphology and nature of micro-organisms
Key points
• Agents of infection include cellular organisms belonging to two of the three recently defined domains of life, the Bacteria and the Eukarya. The latter include fungi and protozoa. The subcellular entities viruses, viroids and prions also cause infection but depend on host cells and tissues for propagation.
• Bacterial and eukaryotic micro-organisms can be detected by light microscopy, whereas electron microscopy is required for viruses. Adult stages of multicellular eukaryotic agents of infection or infestation, such as helminths (worms) and insects, are generally visible to the naked eye.
• Most pathogenic bacteria can be recognized as either Gram-positive or Gram-negative after staining. These properties reflect, respectively, the relatively thick peptidoglycan layer and the thin peptidoglycan plus outer membrane cell wall structures possessed by cells belonging to these two groups.
• The mycobacteria, which include the global pathogens causing tuberculosis and leprosy, have a different staining property, described as acid fast.
• In addition to the cell wall, key bacterial structures with biological and medical significance include the nucleoid, inclusion granules and spores within the cell, and flagella, fimbriae or pili, and capsules on the cell surface.
• Bacterial endospores are highly resistant bacterial cells that result from a differentiation process in some Gram-positive bacteria.
• Viruses are obligate intracellular parasites that use the host cell’s machinery to replicate. They contain a nucleic acid core comprising DNA or RNA (not both) in single- or double-stranded form.
• The core is surrounded by a protein capsid comprising multiple capsomeres; an envelope derived from the host cell membrane surrounds the capsid of some viruses.
Micro-organisms are beyond doubt the most successful forms of life; they have been here longest, they are the most numerous and their distribution defines the limits of the biosphere, encompassing environments previously thought incapable of sustaining life. Here, we are concerned with the tiny fraction of micro-organisms that form associations with human beings and these encompass the cellular entities, bacteria, archaea, fungi and protozoa, and the subcellular entities, viruses, viroids and prions. Whether the last three can be considered organisms or even living entities is a matter for debate. None the less, their transmissible nature, the immune responses they provoke and our inability to detect them with the naked eye place them firmly within the province of medical microbiology. The first two of these criteria also require us to consider some multicellular macroscopically visible organisms (members of the helminths) as agents of infection (see Ch. 63). Most of this chapter is concerned with bacteria. The subcellular entities are introduced briefly and the remaining medically significant groups are considered in more specialized chapters.
Medical microbiology has been founded on recognizing micro-organisms that are associated with human disease. This recognition has relied predominantly on two techniques:
Over the last quarter of the 20th century, it became possible to detect, describe and differentiate micro-organisms by biochemical and genetic methods, and this had two profound effects on microbiology. First, due largely to the work of Carl Woese (see Ch. 1), it is now possible to make a reasonable assessment of the evolutionary relationships between micro-organisms, a task that previously could be achieved for macro-organisms only by examining fossil records (micro-organisms have not left interpretable fossils). Woese’s approach led to the recognition of a whole ‘new’ domain (a group ranking above kingdom level) of cellular organisms, the Archaea, and a fundamental review of microbial classification (Fig. 2.1). Second, we have now amassed an enormous body nucleotide sequence and other molecular data describing microbes. Complete genome sequences are available for all the major bacterial and viral pathogens and most of the pathogenic protozoa and fungi. At the time of writing, the US National Center for Biotechnology Information provides access to 1356 bacterial genomes and, given that current technologies allow the raw data for a complete genome to be generated, within a day. We can stop counting now as it will soon be easier to obtain a complete sequence than subject a new strain of an isolate to a typing analysis. Thus morphology and cultural characteristics are regarded by many as secondary characteristics in the context of classification and identification, while molecular detection methods are steadily encroaching on microscopy and culture in clinical laboratories. Indeed genomic information has repeatedly revealed that some bacteria have structures that have not been revealed by conventional laboratory studies. The capsule (see below) of Campylobacter jejuni provides one prominent example of this.
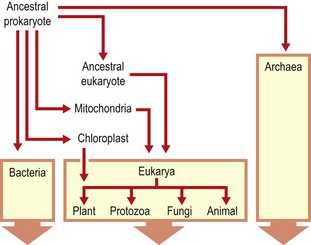
Fig. 2.1 Diagram illustrating proposed evolutionary pathways from a putative common ancestral prokaryote to the present. The Archaea were not recognized as a separate lineage until Woese’s work.
We are currently in a transitional, information gathering phase. Molecular descriptions and detection methods are coming to dominate our view of the microbial world. However, at present it must be emphasized that the morphological classification provides a basic structure that is understood by clinicians and will remain as a basis for communication, at least in the medium term. Moreover, although this will change, light and fluorescence microscopy remain competitively cheap and rapid compared with molecular methods as means of providing information relevant to the clinical management of infection. Finally, the discipline of medical microbiology requires the understanding of the basic structural properties and physiology of micro-organisms to underpin our approach to infections.
Prokaryotic and eukaryotic cells
Micro-organisms are microscopic in size and are usually unicellular. The diameter of the smallest body that can be resolved and seen clearly with the naked eye is about 100 µm. All medically relevant bacteria are smaller than this and a microscope is therefore necessary to see individual cells. When propagated on solid media, bacteria (and fungi) form macroscopically visible structures comprising at least 108 cells, which are known as colonies.
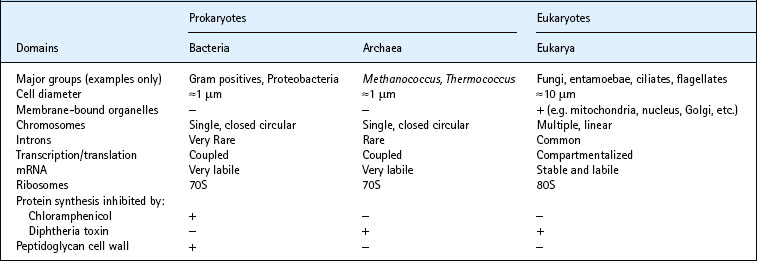
Woese’s insights provided for the first time a coherent view of the evolutionary pathways behind the diversity of all living organisms. In particular, a satisfactory explanation is offered for the existence and diversity of prokaryotic and eukaryotic cells, and all living forms are seen to fall within three domains of life: the Bacteria, the Archaea and the Eukarya. Although the first two of these are prokaryotes (organisms without a membrane-bound nucleus), the Archaea share many characteristics with the Eukarya. This division is of practical significance, as the earlier the point of divergence, the greater the difference in metabolic properties between the present-day representatives of the two lineages. These differences can be exploited by directing treatments at processes unique to the target organism. Some key differences between the three domains of life are summarized in Table 2.1.
Anatomy of the bacterial cell
The principal structures of a typical bacterial cell revealed by long established methods are shown in Figure 2.2. Over the last two decades, developments in genetic manipulation combined with advances in fluorescence and electron microscopy have opened up our understanding of microbial cytology, to the point where many key macromolecular assemblies can be described in great and dynamic detail. Some of these will be mentioned below but, for the most part, this account is rooted in well established descriptions.
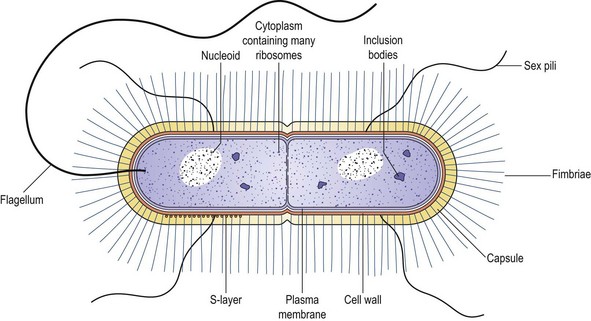
Fig. 2.2 Diagram illustrating the key features of bacterial cells. The S-layer is a variably demonstrated ordered protein layer.
The cytoplasm is bounded peripherally by a very thin, elastic and semi-permeable cytoplasmic (or plasma) membrane (a conventional phospholipid bilayer). Outside, and closely covering this, lies the rigid, supporting cell wall, which is porous and relatively permeable. Cell division occurs by the development of constrictions mediated by the assembly of an actin-like protein, FtsZ. The constrictions proceed from the periphery inwards and, in some cases, produce a transverse cell wall known as a septum or cross-wall.
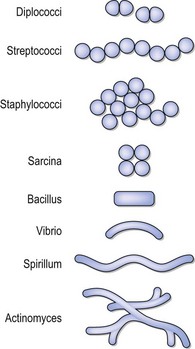
The exact pattern of cell division and the structures associated with the cell wall and cytoplasmic membrane (collectively the cell envelope) combine to produce the cell morphology and characteristic patterns of cell arrangement. The recognition of these features by oil-immersion light microscopy remains of great practical value in making presumptive identifications of bacteria associated with human infections. Bacterial cells may have two basic shapes, spherical (coccus) or rod shaped (bacillus); the rod-shaped bacteria show variants that are comma shaped (vibrio), spiral (spirillum and spirochaete) or filamentous (Fig. 2.3).
The cytoplasm is a predominantly aqueous environment packed with ribosomes and numerous other protein and nucleotide–protein complexes. The cytoplasmic contents are not normally visible by light microscopy, which can only resolve objects more than 0.2 µm in diameter. Although bacterial cytoplasm has traditionally been viewed as devoid of structure, it is now clear that, like eukarya, bacteria have an extensive cytoskeletal network which, in different genera, include structures corresponding to eukaryotic actin, tubulin and intermediate filaments (e.g. MreB, FtsZ and crescentin, respectively). The importance of these is emerging in determining cell shape, division and spore formation (see below). As predicted in the previous edition of this book, antimicrobials targeting these functions, specifically Ftz, have recently been described.
Some larger structures such as spores or inclusion bodies of storage products such as volutin (polyphosphate), lipid (e.g. poly-β-hydroxyalkanoate or triacylglycerol), glycogen or starch occur in some species under specific growth conditions. Specialized labelling techniques (generally requiring fluorescence imaging) enable visualization of the nuclear material or nucleoid and other structures (e.g. the forming cell division annulus). Figure 2.4 is an electron micrograph of a thin section of a dividing bacterial cell. Outside the cell wall there may be a protective gelatinous covering layer called a capsule or, when it is too thin to be resolved with the light microscope, a microcapsule. Soluble large-molecular material may be dispersed by the bacterium into the environment as loose slime. Some bacteria bear, protruding outwards from the cell wall, one or more kinds of protein-based filamentous appendages called flagella, which are organs of locomotion, and hair-like structures termed fimbriae or pili, which, via specific receptor–ligand interactions at their tip, mediate adhesion.

Fig. 2.4 Thin section of a dividing bacillus showing cell wall, cytoplasmic membrane, ribosomes, a developing cross-wall and a mesosome, a membranous structure now generally considered to be an artifact or reflecting some form of cell damage. Magnification ×50 000.
By courtesy of Dr PJ Highton and the editors of Journal of Ultrastructure Research.
Bacterial nucleoid
The genetic information of a bacterial cell is mostly contained (with a few exceptions) in a single, long molecule of double-stranded DNA, which can be extracted in the form of a closed circular thread about 1 mm long. The cell solves the problem of packaging this enormous macromolecule by condensing and looping it into a supercoiled state. As well as the chromosome, the bacterium may contain one or more additional fragments of episomal (extrachromosomal) DNA, known as plasmids. Bacteria are essentially haploid organisms with only one allele of each gene per cell, although there may be multiple copies of chromosomes and plasmids. Unlike the mitotic or meiotic divisions of eukaryotic cells, chromosomal segregation in bacteria at the time of cell division (or fission) does not involve structures that can be resolved by light microscopy. None the less, the speed at which replication can occur (cell divisions more frequently than one every 15 min in some cases) and the simultaneous requirement for multiple rounds of chromosome replication in one cell provide a mind-boggling challenge for the segregation machinery. (Imagine unravelling two 1 m threads inside a 1 mm sphere!)
The bacterial nucleoid lies within the cytoplasm and is associated with several nucleoid-associated proteins (NAPs). Although these were originally referred to as histone-like, they are not structurally related and they function in the cytoplasm rather than within a membrane-bound nucleus. The NAPs such as H-NS and IHF exert important influences on prokaryotic gene expression (see Chapters 4 and 6). The cytoplasmic location of the nucleoid means that as DNA-dependent RNA polymerase makes RNA, ribosomes may attach and initiate protein synthesis on the still attached (nascent) messenger RNA. Synthesis of mRNA and protein (transcription and translation) are therefore seen to be directly coupled in bacteria. In contrast, complete transcripts in eukaryotic cells have to be spliced (to remove the non-coding introns) and capped with polyadenine (this rarely occurs with bacterial mRNA) before the post-transcriptionally modified message is translocated to the cytoplasm.
Ribosomes
Bacterial ribosomes are slightly smaller (10–20 nm) than those of eukaryotic cells and they have a sedimentation coefficient of 70S, being composed of a 30S and a 50S subunit (cf. 40S and 60S in the 80S eukaryotic counterparts). They may be seen with the electron microscope, and number tens of thousands in growing cells. Multiple ribosomes attach to single mRNA molecules to form polysomes. It was the nucleotide sequencing of DNA encoding small-subunit ribosomal RNA (rDNA) that led Woese to postulate the evolutionary pathways shown in Figure 2.1. Essentially all cellular organisms can now be classified at least down to genus level by their small-subunit rRNA (SSrRNA) nucleotide sequences. Subsequently it was recognized that, as growing cells contain so many ribosomes, it should be possible to detect unique identifying (or determinative) SSrRNA sequences by complementary in-situ hybridization with fluorescently labelled oligonucleotide probes. Indeed, it is now possible to apply this approach to natural samples and this has enabled recognition of bacteria that have never been grown in laboratory culture. Finally it should be noted that two key organelles in eukaryotes, mitochondria and chloroplasts, both contain 70S ribosomes. When the SSrRNA-encoding genes on the circular chromosomes of these organelles were sequenced, their original free-living bacterial origin was clearly indicated and the proposed endosymbiotic route by which they became organelles confirmed.
Cytoplasmic membrane
The bacterial cytoplasmic membrane is 5–10 nm thick and consists mainly of phospholipids and proteins. Its structure can be resolved in some ultra-thin sections examined by electron microscopy. Membranes generally appear in suitably stained electron microscope preparations as two dark lines about 2.5 nm wide, separated by a lighter area of similar width. Integral, transmembrane, and peripheral or anchored proteins occur in abundance and perform similar functions to those described in eukaryotes (e.g. transport and signal transduction). A key feature differentiating prokaryotic cytoplasmic membranes from those of eukaryotes is their multifunctional nature. Thus, while in eukaryotic cells the endoplasmic reticulum and Golgi apparatus are involved in protein secretion, packaging and processing, and the mitochondrial inner membrane is the site of electron transport and oxidative phosphorylation, all of these functions must be performed by one membrane in prokaryotes. It is hardly surprising that prokaryotic cell membranes are relatively protein rich, allowing relatively little space for phospholipids.
Cell wall
The cell wall (Figs 2.5 & 2.6) lies immediately external to the cytoplasmic membrane. It is 10–25 nm thick, strong and relatively rigid, though with some elasticity, and openly porous, being freely permeable to solute molecules smaller than 10 kDa in mass and 1 nm in diameter. It is strong but elastic, and supports the weak cytoplasmic membrane against the high internal osmotic pressure (25 and 5 atm in Gram-positive and Gram-negative cells respectively) and maintains the characteristic shape of the bacterium in its coccal, bacillary, filamentous or spiral form.
Except under defined osmotic conditions, cell survival is dependent on the integrity of the cell wall. If the wall is weakened or ruptured, the cytoplasm may swell from osmotic inflow of water and burst the weak cytoplasmic membrane. This process of lethal disintegration and dissolution is termed lysis.
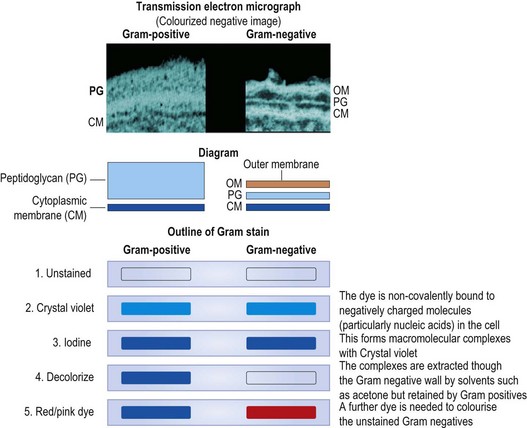
The chemical composition of the cell wall differs considerably between different bacterial species, but in all species the main strengthening component is peptidoglycan (syn. mucopeptide or murein). Peptidoglycan is composed of N-acetylglucosamine and N-acetylmuramic acid molecules linked alternately in a chain (Fig. 2.7). This heteropolymer forms a single molecular continuous sac external to the cytoplasmic membrane (described as the murein sacculus). The thickness of the peptidoglycan layer turns out to be of great practical importance in differentiating medically significant bacteria. In the late nineteenth century a Danish physician, Christian Gram, immortalized himself by devising a staining procedure that we now know distinguishes bacteria with a thick (Gram-positive) and a thin (Gram-negative) murein sacculus (see Fig. 2.5). The traditional classification of bacteria is fundamentally rooted in this dichotomy, which has, fortunately, largely been supported by rRNA-based classification.
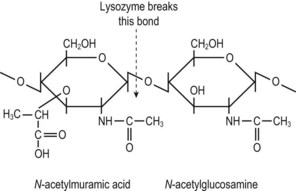
Fig. 2.7 The basic building block of bacterial cell wall peptidoglycan. N-acetylmuramic acid is derived from N-acetylglucosamine by the addition of a lactic acid unit. Each N-acetylmuramic acid molecule is substituted with a pentapeptide; an N-acetylglucosamine molecule is joined to the muramylpentapeptide within the cell membrane and the unit is transferred to growth points in the existing peptidoglycan, where adjacent strands are cross-linked. (See also Fig. 2.8.)
The rapid (<5 min) Gram-stain procedure remains a cornerstone of day-to-day practice in detecting and identifying bacteria in clinical laboratories. Gram’s stain distinguishes bacteria as ‘Gram-positive’ or ‘Gram-negative’, according to whether or not they resist decolouration with acetone, alcohol or aniline oil after staining with a triphenyl methane dye, such as crystal violet, and subsequent treatment with iodine. The Gram-positive bacteria resist decolouration and remain stained a dark purple colour. The Gram-negative bacteria are decolorized, and are then counterstained light pink by the subsequent application of safranin, neutral red or dilute carbol fuchsin. In routine diagnostic work a Gram-stained smear is often the only preparation examined microscopically, as it shows clearly the general morphology of the bacteria as well as revealing their Gram reaction. It should be noted that characteristically Gram-positive species may sometimes appear Gram-negative under certain conditions of growth, especially in ageing cultures on nutrient agar or after exposure to antibiotics.
The N-acetylmuramic acid units of peptidoglycan each carry a short peptide, usually consisting of L-alanine, D-glutamic acid, either meso-diaminopimelic acid (in Gram-negative bacteria) or L-lysine (in Gram-positive bacteria) and D-alanyl-D-alanine. The wall is given its strength by cross-links that form between adjacent strands. These may be formed directly between the meso-diaminopimelic acid or L-lysine of one strand and the penultimate D-alanine of the next, or (the usual form in Gram-positive organisms) through an interpeptide bridge composed of up to five amino acids; in either case, the terminal D-alanine is lost in the cross-linking reaction (Fig. 2.8). Several antibiotics interfere with the construction of the cell wall peptidoglycan (see Ch. 5).
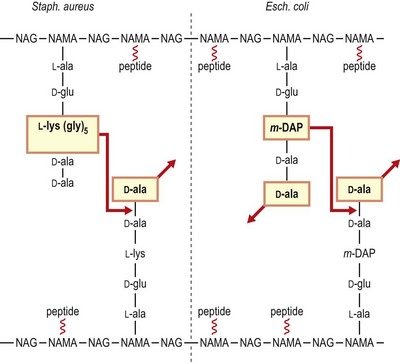
Fig. 2.8 Schematic representation of the peptidoglycan of a representative Gram-positive organism (Staphylococcus aureus) and a representative Gram-negative organism (Escherichia coli). Note that in the Gram-positive bacterium cross-linking occurs through a peptide bridge (pentaglycine in Staph. aureus), whereas direct cross-linking occurs in E. coli. In both cases the terminal D-alanine is lost. Not all peptides are engaged in cross-linking in E. coli, and carboxypeptidases remove redundant D-alanine residues. NAG, N-acetylglucosamine; NAMA, N-acetylmuramic acid; m-DAP, meso-diaminopimelic acid.
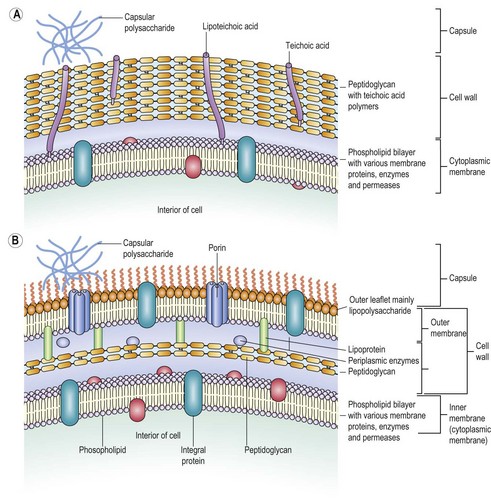
The bacterial cell wall also contains other components whose nature and amount vary with the species. Many Gram-positive bacteria contain relatively large amounts of teichoic acid (a polymer of ribitol or glycerol phosphate complexed with sugar residues) interspersed with the peptidoglycan; some of this material (lipoteichoic acid) is linked to lipids buried in the cell membrane.
Electron microscopy reveals that Gram-negative bacteria possess a second outer membrane external to the peptidoglycan layer. This is essentially another unit membrane in which the outer leaflet is largely composed of a molecule referred to as lipopolysaccharide (LPS). Like the cytoplasmic membrane, this membrane contains many associated proteins whose functions include selective permeability (porins) and attachment (adhesins). The outer membrane confers several important properties on Gram-negative bacteria:
• It protects the peptidoglycan from the effects of lysozyme (a natural body defence substance that cleaves the link between N-acetylglucosamine and N-acetylmuramic acid (see Fig. 2.7).
Components of the LPS, in particular the core structure, lipid A, form endotoxin, which, when released into the bloodstream, may give rise to endotoxic shock (see Ch. 13).
In addition to the basic Gram stain-related properties outlined above, a third type of cell envelope is characteristic of mycobacteria, a group that includes the causal agents of tuberculosis and leprosy. Mycobacteria are related to Gram-positive bacteria, though they can rarely be demonstrated as such. The peptidoglycan layer is covalently linked on its outer aspect to arabinogalactan, which is itself substituted with unique lipids known as mycolic acids. These β-hydroxy fatty acids consist of 60 to 90 carbon residues and, together with non-covalently linked free lipids, form an extremely hydrophobic external layer. This layer has some properties in common with the Gram-negative outer membrane (indeed, porins have recently been detected therein). The whole envelope structure confers the property of acid-fast staining by methods such as the Ziehl–Neelsen (ZN) and phenol–auramine procedures.
The ZN method is of great value in the detection of the tubercle bacillus and other mycobacteria. The mycolic acids referred to above provide a barrier to simple aqueous stains, but when permeability is altered by heating or phenol (or both), concentrated solutions of basic fuchsin, and the fluorescent dyes auramine and rhodamine, can produce well-stained cells that subsequently resist decolorization by strong acid in alcohol. Any decoloured non-acid-fast organisms are counterstained in a contrasting colour with methylene blue or malachite green. Modifications of the ZN method are also useful for the demonstration of bacterial endospores and organisms such as Nocardia spp. and cysts of some protozoa, notably Cryptosporidium spp.
The recognition that some bacterial genera have two boundary lipid layers while others have one has led to the recommendation that the descriptive term, diderm (two skin) be applied to Gram-negative bacteria and to a number of other groups, such as the mycobacteria. In this nomenclature bacteria with a single lipid boundary such as conventional Gram-positive bacteria, are referred to as monoderms.
The cell envelope is a highly dynamic structure in growing bacterial cultures. Its components are subject to rapid turnover (synthesis, assembly, disassembly and degradation) and there is busy molecular traffic in and out of the cytoplasm. Although the diagrammatic and photographic representations here give it a somewhat monolithic and immutable character, the envelope and other surface structures can change very rapidly (within minutes) in response to environmental signals. The cell surface receives and transmits many signals from the surrounding environment, including those involving other bacteria, particularly those belonging to the same strain. This latter phenomenon is known as quorum sensing and appears to be important in regulating gene expression in groups of bacteria.
The structures involved in the molecular traffic through the cell envelope are the subject of intense investigation. In particular, the outward secretion of proteins attracts much current interest. At least seven distinct processes have been identified; all involve impressive macromolecular complexes anchored in the cytoplasmic membrane. Of particular interest here are the type III secretion systems in Gram-negative bacteria. The fully assembled multiprotein complex spans the cytoplasmic membrane, the periplasmic space, the murein sacculus and the outer membrane, and in some cases projects from the cell surface into an adjacent host (human) cell. These impressive delivery systems are capable of injecting effector molecules into the host cell and thereby subvert the latter’s function to the advantage of the microbe.
Extracellular polysaccharides: capsules, microcapsules and loose slime
Many bacteria have been demonstrated to possess a more or less continuous but relatively amorphous layer external to the Gram-negative and Gram-positive envelopes described above. Although these are detected quite readily in some bacteria grown under laboratory conditions, they are somewhat ephemeral in others. These structures appear to be important in mediating contact with potentially hostile environments and may be subject to strict environmental control.
When this layer is fully hydrated and resolvable by light microscopy, it is called a capsule (Fig. 2.9). When it is narrower, and detectable only by indirect, serological means or by electron microscopy, it may be termed a microcapsule. The capsular gel consists largely of water and has only a small content (e.g. 2%) of solids. In most species, the solid material is a complex polysaccharide, although in some species its main constituent is polypeptide.
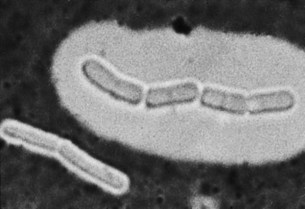
Fig. 2.9 Bacillus megaterium. A chain of bacilli with a large capsule, and a pair with a very small capsule. Wet film with India ink, ×3500.
Loose slime, or free slime, is an amorphous, viscid, colloidal material that is secreted extracellularly by some bacteria. In bacteria that also possess a demonstrable capsule, the slime is generally similar in chemical composition and antigenic character to the capsular substance. When slime-forming bacteria are grown on a solid culture medium, the slime remains around the bacteria as a matrix in which they are embedded, and its presence confers on the growths a watery and sticky ‘mucoid’ character. The slime is freely soluble in water and, when the bacteria are grown or suspended in a liquid medium, it passes away from them and disperses through the medium.
All of these features appear to have some role in interactions with the external environment. In some cases capsules have been shown to protect against phagocytosis, the lytic action of complement and bacteriophage invasion. In at least three instances antibodies directed against capsular antigens have been shown to protect against infection and, indeed, capsular preparations are used in several vaccines. Capsules also appear to have a role in protecting cells against desiccation. The production of extracellular polysaccharides in general provides a matrix within which biofilm formation can take place.
S-layers
A rather more structured (paracrystalline) protein layer has been demonstrated in some bacteria. This S-layer can be shown by electron microscopy and appears to share at least some functional properties with capsules.
Flagella and motility
Motile bacteria possess filamentous appendages known as flagella (singular, flagellum), which act as organs of locomotion. The flagellum is a long, thin filament, twisted spirally in an open, regular wave-form. It is about 0.02 µm thick and is usually several times the length of the bacterial cell. It originates in the bacterial cytooplasm and the structure projects through the cell envelope. According to the species, there may be one, or up to 20, flagella per cell. In elongated bacteria the arrangement of the flagella may be peritrichous, or lateral, when they originate from the sides of the cell, or polar, when they originate from one or both ends. Where several occur on a cell, they may function coiled together as a single ‘tail’. The external portion of a flagellum is essentially a polymer of a single protein, flagellin, whereas the basal region inserted into the cytoplasmic membrane comprises multiple subunits that anchor and power the organ. In a remarkably elegant manner, the flagellar motor is powered directly (as opposed to indirectly via adenosine triphosphate) by the proton gradient created across the cytoplasmic membrane by electron transport. In Escherichia coli, alternation between the anticlockwise and clockwise motion of the flagella effects, respectively, linear or tumbling motility. The intervals between these two patterns are modulated by chemical signals in the environment and the end result is that the bacterium shows chemotactic behaviour (movement towards or away from certain stimuli).
Flagella are invisible in ordinary light microscope preparations, but may be shown by the use of special staining methods, and in special circumstances by dark-ground illumination. Because of the difficulties of these methods, the presence of flagella is commonly inferred from the observation of motility. They can be demonstrated easily and clearly with the electron microscope, usually appearing as simple fibrils without internal differentiation (Fig. 2.10). In some preparations the flagellum appears as a hollow tube formed of helically twisted fibrils, and the flagella of some bacteria (e.g. vibrios) have an outer sheath. In the spirochaetes the flagellum is located in the periplasm and hence is referred to as an endo-flagellum, and this presumably underpins their characteristic spiral motion.
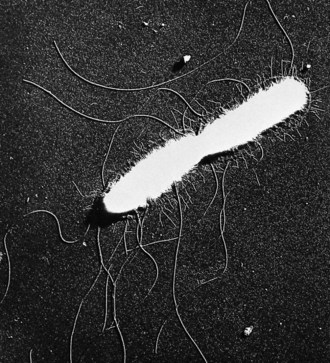
Fig. 2.10 Salmonella enterica serotype Typhi. Dividing bacillus from log-phase culture bears about 15 long wavy flagella and more than 100 short fimbriae. Note the dense (white) protoplast shrunken away from the cell wall. Whole bacillus dried and shadow-cast. Electron micrograph, ×16 000.
From Duguid JP, Wilkinson JF 1961 Environmentally induced changes in bacterial morphology. Symposia of the Society for General Microbiology 11: 69–99.
Motility is clearly important to many bacteria and probably serves mainly to place the cell in environments favourable to growth and free from noxious influences. In some cases possession of flagella is thought to contribute to the pathogenesis of disease.
Fimbriae and pili
Many bacteria possess filamentous appendages called fimbriae or pili. These terms are often used interchangeably, although the latter was originally reserved for structures involved in genetic exchange between bacteria (sex pili; see below). Fimbriae are far more numerous than flagella (e.g. 100–500, being borne peritrichously by each cell), and are much shorter and only about half as thick (e.g. varying from 0.1 to 1.5 µm in length and having a uniform width between 4 and 8 nm). They do not have the smoothly curved spiral form of flagella and are mostly more or less straight. They cannot be seen with the light microscope but are clearly seen with the electron microscope in preparations (see Fig. 2.10). The molecular structure of pili has been studied extensively and reveals many features required to span the bacterial cell envelope that are common to different bacterial groups and difference cells structures.
Multiple types (e.g. types 1 and 2, P type, etc.) of fimbriae have been recognized according to their dimensions, antigenic and phenotypic properties. Until recently it was thought that they were confined to Gram-negative bacteria but now they have been demonstrated in many Gram-positive bacteria. In medical and veterinary contexts, fimbriae are recognized to be important in mediating adhesion between the bacterium and host cells (classically this was recognized in the phenomenon of haemagglutination, a property of type 1, mannose-sensitive pili). In contrast, sex pili are structurally similar to other fimbriae but are longer and confer the ability to attach specifically to other bacteria that lack these appendages. Sex pili initiate the process of conjugation (see Ch. 6); they also act as receptor sites for certain bacteriophages described as being ‘donor specific’.
It should be noted that fimbriae are not the only means by which bacteria can be involved in specific adhesion events. Non-fimbrial adhesins (generally proteins or glyco-proteins) are also important in this regard. Receptor-specific interactions are very important in infective disease, as they are thought to determine much of the tissue tropism of the pathological process.
Importance of microbial surface structures in infection
The structures described have significance for the function of bacteria and their identification by clinical microbiologists, but the surface structures of all micro-organisms (not just bacteria) are of critical importance in the process of infection. They are vital in initiating the contact that occurs during the encounter and establishment of infection (see Ch. 13). Moreover, in addition to substances secreted by micro-organisms, surface structures are exposed to the actions of the innate and adaptive immune systems (see Ch. 8) and, among pathogens, their composition, variability and function reflect these selection pressures and provide opportunities for immunization.
The bacterial ‘life cycle’
Multicellular organisms have long been recognized to pass through many different stages. These may include many immature forms (cf. the larval stages of helminth parasitic worms) or dormant forms (e.g. plant seeds). Even protozoa and fungi may show multiple developmental stages. In contrast, bacteria have been viewed as growing (vegetative), stationary or dead. With the exception of spore-forming genera (see below), because they do not undergo morphological differentiation, bacterial cells have been considered essentially uniform in their properties. As indicated above, it is now recognized that all bacteria adapt extensively and rapidly to their environment. This adaptation takes place at both phenotypic (gene expression) and genotypic (genetic complement and arrangement) levels. It seems clear that there are many more physiological states in which bacteria can exist than previously acknowledged and that these states may influence the capacity of the immune system and antimicrobial agents to eliminate them. In particular, the possibility that non-sporulating bacteria are capable of dormancy (a reversible state of metabolic shutdown) has attracted much interest. Spore formation, however, is the key paradigm of differentiation and dormancy in bacteria.
The different forms taken on by organisms at different stages in the life cycle are of course important in their recognition. The range of basic cellular forms of bacteria was mentioned above. It is difficult to generalize, but important to mention that bacterial cell morphology does alter with the physiological state. Characteristically, the cells of bacteria that are growing rapidly are larger than their non- or slowly growing counterparts. Although this does not alter basic coccal morphology, it may make bacilli appear more intermediate (cocco-bacillary) or spherical (coccoid) in shape. Bacilli exposed to certain noxious influences (notably some antibiotics) may produce extended forms that are sometimes described as filamentous. Bacteria that are characterized as essentially filamentous produce a mat of intertwining filaments known as a mycelium (one characteristic form of fungal growth). This form of growth is also associated with fragmentation in which coccal forms may be released, and this results in highly pleomorphic cultures. All of these alternative growth forms are undoubtedly under the influence of environmental signals, although their identities have yet to be determined in most cases.
Bacterial spores
Some bacteria, notably those of the genera Bacillus and Clostridium, develop a highly resistant resting phase or endospore, whereby the organism can survive in a dormant state through a long period of starvation or other adverse environmental conditions (resuscitation of spores several thousand years old has been claimed). The process does not involve multiplication: in sporulation, each vegetative cell forms only one spore, and in subsequent germination each spore gives rise to a single vegetative cell. Geneticists have viewed sporulation as a paradigm of a simple differentiation process, and the key molecular processes required in Bacillus subtilis are now understood in great detail. In the face of sporulation stimuli, classically starvation or transition from growth to stationary phase, a programme of sequential expression of specific genes is triggered. The end result is a morphologically distinct structure, the endospore, within the mother cell.
In unstained preparations the spore is recognized within the parent cell by its greater refractility. It is larger than lipid inclusion granules and is often ovoid, in contrast to the spherical shape of the lipid granules. Mature ungerminated spores are ‘phase bright’ when viewed by phase-contrast microscopy; immature or germinated spores are ‘phase dark’. When mature, the spore resists coloration by simple stains, appearing as a clear space within the stained cell. Spores are slightly acid-fast and may be stained differentially by a modification of the ZN method. The appearance of the mature spores varies according to the species, being spherical, ovoid or elongated, occupying a terminal, subterminal or central position, and being narrower than the cell, or broader and bulging it. Spores of some species have an additional, apparently loose, covering known as the exosporium (Fig. 2.11).
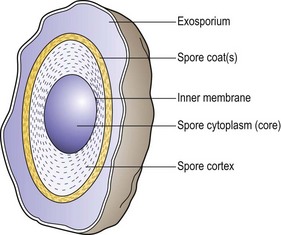
Fig. 2.11 Cross-section of a bacterial spore. The core is surrounded by the inner spore membrane. The cortex, a laminated structure, is protected by a more resistant layer or multiple layers forming the spore coat. In some cases, a loose outer covering (exosporium) can be defined.
Spores are much more resistant than the vegetative forms to exposure to disinfectants, drying and heating. Thus, application of moist heat at 100–120°C or more for a period of 10–20 min may be needed to kill spores, whereas heating at 60°C suffices to kill vegetative cells. In the dry state, or in moist conditions unfavourable to growth, spores may remain viable for many years. The marked resistance of spores has been attributed to several factors in which they differ from vegetative cells: the impermeability of their cortex and outer coat, their high content of calcium and dipicolinic acid, their low content of water, and their very low metabolic and enzymic activity.
Reactivation of the spore is termed germination and it should be noted that this is not just a reversal of the process by which the spore was formed. Germination of the spore occurs in response to specific stimuli that are generally related to external conditions favourable to growth. It is irreversible and involves rapid degradative changes. The spore successively loses its heat resistance and its dipicolinic acid; it loses calcium, it becomes permeable to dyes and its refractivity changes. Spores that have survived exposure to severe adverse influences such as heat are much more exacting than normal spores in their requirements for germination. For this reason, specially enriched culture media are used when testing the sterility of materials, such as surgical catgut, that have been exposed to disinfecting procedures. In the process of germination, the spore swells, its cortex disintegrates, its coat is broken open and a single vegetative cell emerges.
The initiation of germination (activation) is incompletely understood. It is clear that the state of dormancy of spores may be altered by various treatments, such as transient exposure to heat at 80°C, so that germination can then proceed more rapidly in the individual cells or more completely in a spore population. Activation is distinct from germination and is reversible if germination does not proceed.
After germination, cell growth leading up to the formation of the first vegetative cell and before the first cell division is referred to as outgrowth. The conditions required for successful outgrowth may differ markedly from those that allow germination.
Conidia (exospores)
Some of the mycelial bacteria (Actinomycetales) and many filamentous fungi form conidia, resting spores of a kind different from endospores. The conidia are borne externally by abstriction from the ends of the parent cells (conidiophores), and are disseminated by the air or other means to fresh habitats. They are not especially resistant to heat and disinfectants.
Pleomorphism and involution
During growth, bacteria of a single strain may show considerable variation in size and shape, or form a proportion of cells that are swollen, spherical, elongated or pear shaped. This pleomorphism occurs most readily in certain species (e.g. Streptobacillus moniliformis and Yersinia pestis) in ageing cultures, on artificial medium and especially in the presence of antagonistic substances such as penicillin, glycine, lithium chloride, sodium chloride in high concentrations, and organic acids at low pH. The abnormal cells are generally regarded as degenerate or involution forms; some are non-viable, whereas others may grow and revert to the normal form when transferred to a suitable environment. In many cases the abnormal shape seems to be the result of defective cell wall synthesis and produces a grotesquely swollen cell, comparable to a spheroplast (see below), that later usually bursts and lyses.
Spheroplasts, protoplasts and L-forms
If bacteria have their cell walls removed or weakened while they are held in a solution of sufficient osmolarity to prevent them taking up water by osmosis, they may escape being lysed and, instead, may become converted into viable spherical bodies. If all the cell wall material has been removed from them, the spheres are free protoplasts. If they remain enclosed by an intact, but weakened, residual cell wall, they are called spheroplasts. Protoplasts and spheroplasts are osmotically sensitive; they vary in size with the osmotic pressure of the suspending medium and, if the medium is much diluted, they swell up and perish by lysis. In contrast to these laboratory generated forms (which may be made deliberately for research or biotechnological applications), L-forms of bacteria may arise spontaneously and are also cell wall deficient. There is much controversy about the contribution of L-forms to infection. They are difficult to demonstrate as they do not stain with Gram or acid-fast methods and may not propagate in vitro. In common with another controversial area in bacteriology, the ‘nanobacteria’, L-forms may pass through standard bacteria-stopping filters.
The nature and composition of viruses
Structure
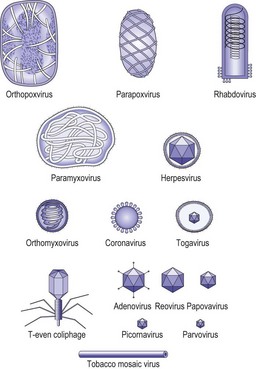
The basic infectious particle of a virus is known as the virion. In the simplest viruses this consists of nucleic acid and a surrounding coat of protein called the capsid. Some viruses are enclosed within an envelope, derived from host cell membranes but modified by the inclusion of viral glycoproteins. The capsid is composed of distinct morphological units or capsomeres, which are assembled from viral proteins. Depending on the arrangement of these proteins, the capsomeres may be spherical, cylindrical or ring-like in appearance. The nucleocapsid is the combination of nucleic acid and capsid. The arrangement of the capsomeres around the nucleic acid determines the symmetry of the virion. When the capsomeres are applied directly to the helical nucleic acid, a coil-like structure with the appearance of a hollow tube is formed. Viruses with this arrangement are said to have helical symmetry. Most helical viruses enclose the nucleocapsid within an envelope and thus do not have a rigid appearance. The other major type is shown by the viruses with icosahedral symmetry (Fig. 2.12), in which the capsomeres are arranged as if lying on the faces of an icosahedron with 20 equilateral triangular faces and 12 corners or apices (Fig. 2.13). Capsomeres on the faces and edges of this figure are called hexons, as they always link with six adjacent capsomeres; those positioned at the apices are the pentons, as they always join to five capsomeres. Viruses with icosahedral symmetry have a rigid structure and, under the electron microscope, have a characteristic hexagonal outline with triangular faces. However, if the diameter of the virion is less than about 50 nm the particle will appear spherical. Many viruses with icosahedral symmetry are enclosed by an outer envelope. The poxviruses are large and complex, and do not show either type of symmetry; they are referred to as complex. The size of virions varies considerably, from 25–300 nm, in different families (Fig. 2.14).
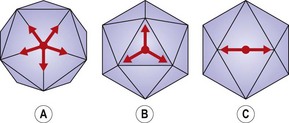
Fig. 2.12 An icosahedron viewed along its (A) five-fold, (B) three-fold and (C) two-fold axes of symmetry.
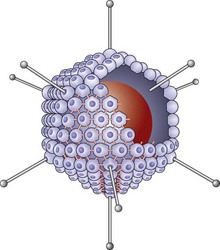
Fig. 2.13 Icosahedron of an adenovirus. The core of DNA is represented by a circular mass. Some of the pentamers at the 12 vertices have been indicated with protruding fibres and terminal knobs. The remaining 240 hexamer capsids are, for the most part, shown as compressed into hollow spheres linked to one another by divalent bonds. The hexagonal shape of a few of the capsomeres is seen in the centre of the diagram.
Viral nucleic acid
The most common types of nucleic acid in viruses of human beings are single-stranded RNA and double-stranded DNA. However, both double-stranded RNA and single-stranded DNA occur in the reoviruses and parvoviruses, respectively. The genomes of RNA viruses may be present as a single strand as in paramyxoviruses, or as two copies as in the retroviruses, or exist as a specific number of fragments as in the orthomyxoviruses and reoviruses. Circular molecules of DNA are present in the virions of papovaviruses and hepadnaviruses. The amount of nucleic acid in virions is constant for a particular virus but shows considerable variation; thus the genome can vary from 5 kilobase pairs in parvoviruses to 375 kilobase pairs in the largest poxviruses.
Virion enzymes
Several viruses carry essential enzymes in the virion. As discussed in Chapter 7, an RNA-dependent RNA polymerase or transcriptase is an essential component of the virion in several virus families, including the negative-strand RNA viruses. Among DNA viruses only the poxviruses carry a DNA-dependent RNA polymerase. The hepadnaviruses have a virion polymerase complex that has some similarity to the reverse transcriptase complex found in the retroviruses.
Viral proteins
Analysis of the proteins produced in a cell during viral infection shows that some are essential components of the virion; these are the structural proteins and include capsid proteins and enzymes as well as basic core proteins that may be necessary to package the nucleic acid within the capsid. Other proteins such as enzymes are needed for the production of viral components but are not part of the virion; these are the non-structural proteins. The essential steps of virus attachment and penetration of the host cell are known to depend on regions of the outer capsid, such as the apical fibres and knobs of adenoviruses, or on parts of the envelope glycoproteins of viruses, such as influenza A and B and the human immunodeficiency virus.
Viroids, defective viruses and prions
Our concept of the organismal nature of infectious agents is stretched to the limit by these infective entities. Viroids are essentially circular RNA molecules that have been associated with several plant diseases; they do not encode proteins or possess a capsid. The hepatitis delta agent has some features in common with viroids and defective viruses (viruses that need the help of another virus for the formation of infectious particles). In the case of the delta agent these features result in an infective agent that can be transmitted in parallel with hepatitis B.
Prions are proteinaceous infective agents that are responsible for the transmissible spongiform encephalopathies (see Ch. 60). An increase in the number of prion proteins in a new host seems to result from the capacity of the introduced protein to induce abnormal conformational changes in a closely related host protein, rather than by replication. Accumulation of the protein in the induced conformation produces the characteristic pathology of the disease.
Blair DF. How bacteria sense and swim. Annual Review of Microbiology. 1995;49:489–522.
Cabeen MT, Jacobs-Wagner C. Skin and bones, the bacterial cytoskeleton, cell wall, and cell morphogenesis. Journal of Cell Biology. 2007;179:381–387.
Frey D, Oldfield RJ, Bridger RC. A Colour Atlas of Pathogenic Fungi. London: Wolfe Medical; 1979.
Li Z, Jensen GJ. Electron cryotomograpy: a new view into microbial ultrastructure. Current Opinion in Microbiology. 2009;12:333–340.
Madeley CR, Field AM. Virus Morphology, ed 2. Edinburgh: Churchill Livingstone; 1988.
Moat AG, Foster JW. Microbial Physiology, ed 4. New York: Wiley-Liss; 2002.
Neidhardt FC, Ingraham JL, Schaechter M. Physiology of the Bacterial Cell: A Molecular Approach. Sunderland, MA: Sinauer; 1990.
Nikaido H, Vaara M. Molecular basis of bacterial outer membrane permeability. Microbiological Reviews. 1985;49:1–32.
Olds RJ. A Colour Atlas of Microbiology. London: Wolfe Medical; 1975.
Proft T, Baker EN. Pili in Gram negative and Gram positive bacteria – structure, assembly and their role in disease. Cellular and Molecular Life Sciences. 2009;66:613–635.
Wadhams GH, Armitage JP. Making sense of it all: Bacterial chemotaxis. Nature Reviews in Molecular Cell Biology. 2004;5:1024–1037.
CELLS alive! http://www.cellsalive.com/
Tree of Life Web Project http://tolweb.org/tree/
University of California, Berkeley. More on bacterial morphology http://www.ucmp.berkeley.edu/bacteria/bacteriamm.html